Research Articles:
Microbial Cell, Vol. 3, No. 10, pp. 511 - 521; doi: 10.15698/mic2016.10.534
Phylogenetic profiles of all membrane transport proteins of the malaria parasite highlight new drug targets
1 Department of Immunology, Max Planck Institute for Infection Biology, Berlin, Germany.
2 Department of Medical Microbiology & Centre for Molecular and Biomolecular Informatics, Radboud Institute for Molecular Life Sciences, Radboud University Medical Centre, Nijmegen, The Netherlands.
Keywords: drug target, experimental genetics, malaria parasite, membrane transport protein, orthology, phylogeny, Plasmodium.
Received originally: 01/04/2016 Received in revised form: 30/07/2016
Accepted: 01/08/2016
Published: 30/08/2016
Correspondence:
Taco W.A. Kooij, Department of Medical Microbiology & Centre for Molecular and Biomolecular Informatics, Radboud Institute for Molecular Life Sciences, Radboud University Medical Centre; P.O. Box 9101, 6500 HB Nijmegen, The Netherlands taco.kooij@radboudumc.nl
Conflict of interest statement: The authors declare no existing conflicts of interest.
Please cite this article as: January Weiner 3rd and Taco W.A. Kooij (2016). Phylogenetic profiles of all membrane transport proteins of the malaria parasite highlight new drug targets. Microbial Cell 3(10): 511-521.
Abstract
In order to combat the on-going malaria epidemic, discovery of new drug targets remains vital. Proteins that are essential to survival and specific to malaria parasites are key candidates. To survive within host cells, the parasites need to acquire nutrients and dispose of waste products across multiple membranes. Additionally, like all eukaryotes, they must redistribute ions and organic molecules between their various internal membrane bound compartments. Membrane transport proteins mediate all of these processes and are considered important mediators of drug resistance as well as drug targets in their own right. Recently, using advanced experimental genetic approaches and streamlined life cycle profiling, we generated a large collection of Plasmodium berghei gene deletion mutants and assigned essential gene functions, highlighting potential targets for prophylactic, therapeutic, and transmission-blocking anti-malarial drugs. Here, we present a comprehensive orthology assignment of all Plasmodium falciparum putative membrane transport proteins and provide a detailed overview of the associated essential gene functions obtained through experimental genetics studies in human and murine model parasites. Furthermore, we discuss the phylogeny of selected potential drug targets identified in our functional screen. We extensively discuss the results in the context of the functional assignments obtained using gene targeting available to date.
INTRODUCTION
The malaria parasite has adopted a highly complex life cycle involving a continuous switching between vertebrate hosts and anopheline mosquitoes. Within humans, Plasmodium species are obligate intracellular parasites moving through three different life-cycle stages. After an infectious mosquito bite, a single phase of preclinical growth within the host liver cells commences [1]. Next, fast proliferating blood-stage parasites are the cause of malaria-associated pathology and severe disease outcome [2]. Finally, some of the asexual blood-stage parasites are triggered to develop into male or female gametocytes, which are required for sexual reproduction following a mosquito blood meal [3][4].
–
Despite a gradual decline in annual malaria cases and deaths, the parasite remains one of the largest global killers. In 2015, WHO reported 150-300 million cases and 438,000 deaths [5]. Difficulty in combating the disease is exacerbated by the growing resistance to anti-malarial drugs and the absence of an effective vaccine. The continuous need for new therapeutic anti-malarial drugs is unquestioned. Yet, due to limited availability, there is a much more pressing need for drugs that can prevent infections by acting prophylactically on the liver stage of the parasite, and transmission-blocking compounds, which kill the gametocytes thus helping to prevent the spread of the disease. Chemoprophylaxis is not only important for travellers [6] but also particularly for women in endemic areas in their first or second pregnancy [7]. Thus, there is a critical need for novel drugs that may be used prophylactically, therapeutically, or to block transmission [8].
–
As defined by the Medicines for Malaria Venture in there 2015 annual report [9], ideal medicines for treatment and protection would both be suitable for mass drug administration programs and require a single encounter treatment, or better still a single exposure treatment, to help improve compliance. Treatment of infection should be effective against all life-cycle stages of all five malaria species infecting humans, including resistant strains, and resistance against the new drug should be difficult to achieve.
–
Generally, it is believed that these features may be best obtained by the combination of at least two active compounds, one fast acting for immediate clearance of the infection and a second, slower-acting compound providing long duration of efficacy. Furthermore, an ideal treatment would be gametocytocidal to prevent the spread of infection to mosquitoes, while sporontocidal or liver-stage activity would provide a prophylactic component. To complete the wish list of the ideal anti-malarial drug, it should also be active against the so-called hypnozoites, the dormant stages of certain malaria parasite species (most notably P. vivax) that are the cause of relapse infection even many years after the infectious mosquito bite. It is obvious that the development of the ideal anti-malarial drug will not be straightforward and will result in a compromise between the long list of desirable attributes and features. In this light, it is particularly important that on-going functional studies of malaria parasite biology continue to highlight potential new drug targets that have important roles in the different life-cycle stages of the parasite and are conserved among all malaria parasite species but are absent from or have diverged significantly in humans, such that compounds acting on these important Plasmodium proteins may do so effectively as well as selectively.
–
Plasmodium membrane transport proteins (MTP), such as the chloroquine resistance transporter (CRT) and the ATP-binding cassette (ABC) transporter family, including the multidrug resistance proteins (MDR) and the multidrug resistance-associated proteins (MRP), are well known for their roles in anti-malarial drug resistance [10][11]. MTPs are also generally considered potential drug targets in their own right [12][13]. Spiroindolones and dihydroisoquinolones are new classes of potent anti-malarial drugs, currently under clinical testing, that have both been shown to act via the Plasmodium falciparum cation ATPase, ATP4, causing severe disturbance of Na+ homeostasis in the parasite [14][15][16][17].
–
To survive and thrive, malaria parasites utilize a range of transport processes to import nutrients, export waste, and redistribute ions and small organic molecules between different sites and organelles [12][18]. MTPs of different classes facilitate these processes. Following the functional and phylogenetic classification of MTPs from the Transporter Classification Database (http://www.tcdb.org) [19], Plasmodium MTPs can be classified as: α-type channels and β-barrel porins (TCDB Class 1.A/B); P-P-bond hydrolysis-driven transporters, here referred to as pumps (TCDB Class 3.A); porters, including uniporters, symporters, and antiporters (TCDB Class 2.A); and unclassified, putative MTPs.
–
Experimental genetics is a powerful means to further explore critical functions of Plasmodium MTPs for parasite survival throughout its complex life cycle [20][21]. Due to more efficient experimental and computational methods and access to the entire in vivo life cycle, the majority of such studies have been performed using murine malaria model species, notably Plasmodium berghei, and have focussed on single MTPs (Table S1). Two of our most recent studies have more than doubled the number of targeted genes, and have generated loss-of-function mutants in both P. falciparum and P. berghei [22][23]. A systematic study of the MDR family demonstrated that four of seven members fulfil essential functions during blood-stage development, highlighting these as potential drug targets [22]. While targeting 35 orphan MTPs, Kenthirapalan et al. produced the largest collection of P. berghei knock-out parasites available to date [23]. The 29 available mutant lines provide a powerful resource for further studies of malaria parasite transport processes. In addition to highlighting six genes essential for blood-stage survival including five pumps, they also revealed potential prophylactic (MFS6) and transmission-blocking (ZIP1) drug targets [23].
–
Here, we performed comprehensive orthology profiling by reciprocal Blast of all identified 139 Plasmodium MTPs (consisting of the list published by Martin et al. [24] expanded with newly identified candidates) against a selection of 41 species from the entire breadth of the eukaryotic kingdom. To further validate the potential of the eight newly identified drug targets, we have explored their phylogenetic relationships in detail. These results are discussed extensively in the context of available insights from functional genetics studies of malaria parasite MTPs.
RESULTS AND DISCUSSION
Comprehensive orthology assignments of Plasmodium MTPs
To identify the levels of conservation of the 139 P. falciparum MTPs (Table S1) within the eukaryotic domain, we first performed an extensive orthology assignment using Blast against a set of 41 species (Table S2). We selected a subset of species to represent the enormous diversity in the eukaryotic domain by including sequences from all kingdoms, with an emphasis on protozoan species, commonly used model species, and parasite species of great medical or veterinary importance. These include the clinically second most important human malaria parasite Plasmodium vivax and the murine malaria model parasite P. berghei. Plasmodium species belong to a large monophyletic group of largely obligate intracellular parasites, the Apicomplexa. We included six additional apicomplexan parasites in our analysis: two piroplasms (Babesia bovis and Theileria annulata) that cause cattle fever and like Plasmodium species belong to the Aconoidasida, and four Conoidasida, including parasites of medical (Toxoplasma gondii and Cryptosporidium parvum) and veterinary (Neospora caninum and Eimeria tenella) importance.
–
Apicomplexan parasites are characterized by a complex at the apical end that is used to penetrate and enter a wide variety of host cells. However, this is not the only peculiar subcellular structure of these eukaryotic single cell parasites. Being eukaryotes, one would naturally expect most of the commonly shared organelles to be present, but while most do retain a single and thoroughly reduced mitochondrion, some have only vague remnants of the cell’s power house in the form of mitosomes [25]. An additional organelle of endosymbiotic heritage present in the majority of apicomplexan parasites (though it has been lost in Cryptosporidium species) is the former-photosynthetic apicoplast [26]. This plastid is of red algal origin, a trait shared across a variety of highly divergent species, including other alveolates, such as Chromerida and dinoflagellates, but not ciliates. For our analysis, we included four alveolates, including two ciliates and one of the closest relatives of the Apicomplexa, Vitrella brassicaformis.
–
The true evolutionary relationships between highly diverse protozoan lineages are unresolved and still a matter of dispute but many interesting parallels can be observed. Thus Cryptomonads, Haptophyta, and a significant proportion of Stramenopiles, all harbour plastids of red algal origin, although at present it is not clear whether these plastids originate from single or multiple secondary endosymobiotic events [27][28][29]. In our analysis, we included one cryptomonad, Guillardia theta, as well as four stramenopiles including the oomycete Phytophthora infestans. The latter is a plant parasite causing potato blight, which does not harbour a plastid, but it does share another interesting feature with malaria parasites. Both oomycetes and malaria parasites extensively remodel the host they infect by exporting a large repertoire of effector proteins using comparable strategies [30]. Further protist representative sequences were taken from two Rhizaria, six Excavata, including three kinetoplastid parasites, and two Amoebozoa.
–
From the plant kingdom, we took a green and a red alga and the most commonly used model plant Arabidopsis thaliana. Ophistokonts were represented by two fungi (baker’s yeast and the pathogenic Cryptococcus neoformans) and by nine animals, including the parasitic flatworm Schistosoma haematobium, the malaria mosquito Anopheles gambiae, a number of widely used model organisms, and of course mouse and human.
–
Potential prophylactic drug targets
In our recent functional screen of orphan MTPs, we identified a critical role for a major facilitator superfamily member, MFS6, during liver-stage development, in addition to important functions in the blood [23]. Though we identified a putative orthologue in V. brassicaformis (Figure 1), MFS6 is largely Plasmodium-specific and appears absent from humans. Its important functions during both liver- and blood-stage development justify further exploration of the protein as a target for compounds with combined prophylactic and therapeutic activity. P. berghei MRP2-deficient parasites demonstrate a complete arrest in the liver, a phenotype that was also observed in P. falciparum counterpart MRPs [31]. Like MFS6, MRP2 appears to be largely Plasmodium-specific (Figure 1) providing another promising drug target. However, for these two MTPs both their function and their respective substrates remain unresolved.
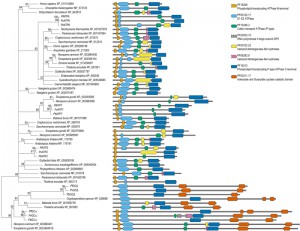
 | FIGURE 1A: Orthologies of membrane transport proteins of the malaria parasite. Extensive amino acid based reciprocal homology searches were performed to establish orthologues of the all Plasmodium channels/pores (TCDB Class 1.A/B [19]; pink), pumps (3.A; purple), porters (2.A; cyan), and unclassified, putative MTPs (yellow) in representative species of the entire breadth of the eukaryotic domain. Dot sizes indicate the fraction of the sequence length over which homology was detected; red dots indicate reciprocal orthologues. When a protein has not been given a name, these are indicated by their P. falciparum geneID number. MTPs for which the encoding genes were deleted successfully are indicated in blue, unsuccessful gene deletions suggesting essential functions during blood-stage development are highlighted in red. (®, species harbouring plastids of red algal origin; *, putative heavy metal transporting MTPs) Please see the continuation of the table on the next page. |
 | FIGURE 1B: Orthologies of membrane transport proteins of the malaria parasite. Extensive amino acid based reciprocal homology searches were performed to establish orthologues of the all Plasmodium channels/pores (TCDB Class 1.A/B [19]; pink), pumps (3.A; purple), porters (2.A; cyan), and unclassified, putative MTPs (yellow) in representative species of the entire breadth of the eukaryotic domain. Dot sizes indicate the fraction of the sequence length over which homology was detected; red dots indicate reciprocal orthologues. When a protein has not been given a name, these are indicated by their P. falciparum geneID number. MTPs for which the encoding genes were deleted successfully are indicated in blue, unsuccessful gene deletions suggesting essential functions during blood-stage development are highlighted in red. (®, species harbouring plastids of red algal origin; *, putative heavy metal transporting MTPs) Please see the beginning of the table above. |
Evidence is mounting that transport processes of metal ions, in particular of heavy metals such as iron, copper, and zinc, could provide efficient targets for chemoprophylactic treatments. Although the exact mechanisms are unclear, P. berghei liver-stage development is influenced by host iron homeostasis [32][33]. Recently, a vacuolar iron transporter was described that plays an important (although not critical) role during liver infection in vivo and in vitro [34]. Parasites lacking an alternative zinc–iron permease (ZIPCO) are severely affected in liver-stage development in vitro and show a delay in prepatency of two days [35]. A similar delay in prepatency was observed following needle injection of ctr2– sporozoites despite developing normally in culture [23]. Interestingly, these parasites were most severely affected during natural transmission by infectious mosquito bites or following subcutaneous injection of the sporozoites. Parasites deficient in the copper channel 1 gene (CTR1) completely failed to transmit, but this was at least in part attributable to a much reduced and delayed sporozoite production, and the infectivity of these sporozoites remains to be determined in more detail to establish whether CTR1 like CTR2 plays an important role in the liver [23]. Despite the fact that the putative heavy metal transporting MTPs with a demonstrated important role during the establishment of new infection are not strictly essential, they may still prove interesting targets for prophylactic interventions, since none of the MTPs discussed above appear to have a clear reciprocal orthologue in humans and show only partial sequence matches (Figure 1).
–
Potential transmission-blocking drug targets
The importance of heavy metal homeostasis appears not to be restricted to mosquito-to-mouse/human transition. Also sexual blood-stage parasites, parasite fertility, and effective colonization of the mosquito midgut appear to be strongly dependent on the correct distribution of these cations.
–
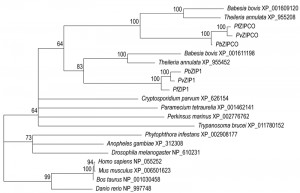
Where the two copper channels may act in copper transport in the liver stages, a copper-transporting P-type ATPase (CuTP) was shown to be central to fertility of both male and female gametes [36]. Activation of cutp– male microgametes, a process known as exflagellation, is reduced to ~10% of wild-type and this reduction could be phenocopied using an intracellular copper chelator. Cross-fertilization studies and an even more pronounced reduction in oocyst numbers indicate additional important roles in the female gametes. ZIP1, a paralogue of the liver stage-specific ZIPCO, is crucial for mosquito colonization [23]. Exflagellation in zip1– parasites is reduced to naught, which is directly attributable to a nearly complete absence of male gametocyte formation. Since ZIP1-deficient parasites also have a slightly reduced blood-stage multiplication rate, ZIP1 is potentially a very attractive transmission-blocking drug target. Indeed, if a compound would be able to target ZIPCO and ZIP1, this would be a triple-acting drug that could be used prophylactically, therapeutically, and to block spread of the disease. One disadvantage could be that humans harbour fourteen zinc-iron permeases [37], although only one of these bears any sequence similarity to the Plasmodium copies. Nevertheless, caution is needed to ensure that the compound is sufficiently parasite-specific. Phylogenetic profiling of the identified ZIP sequences supports a relatively recent gene duplication at the root of the Aconoidasida (Figure 2), and the relatively long distance from the vertebrate ZIPs suggest that specific targeting of the Plasmodium ZIPs might be feasible.
Other MTPs with roles during mouse/human-to-mosquito host transition have been identified, such as the Ca2+/H+ exchanger (CAX) [38], a cation diffusion factor or putative zinc transporter (CDF) [23], and the pantothenate transporter [23][39]. Unfortunately, not a single MTP has been identified that is critical for both male and female gametocyte formation, although it remains to be tested if zip1– female gametocytes remain fertile or have lost the capacity to reproduce.
–
Potential therapeutic drug targets
Two types of channels have been discussed as potential therapeutic drug targets. Two potassium channels (K1 and K2) were refractory to gene deletion in P. falciparum [40]. However, absence of supporting evidence that this was not merely due to the technical difficulties of targeting P. falciparum genes and the fact that PbK1 could be readily deleted [41] suggest that these channels may not be strictly necessary for blood-stage survival. The suitability of aquaglyceroporin (AQP) has also been subject of controversy. Initial studies demonstrated that blood-stage growth of aqp– parasites was strongly affected [42]. Independently, we were only able to replicate a minor defect in Swiss-Webster mice [43]. Using a sensitive flow cytometry – based method in NMRI mice, we saw no difference in growth rates of WT and aqp– parasites either growing in direct competition or in individual mice [44]. The relevant difference between NMRI and Swiss-Webster mice that may lead to this observation is unclear.
–
Interestingly, a vast majority of the currently identified resistance markers, e.g. the ABC family [11], as well as the single validated druggable MTP, ATP4 [16][17], are primary active transporters that require ATP to fuel their activity. As the parasite invests energy in their functioning, it is perhaps not surprising that many pumps play crucial roles at some stages during the parasite’s life cycle. Indeed, of all nineteen targeted pumps, eleven were shown to be refractory to gene deletion (Table S1). In addition to the four MDRs (MDR1, MDR4, MDR6, and MDR7) [22], these include the putative cation transporting ATPase, ATP4, (S. Kenthirapalan, K. Matuschewski, T.W.A. Kooij, unpublished data), ATP6 [45] and the V-type proton ATPase subunit G [46], ABCI3, and four predicted aminophospholipid transporters [23].
–
Our orthology profiling indicates that the ABC transporters, with the exception of MDR1 (and to lesser extent MDR2), are poorly conserved across the eukaryotic kingdom, further highlighting their potential as anti-malarial drug targets (Figure 1). ABCI3 appears to be a unique, Plasmodium-specific ABC family member and is characterized by the presence of two transmembrane domains consisting of multiple transmembrane helices interspersed by a single nucleotide-binding domain. An extensive phylogenetic profiling of all ABC transporters identified in 16 eukaryotic and 55 prokaryotic genomes assigned PfABCI3 to a poorly supported clade together with sequences from two archaea from two different kingdoms and sequences from five bacteria from four different phyli [47]. Thus, the orthology and phylogeny of ABCI3, along with demonstrated essential function during blood-stage development, strengthen its potential as a therapeutic drug target.
–
Another class of primary active transporters that was highlighted for its druggable potential consists of putative aminophospholipid-transporting P4-type ATPases, from hereon referred to as flippases [23]. In addition to ATP2, ATP7, and ATP8, these also include two putatively bifunctional proteins that also harbour guanylyl cyclase (GC) activity [48]. GCα, like the other flippases, was essential for blood-stage development, whereas GCβ plays a critical role in colonization of the mosquito midgut [23][49][50]. While parts of the GCs are conserved, no bifunctional orthologues were found outside the apicomplexan clade (Figure 1). The phylogenetic tree of the flippases confirmed this notion (Figure 3). Orthology and phylogenetic profiles further indicate that ATP2 and ATP8 are rather well conserved including human orthologues, whereas ATP7 is largely apicomplexan-specific (Figure 1). Combined, these data suggest that ATP7 and GCα may form attractive targets for novel anti-malarial compounds. Of note, for one P. falciparum gene encoding a putative flippase (PF3D7_1468600), no orthologue exists in P. berghei and a possible essential role has not yet been established.
Of the largest group of MTPs, the porters (TCDB Class 2.A) [19], 28 genes have been targeted in P. berghei, only three times without success (Table S1). In addition to the well-studied CRT [51], these include a hexose transporter HT [52], which is well conserved including in humans (Figure 1), and the putative drug metabolite transporter DMT2 [23]. The latter presents a particularly interesting case considering the orthology discovery. Orthologues were identified in all apicomplexan parasites, with the exception of Cryptosporidium, in other alveolates, but not ciliates, and in the cryptomonad, G. theta, and most stramenopiles. Most species have a single DMT2 but T. gondii, N. caninum, and V. brassicaformis have multiple copies. While the exact phylogenetic relationships of the alveolates, cryptomonads, and stramenopiles is still a topic of debate, many of these chromalveolates, as they are commonly referred to, harbour a plastid of red algal origin. Ciliates and Cryptosporidium species do not have such a plastid and thus it is tempting to speculate that DMT2 localizes to this endosymbiotic organelle. Indeed, the red alga Cyanidioschizon merolae appears to harbour a sequence with a very weak similarity that was initially just below the cut-off E-value applied to establish significant homologies. However, the presence of a rather well conserved copy in P. infestans, that lacks a relic plastid, and presence of weak similarities in three other unrelated eukaryotes (Giardia lamblia, Dictyostelium discoideum, and Chlamydomonas reinhardtii) suggest that the evolutionary history of this gene may be more complicated. When building a maximum likelihood tree for all identified homologues, including the few hits from unrelated species, the tree does not resolve well (data not shown). However, when only including sequences from the chromalveolates and the red alga, the tree is well-supported showing phylum-specific clades with Cyanidioschizon merolae as the outmost group of the tree (data not shown). Attempts to include other sequences as outgroup, e.g. Plasmodium DMT1 sequences, were unsuccessful due to a lack in sequence similarity and the consequently poorly resolving sequence alignments. The most consistent results were obtained when using a single DMT2 homologue of one of the three unrelated species as an outgroup (Figure 4).
Despite the significant uncertainties and remaining unresolved questions about the origin of DMT2, these data could well suggest that this essential MTP is localized in the parasite’s apicoplast. Our initial localization studies were hampered by very low expression levels of the protein and hence difficult to interpret, but appeared to indicate an intraparasitic staining including a specific, small structure that may well be the apicoplast [23]. In blood-stage parasites, the single critical role of the apicoplast was shown to be the production of isopentenyl pyrophosphate (IPP) [53] and it is tempting to speculate that DMT2 is the dedicated IPP transport protein. However, DMT2 may also be involved in critical processes for the maintenance of the organelle and the IPP biosynthesis pathway, e.g. through the import of sulphur or iron into the organelle for essential iron-sulphur cluster biosynthesis [54].
–
Conclusions
In conclusion, our extensive orthology assignment and phylogenetic profiling, in combination with published experimental genetics studies, particularly in the murine malaria model parasite P. berghei, support the candidacy of a number of prophylactic, therapeutic, and transmission-blocking drug targets. Further studies into the biochemical and structural properties of these MTPs are required and deserve prioritization.
MATERIALS AND METHODS
Identification of putative orthologues
We first selected candidate protein sequences by homology search using Blast (blastp, version 2.2.29+) [55], against a set of pre-selected proteomes (Table S2) applying a threshold for being considered significant of E=5e-4. Next, for each candidate, we performed a reciprocal Blast search against the Plasmodium falciparum genome. To circumvent incomplete annotations of the genomes, we have additionally searched each query against the NCBI non-redundant (NR) database from March 19th, 2016. Hits on the same sequence were analysed and the overall coverage of the identified homology was calculated as the fraction of the query length. Candidates which returned the same protein as the original query (i.e., reciprocal blast hits) were kept as putative orthologues, while remaining hits were retained as unspecified homologues. We further inspected the putative homologues using the RADS algorithm [56] and the Needleman-Wunsch global alignment algorithm.
–
Phylogenetic reconstruction
Groups of putative orthologues for selected proteins were aligned using the programme Clustal Omega v. 1.2.1 [57] with default parameters. The alignments were then manually inspected and regions of low coverage and poor conservation were masked. Maximum likelihood phylogeny reconstruction was performed with the TreePuzzle [58] with 10,000 iterations and a mixed model of rate homogeneity.
–
Domain analysis
Domains of protein sequences were detected with hmmscan from the HMMer package v. 3.1b1 (http://hmmer.org) and the HMM profile collection from the PfamA database (October 2015). Significant overlaps were solved by E-value precedence. Domain architectures were analysed in combination with phylogenetic trees using the DoMosaic programme [59].
References
- S.E. Lindner, J.L. Miller, and S.H.I. Kappe, "Malaria parasite pre-erythrocytic infection: preparation meets opportunity", Cellular Microbiology, vol. 14, pp. 316-324, 2012. http://dx.doi.org/10.1111/j.1462-5822.2011.01734.x
- L.H. Miller, H.C. Ackerman, X. Su, and T.E. Wellems, "Malaria biology and disease pathogenesis: insights for new treatments", Nature Medicine, vol. 19, pp. 156-167, 2013. http://dx.doi.org/10.1038/nm.3073
- G.A. Josling, and M. Llinás, "Sexual development in Plasmodium parasites: knowing when it's time to commit", Nature Reviews Microbiology, vol. 13, pp. 573-587, 2015. http://dx.doi.org/10.1038/nrmicro3519
- T.W. Kooij, and K. Matuschewski, "Triggers and tricks of Plasmodium sexual development", Current Opinion in Microbiology, vol. 10, pp. 547-553, 2007. http://dx.doi.org/10.1016/j.mib.2007.09.015
- . WHO, "World malaria report 2015", Available: http://www.who.int/malaria/publications/world-malaria-report-2015/report/en/ Accessed 27.08.2016, 2015.
- P. Schlagenhauf, and E. Petersen, "Malaria Chemoprophylaxis: Strategies for Risk Groups", Clinical Microbiology Reviews, vol. 21, pp. 466-472, 2008. http://dx.doi.org/10.1128/CMR.00059-07
- D. Radeva-Petrova, K. Kayentao, F.O. ter Kuile, D. Sinclair, and P. Garner, "Drugs for preventing malaria in pregnant women in endemic areas: any drug regimen versus placebo or no treatment", Cochrane Database of Systematic Reviews, vol. 2014, 2014. http://dx.doi.org/10.1002/14651858.CD000169.pub3
- T.N.C. Wells, R.H. van Huijsduijnen, and W.C. Van Voorhis, "Malaria medicines: a glass half full?", Nature Reviews Drug Discovery, vol. 14, pp. 424-442, 2015. http://dx.doi.org/10.1038/nrd4573
- . Medicines for Malaria Venture, "MMV annual report 2015", Available: http://www.mmv.org/newsroom/publications/annual-report-2015 Accessed 27.08.2016, 2015.
- A. Ecker, A.M. Lehane, J. Clain, and D.A. Fidock, "PfCRT and its role in antimalarial drug resistance", Trends in Parasitology, vol. 28, pp. 504-514, 2012. http://dx.doi.org/10.1016/j.pt.2012.08.002
- J.B. Koenderink, R.A. Kavishe, S.R. Rijpma, and F.G. Russel, "The ABCs of multidrug resistance in malaria", Trends in Parasitology, vol. 26, pp. 440-446, 2010. http://dx.doi.org/10.1016/j.pt.2010.05.002
- K. Kirk, and A.M. Lehane, "Membrane transport in the malaria parasite and its host erythrocyte", Biochemical Journal, vol. 457, pp. 1-18, 2013. http://dx.doi.org/10.1042/BJ20131007
- K. Kirk, "Channels and transporters as drug targets in the Plasmodium-infected erythrocyte", Acta Tropica, vol. 89, pp. 285-298, 2004. http://dx.doi.org/10.1016/j.actatropica.2003.10.002
- M. Rottmann, C. McNamara, B.K.S. Yeung, M.C.S. Lee, B. Zou, B. Russell, P. Seitz, D.M. Plouffe, N.V. Dharia, J. Tan, S.B. Cohen, K.R. Spencer, G.E. González-Páez, S.B. Lakshminarayana, A. Goh, R. Suwanarusk, T. Jegla, E.K. Schmitt, H. Beck, R. Brun, F. Nosten, L. Renia, V. Dartois, T.H. Keller, D.A. Fidock, E.A. Winzeler, and T.T. Diagana, "Spiroindolones, a Potent Compound Class for the Treatment of Malaria", Science, vol. 329, pp. 1175-1180, 2010. http://dx.doi.org/10.1126/science.1193225
- B.K.S. Yeung, B. Zou, M. Rottmann, S.B. Lakshminarayana, S.H. Ang, S.Y. Leong, J. Tan, J. Wong, S. Keller-Maerki, C. Fischli, A. Goh, E.K. Schmitt, P. Krastel, E. Francotte, K. Kuhen, D. Plouffe, K. Henson, T. Wagner, E.A. Winzeler, F. Petersen, R. Brun, V. Dartois, T.T. Diagana, and T.H. Keller, "Spirotetrahydro β-Carbolines (Spiroindolones): A New Class of Potent and Orally Efficacious Compounds for the Treatment of Malaria", Journal of Medicinal Chemistry, vol. 53, pp. 5155-5164, 2010. http://dx.doi.org/10.1021/jm100410f
- N. Spillman, R. Allen, C. McNamara, B. Yeung, E. Winzeler, T. Diagana, and K. Kirk, "Na+ Regulation in the Malaria Parasite Plasmodium falciparum Involves the Cation ATPase PfATP4 and Is a Target of the Spiroindolone Antimalarials", Cell Host & Microbe, vol. 13, pp. 227-237, 2013. http://dx.doi.org/10.1016/j.chom.2012.12.006
- M.B. Jiménez-Díaz, D. Ebert, Y. Salinas, A. Pradhan, A.M. Lehane, M. Myrand-Lapierre, K.G. O’Loughlin, D.M. Shackleford, M. Justino de Almeida, A.K. Carrillo, J.A. Clark, A.S.M. Dennis, J. Diep, X. Deng, S. Duffy, A.N. Endsley, G. Fedewa, W.A. Guiguemde, M.G. Gómez, G. Holbrook, J. Horst, C.C. Kim, J. Liu, M.C.S. Lee, A. Matheny, M.S. Martínez, G. Miller, A. Rodríguez-Alejandre, L. Sanz, M. Sigal, N.J. Spillman, P.D. Stein, Z. Wang, F. Zhu, D. Waterson, S. Knapp, A. Shelat, V.M. Avery, D.A. Fidock, F. Gamo, S.A. Charman, J.C. Mirsalis, H. Ma, S. Ferrer, K. Kirk, I. Angulo-Barturen, D.E. Kyle, J.L. DeRisi, D.M. Floyd, and R.K. Guy, "(+)-SJ733, a clinical candidate for malaria that acts through ATP4 to induce rapid host-mediated clearance of Plasmodium", Proceedings of the National Academy of Sciences, vol. 111, 2014. http://dx.doi.org/10.1073/pnas.1414221111
- K. Kirk, "Ion Regulation in the Malaria Parasite", Annual Review of Microbiology, vol. 69, pp. 341-359, 2015. http://dx.doi.org/10.1146/annurev-micro-091014-104506
- M.H. Saier, V.S. Reddy, D.G. Tamang, and �. Västermark, "The Transporter Classification Database", Nucleic Acids Research, vol. 42, pp. D251-D258, 2013. http://dx.doi.org/10.1093/nar/gkt1097
- J.M. Matz, and T.W.A. Kooij, "Towards genome-wide experimental genetics in thein vivomalaria model parasitePlasmodium berghei", Pathogens and Global Health, vol. 109, pp. 46-60, 2015. http://dx.doi.org/10.1179/2047773215Y.0000000006
- T.F. de Koning-Ward, P.R. Gilson, and B.S. Crabb, "Advances in molecular genetic systems in malaria", Nature Reviews Microbiology, vol. 13, pp. 373-387, 2015. http://dx.doi.org/10.1038/nrmicro3450
- S.R. Rijpma, M. van der Velden, T. Annoura, J.M. Matz, S. Kenthirapalan, T.W.A. Kooij, K. Matuschewski, G. van Gemert, M. van de Vegte‐Bolmer, R. Siebelink‐Stoter, W. Graumans, J. Ramesar, O. Klop, F.G.M. Russel, R.W. Sauerwein, C.J. Janse, B.M. Franke‐Fayard, and J.B. Koenderink, "Vital and dispensable roles of Plasmodium multidrug resistance transporters during blood‐ and mosquito‐stage development", Molecular Microbiology, vol. 101, pp. 78-91, 2016. http://dx.doi.org/10.1111/mmi.13373
- S. Kenthirapalan, A.P. Waters, K. Matuschewski, and T.W.A. Kooij, "Functional profiles of orphan membrane transporters in the life cycle of the malaria parasite", Nature Communications, vol. 7, 2016. http://dx.doi.org/10.1038/ncomms10519
- R.E. Martin, H. Ginsburg, and K. Kirk, "Membrane transport proteins of the malaria parasite", Molecular Microbiology, vol. 74, pp. 519-528, 2009. http://dx.doi.org/10.1111/j.1365-2958.2009.06863.x
- A.B. Vaidya, and M.W. Mather, "Mitochondrial Evolution and Functions in Malaria Parasites", Annual Review of Microbiology, vol. 63, pp. 249-267, 2009. http://dx.doi.org/10.1146/annurev.micro.091208.073424
- G.G. van Dooren, and B. Striepen, "The Algal Past and Parasite Present of the Apicoplast", Annual Review of Microbiology, vol. 67, pp. 271-289, 2013. http://dx.doi.org/10.1146/annurev-micro-092412-155741
- J. Archibald, "Endosymbiosis and Eukaryotic Cell Evolution", Current Biology, vol. 25, pp. R911-R921, 2015. http://dx.doi.org/10.1016/j.cub.2015.07.055
- V. Zimorski, C. Ku, W.F. Martin, and S.B. Gould, "Endosymbiotic theory for organelle origins", Current Opinion in Microbiology, vol. 22, pp. 38-48, 2014. http://dx.doi.org/10.1016/j.mib.2014.09.008
- P.J. Keeling, "The Number, Speed, and Impact of Plastid Endosymbioses in Eukaryotic Evolution", Annual Review of Plant Biology, vol. 64, pp. 583-607, 2013. http://dx.doi.org/10.1146/annurev-arplant-050312-120144
- R.H. Jiang, R.V. Stahelin, S. Bhattacharjee, and K. Haldar, "Eukaryotic virulence determinants utilize phosphoinositides at the ER and host cell surface", Trends in Microbiology, vol. 21, pp. 145-156, 2013. http://dx.doi.org/10.1016/j.tim.2012.12.004
- S.R. Rijpma, M. Velden, M. González‐Pons, T. Annoura, B.C.L. Schaijk, G. Gemert, J.J.M.W. Heuvel, J. Ramesar, S. Chevalley‐Maurel, I.H. Ploemen, S.M. Khan, J. Franetich, D. Mazier, J.H. Wilt, A.E. Serrano, F.G.M. Russel, C.J. Janse, R.W. Sauerwein, J.B. Koenderink, and B.M. Franke‐Fayard, "Multidrug ATP‐binding cassette transporters are essential for hepatic development of Plasmodium sporozoites", Cellular Microbiology, vol. 18, pp. 369-383, 2015. http://dx.doi.org/10.1111/cmi.12517
- S. Portugal, C. Carret, M. Recker, A.E. Armitage, L.A. Gonçalves, S. Epiphanio, D. Sullivan, C. Roy, C.I. Newbold, H. Drakesmith, and M.M. Mota, "Host-mediated regulation of superinfection in malaria", Nature Medicine, vol. 17, pp. 732-737, 2011. http://dx.doi.org/10.1038/nm.2368
- P. Ferrer, R. Castillo-Neyra, C.N. Roy, and D.J. Sullivan, "Dynamic control of hepatic Plasmodium numbers by hepcidin despite elevated liver iron during iron supplementation", Microbes and Infection, vol. 18, pp. 48-56, 2016. http://dx.doi.org/10.1016/j.micinf.2015.08.018
- K. Slavic, S. Krishna, A. Lahree, G. Bouyer, K.K. Hanson, I. Vera, J.K. Pittman, H.M. Staines, and M.M. Mota, "A vacuolar iron-transporter homologue acts as a detoxifier in Plasmodium", Nature Communications, vol. 7, 2016. http://dx.doi.org/10.1038/ncomms10403
-
T. Sahu, B. Boisson, C. Lacroix, E. Bischoff, Q. Richier, P. Formaglio, S. Thiberge, I. Dobrescu, R. Ménard, and P. Baldacci, "
ZIPCO , a putative metal ion transporter, is crucial for Plasmodium liver‐stage development", EMBO Molecular Medicine, vol. 6, pp. 1387-1397, 2014. http://dx.doi.org/10.15252/emmm.201403868 -
S. Kenthirapalan, A.P. Waters, K. Matuschewski, and T.W.A. Kooij, "Copper‐transporting
ATP ase is important for malaria parasite fertility", Molecular Microbiology, vol. 91, pp. 315-325, 2013. http://dx.doi.org/10.1111/mmi.12461 - T. Kambe, A. Hashimoto, and S. Fujimoto, "Current understanding of ZIP and ZnT zinc transporters in human health and diseases", Cellular and Molecular Life Sciences, vol. 71, pp. 3281-3295, 2014. http://dx.doi.org/10.1007/s00018-014-1617-0
- D.S. Guttery, J.K. Pittman, K. Frénal, B. Poulin, L.R. McFarlane, K. Slavic, S.P. Wheatley, D. Soldati-Favre, S. Krishna, R. Tewari, and H.M. Staines, "The Plasmodium berghei Ca2+/H+ Exchanger, PbCAX, Is Essential for Tolerance to Environmental Ca2+ during Sexual Development", PLoS Pathogens, vol. 9, pp. e1003191, 2013. http://dx.doi.org/10.1371/journal.ppat.1003191
- R.J. Hart, L. Lawres, E. Fritzen, C.B. Mamoun, and A.S.I. Aly, "Plasmodium yoelii Vitamin B5 Pantothenate Transporter Candidate is Essential for Parasite Transmission to the Mosquito", Scientific Reports, vol. 4, 2014. http://dx.doi.org/10.1038/srep05665
- K.L. Waller, S.M. McBride, K. Kim, and T.V. McDonald, "Characterization of two putative potassium channels in Plasmodium falciparum", Malaria Journal, vol. 7, 2008. http://dx.doi.org/10.1186/1475-2875-7-19
- P. Ellekvist, J. Maciel, G. Mlambo, C.H. Ricke, H. Colding, D.A. Klaerke, and N. Kumar, "Critical role of a K + channel in Plasmodium berghei transmission revealed by targeted gene disruption", Proceedings of the National Academy of Sciences, vol. 105, pp. 6398-6402, 2008. http://dx.doi.org/10.1073/pnas.0802384105
- D. Promeneur, Y. Liu, J. Maciel, P. Agre, L.S. King, and N. Kumar, "Aquaglyceroporin PbAQP during intraerythrocytic development of the malaria parasite Plasmodium berghei", Proceedings of the National Academy of Sciences, vol. 104, pp. 2211-2216, 2007. http://dx.doi.org/10.1073/pnas.0610843104
- S. Kenthirapalan, A.P. Waters, K. Matuschewski, and T.W. Kooij, "Flow cytometry-assisted rapid isolation of recombinant Plasmodium berghei parasites exemplified by functional analysis of aquaglyceroporin", International Journal for Parasitology, vol. 42, pp. 1185-1192, 2012. http://dx.doi.org/10.1016/j.ijpara.2012.10.006
- J.M. Matz, K. Matuschewski, and T.W. Kooij, "Two putative protein export regulators promote Plasmodium blood stage development in vivo", Molecular and Biochemical Parasitology, vol. 191, pp. 44-52, 2013. http://dx.doi.org/10.1016/j.molbiopara.2013.09.003
- S. Pulcini, H.M. Staines, J.K. Pittman, K. Slavic, C. Doerig, J. Halbert, R. Tewari, F. Shah, M.A. Avery, R.K. Haynes, and S. Krishna, "Expression in Yeast Links Field Polymorphisms in PfATP6 to in Vitro Artemisinin Resistance and Identifies New Inhibitor Classes", The Journal of Infectious Diseases, vol. 208, pp. 468-478, 2013. http://dx.doi.org/10.1093/infdis/jit171
- J. Fonager, E.M. Pasini, J.A. Braks, O. Klop, J. Ramesar, E.J. Remarque, I.O. Vroegrijk, S.G. van Duinen, A.W. Thomas, S.M. Khan, M. Mann, C.H. Kocken, C.J. Janse, and B.M. Franke-Fayard, "Reduced CD36-dependent tissue sequestration of Plasmodium-infected erythrocytes is detrimental to malaria parasite growth in vivo", Journal of Experimental Medicine, vol. 209, pp. 93-107, 2011. http://dx.doi.org/10.1084/jem.20110762
- J. Xiong, J. Feng, D. Yuan, J. Zhou, and W. Miao, "Tracing the structural evolution of eukaryotic ATP binding cassette transporter superfamily", Scientific Reports, vol. 5, 2015. http://dx.doi.org/10.1038/srep16724
- D.J. Carucci, A.A. Witney, D.K. Muhia, D.C. Warhurst, P. Schaap, M. Meima, J. Li, M.C. Taylor, J.M. Kelly, and D.A. Baker, "Guanylyl Cyclase Activity Associated with Putative Bifunctional Integral Membrane Proteins in Plasmodium falciparum", Journal of Biological Chemistry, vol. 275, pp. 22147-22156, 2000. http://dx.doi.org/10.1074/jbc.M001021200
- R.W. Moon, C.J. Taylor, C. Bex, R. Schepers, D. Goulding, C.J. Janse, A.P. Waters, D.A. Baker, and O. Billker, "A Cyclic GMP Signalling Module That Regulates Gliding Motility in a Malaria Parasite", PLoS Pathogens, vol. 5, pp. e1000599, 2009. http://dx.doi.org/10.1371/journal.ppat.1000599
- M. Hirai, M. Arai, S. Kawai, and H. Matsuoka, "PbGCβ Is Essential for Plasmodium Ookinete Motility to Invade Midgut Cell and for Successful Completion of Parasite Life Cycle in Mosquitoes", The Journal of Biochemistry, vol. 140, pp. 747-757, 2006. http://dx.doi.org/10.1093/jb/mvj205
- A. Ecker, V. Lakshmanan, P. Sinnis, I. Coppens, and D.A. Fidock, "Evidence That Mutant PfCRT Facilitates the Transmission to Mosquitoes of Chloroquine-Treated Plasmodium Gametocytes", The Journal of Infectious Diseases, vol. 203, pp. 228-236, 2011. http://dx.doi.org/10.1093/infdis/jiq036
- K. Slavic, U. Straschil, L. Reininger, C. Doerig, C. Morin, R. Tewari, and S. Krishna, "Life cycle studies of the hexose transporter of Plasmodium species and genetic validation of their essentiality", Molecular Microbiology, vol. 75, pp. 1402-1413, 2010. http://dx.doi.org/10.1111/j.1365-2958.2010.07060.x
- E. Yeh, and J.L. DeRisi, "Chemical Rescue of Malaria Parasites Lacking an Apicoplast Defines Organelle Function in Blood-Stage Plasmodium falciparum", PLoS Biology, vol. 9, pp. e1001138, 2011. http://dx.doi.org/10.1371/journal.pbio.1001138
- J.M. Haussig, K. Matuschewski, and T.W.A. Kooij, "Identification of Vital and Dispensable Sulfur Utilization Factors in the Plasmodium Apicoplast", PLoS ONE, vol. 9, pp. e89718, 2014. http://dx.doi.org/10.1371/journal.pone.0089718
- C. Camacho, G. Coulouris, V. Avagyan, N. Ma, J. Papadopoulos, K. Bealer, and T.L. Madden, "BLAST+: architecture and applications", BMC Bioinformatics, vol. 10, 2009. http://dx.doi.org/10.1186/1471-2105-10-421
- N. Terrapon, J. Weiner, S. Grath, A.D. Moore, and E. Bornberg-Bauer, "Rapid similarity search of proteins using alignments of domain arrangements", Bioinformatics, vol. 30, pp. 274-281, 2013. http://dx.doi.org/10.1093/bioinformatics/btt379
- F. Sievers, A. Wilm, D. Dineen, T.J. Gibson, K. Karplus, W. Li, R. Lopez, H. McWilliam, M. Remmert, J. Söding, J.D. Thompson, and D.G. Higgins, "Fast, scalable generation of high‐quality protein multiple sequence alignments using Clustal Omega", Molecular Systems Biology, vol. 7, 2011. http://dx.doi.org/10.1038/msb.2011.75
- H.A. Schmidt, K. Strimmer, M. Vingron, and A. von Haeseler, "TREE-PUZZLE: maximum likelihood phylogenetic analysis using quartets and parallel computing", Bioinformatics, vol. 18, pp. 502-504, 2002. http://dx.doi.org/10.1093/bioinformatics/18.3.502
- A.D. Moore, A. Held, N. Terrapon, J. Weiner, and E. Bornberg-Bauer, "DoMosaics: software for domain arrangement visualization and domain-centric analysis of proteins", Bioinformatics, vol. 30, pp. 282-283, 2013. http://dx.doi.org/10.1093/bioinformatics/btt640
SUPPLEMENTAL INFORMATION
![]() Download Supplemental Information
Download Supplemental Information
ACKNOWLEDGMENTS
This work was supported by the Max Planck Society and by the Netherlands Organisation for Scientific Research (NWO-VIDI 864.13.009). We would like to thank Gayle McEwen for her help with the manuscript.
COPYRIGHT
© 2016

Phylogenetic profiles of all membrane transport proteins of the malaria parasite highlight new drug targets by January Weiner 3rd and Taco W.A. Kooij is licensed under a Creative Commons Attribution 4.0 International License.