Back to article: Detection of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and its first variants in fourplex real-time quantitative reverse transcription-PCR assays
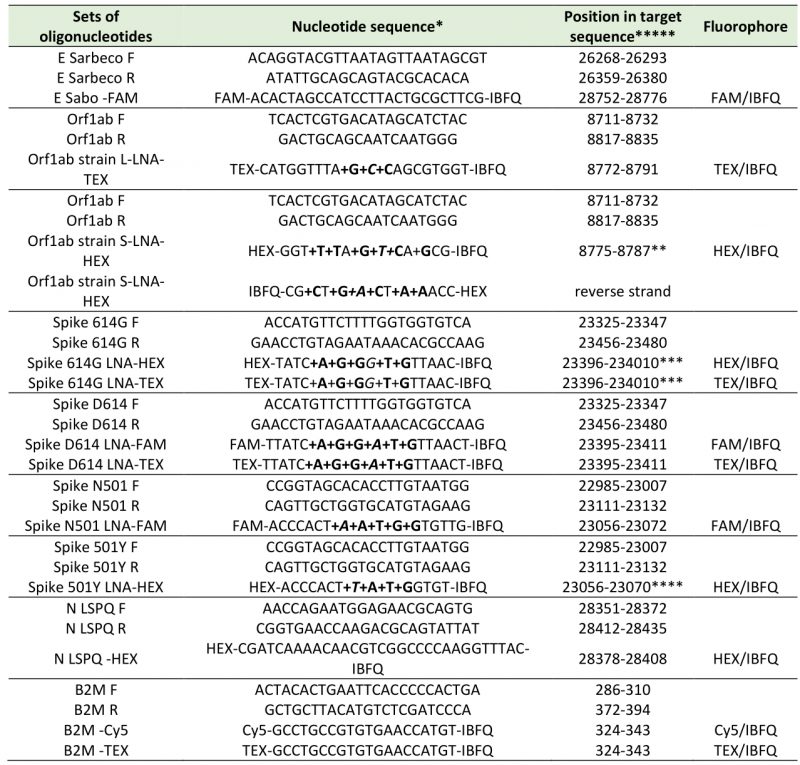
TABLE 1. Oligonucleotide primers and fluorogenic probes used in real-time quantitative RT-PCR assays.
*: LNA-modified nucleotides are designated with prefix ‘+’ and are in bold, e.g. +C for a LNA-modified C.
**: position with variant mismatch (italic) (C instead of T at position 8782) [23].
***: position with variant mismatch (italic) (G instead of A at position 23403) [26].
****: position with variant mismatch (italic) (T instead of A at position 23063) [25].
*****: NC_045512: Reference sequence of SARS-CoV-2 isolate Wuhan-Hu-1, complete genome [1].
NM_004048 : Reference sequence of homo sapiens beta-2-microglobulin (B2M), mRNA.
1. Wu F, Zhao S, Yu B, Chen YM, Wang W, Song ZG, Hu Y, Tao ZW, Tian JH, Pei YY, Yuan ML, Zhang YL, Dai FH, Liu Y, Wang QM, Zheng JJ, Xu L, Holmes EC, and Zhang YZ (2020). A new coronavirus associated with human respiratory disease in China. Nature 579:265-269. 10.1038/s41586-020-2008-3
23. Tang X, Wu C, Li X, Song Y, Yao X, Wu X, Duan, Y, Zhang, H, Wang, Y, Qian, Z, Cui, J, and Lu, J (2020). On the origin and continuing evolution of SARS-CoV-2. Nat Sci Rev 7:1012-1023. 10.1093/nsr/nwaa036
25. Yi C, Sun X, Ye J, Ding L, Liu M, Yang Z, Lu X, Zhang Y, Ma L, Gu W, Qu A, Xu J, Shi Z, Ling Z, and Sun B (2020). Key residues of the receptor binding motif in the spike protein of SARS-CoV-2 that interact with ACE2 and neutralizing antibodies. Cell Mol Immunol 17:621-630. 10.1038/s41423-020-0458-z
26. Korber B, Fischer WM, Gnanakaran S, Yoon H, Theiler J, Abfalterer W, Hengartner N, Giorgi EE, Bhattacharya T, Foley B, Hastie KM, Parker MD, Partridge DG, Evans CM, Freeman TM, and de Silva TI; Sheffield COVID-19 Genomics Group, McDanal C, Perez LG, Tang H, Moon-Walker A, Whelan SP, LaBranche CC, Saphire EO, and Montefiori DC (2020). Tracking changes in SARS-CoV-2 Spike: evidence that D614G increases infectivity of the COVID-19 virus. Cell 182:812-827.e19. 10.1016/j.cell.2020.06.043