Back to article: The logics of metabolic regulation in bacteria challenges biosensor-based metabolic engineering
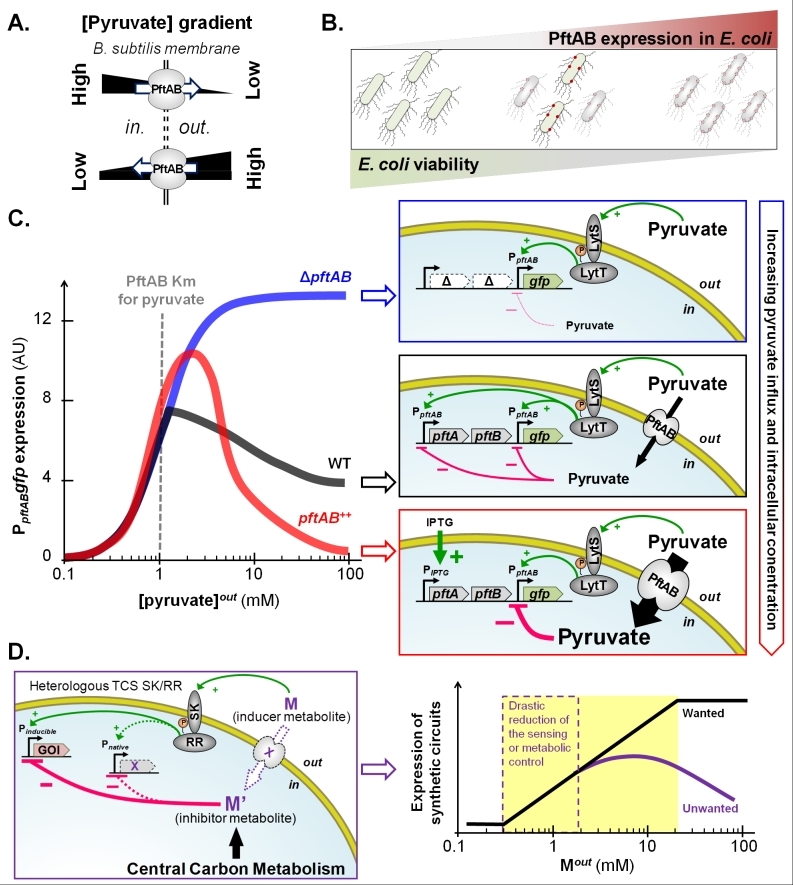
FIGURE 1: The bacterial pyruvate transport system PftAB and its complex regulation by the two-component system LytST. (A) The pyruvate facilitated transporter PftAB of B. subtilis can either import or export pyruvate depending on the concentration gradient of pyruvate across the cell membrane. (B) E. coli cell death or cell growth inhibition increases with the level of heterologous expression of pftAB. (C) Expression of pftAB in WT (grey), ΔpftAB mutant (blue), and pftAB over-expressing (red) B. subtilis cells grown in minimal medium with glutamate and succinate as carbon source, and pyruvate concentrations ranging from 0.1 to 100 mM (left panel). Extracellular pyruvate acts as the signal molecule for LytST, which induces expression of pftAB. However, when the pyruvate influx is high, LytST activity is drastically retro-inhibited. Consistently, in the ΔpftAB mutant, the level of induction is maximal as there is no influx of pyruvate (right panel). (D) In metabolic engineering, the expression of one (or more) gene(s) of interest (GOI) is (are) under the control of promoter(s) that can be activated by the use of inducer metabolite(s) (M). The activity of a heterologously expressed TCS may be retro-inhibited by the inducer or derivative metabolites (M or M’) if naturally present in the host cell (left panel). As a result, the TCS-induced expression of a synthetic circuit will not exhibit a log-linear dose response as M increases. The distortion between the expected (wanted) and effective (unwanted) induction challenges the rational design of novel nature-inspired sensors (right panel).