News and thoughts:
Microbial Cell, Vol. 6, No. 12, pp. 527 - 530; doi: 10.15698/mic2019.12.699
Sulfur dioxide resistance in Saccharomyces cerevisiae: beyond SSU1
1 Food Biotechnology Department, Instituto de Agroquímica y Tecnología de Alimentos (IATA), Consejo Superior de Investigaciones Científicas (CSIC), Valencia, Spain.
Keywords: Saccharomyces cerevisiae, wine, sulfur dioxide, Com2.
Received originally: 12/11/2019 Accepted: 15/11/2019
Published: 21/11/2019
Correspondence:
José Manuel Guillamón, Food Biotechnology Department, Instituto de Agroquímica y Tecnología de Alimentos (IATA), Consejo Superior de Investigaciones Científicas (CSIC), Valencia, Spain; guillamon@iata.csic.es
Estéfani García-Ríos, Food Biotechnology Department, Instituto de Agroquímica y Tecnología de Alimentos (IATA), Consejo Superior de Investigaciones Científicas (CSIC), Valencia, Spain; e.garcia.rios@iata.csic.es
Conflict of interest statement: The authors declare no conflict of interest.
Please cite this article as: Estéfani García-Ríos and José Manuel Guillamón (2019). Sulfur dioxide resistance in Saccharomyces cerevisiae: beyond SSU1. Microbial Cell 6(12): 527-530. doi: 10.15698/mic2019.12.699
Sulfite resistance is an important oenological trait for wine yeasts because this compound is used during winemaking as a microbial inhibitor and antioxidant. The molecular mechanisms by which Saccharomyces cerevisiae responds and tolerates SO2 have been mainly focused on the sulfite efflux pump encoded by SSU1. Different chromosomal rearrangements in the regulatory region of this gene have been correlated with improved sulfite tolerance. However, other molecular factors must contribute to this trait because the SSU1 gene activity does not always fit with sulfite tolerance. An interesting approach to shed light onto this issue could be found by Lage et al. (2019). These authors have combined transcriptomic and genome-wide analysis to describe how the poorly characterized transcription factor Com2 controls, directly or indirectly, the expression of more than 80% of the genes activated by SO2. Additionally, large-scale phenotyping revealed the identification of 50 Com2-targets contributing to the protection against SO2. This information is very interesting for gaining knowledge regarding this important industrial trait.
Microorganisms utilize a great variety of genetic strategies to adapt to natural and human-made environments. The wine strains of S. cerevisiae are a clear example of highly specialized microorganisms that have evolved to use the different ecological niches provided by human activity. The specific genetic characteristics of the wine yeast strains are a consequence of the process of domestication [1][2][3][4][5][6]. Thus, in the age of high throughput sequencing technologies and omics data, a current challenge is to unveil the molecular determinants underlying a specific trait of industrial interest. This knowledge will be of paramount importance for selection and genetic improvement of the industrial strains.
–
Sulfite (SO32−), which is produced by dissolution of sulfur dioxide (SO2) in water, is used during winemaking as a microbial inhibitor and antioxidant. Therefore, sulfite resistance is a desired trait for wine yeast strains [7]. S. cerevisiae cells have different mechanisms to deal with the stress produced by sulfites including the increase in the production of acetaldehyde, which binds to SO32−, the regulation of the sulfite uptake pathway, and sulfite efflux through a plasma membrane pump encoded by the SSU1 gene [8]. Indeed, wine S. cerevisiae strains are considerably more tolerant to SO2 than laboratory strains and the main molecular mechanism connected with this higher resistant phenotype results from a higher transcription of the SSU1 gene. Among this group of strains, three chromosomal rearrangements (VIIItXVI, XVtXVI and Inv-XVI) have been described to up-regulate SSU1 expression and thereby increasing sulfite tolerance [9][10][11][12]. In all cases, the chromosomal rearrangement involves the SSU1 promoter and leads to its transcriptional up-regulation. Surprisingly, SSU1 transcript levels are not responsive to SO2 [9][10][12], indicating that the different levels of resistance in S. cerevisiae wine strains are, in general, explained by the basal transcript levels of SSU1, except for the 71B strain that harbors a sulfur-inducible SSU1 gene that may have gained a new regulatory system [13]. However, in some cases SSU1 mRNA levels did not correlate with sulfite tolerance probably due to the contribution of other factors to yeast sulfite resistance. Thus, a key question remains to be addressed in the field: Which are the molecular mechanisms that explain yeast sulfite tolerance? This question is of paramount importance for understanding much better the regulation network between gene activity and metabolic response. However, this information is also crucial for the wine industry, which is leading a process of sulfite reduction in their wines
–
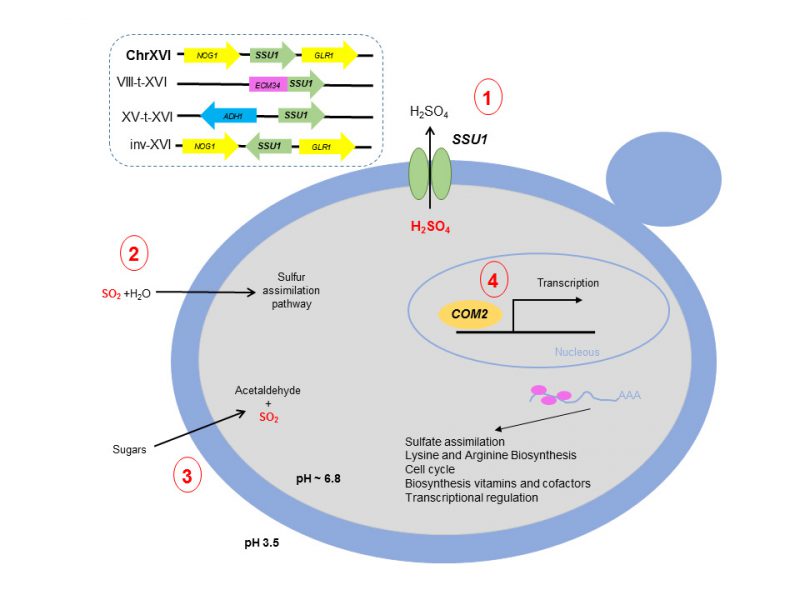
In this scenario, Lage et al., (2019) [14] have recently demonstrated the involvement of Com2 as the main transcriptional factor regulating genes that impart SO2 resistance. The results clearly highlight Com2 as the main player in the reprogramming of yeast genomic expression under SO2 stress, regulating approximately 80% of the SO2-activated genes. Moreover, the involvement in SO2 tolerance of genes regulated by Com2 was validated by a large-scale phenotyping of the corresponding mutants, proving the susceptibility of these mutants to SO2. Figure 1 shows a summary of the main mechanisms involved in SO2 resistance in S. cerevisiae.
–
–
COM2 encodes an orphan homologue of the environmental stress-responsive transcription factors Msn2 and Msn4 [15] and, the genes regulated by Msn2 were also found to be regulated by Com2, suggesting some overlap between them. In a phenotypic assay with and without sulfite, the strain lacking the Com2 gene was more sensitive to sulfite than its wild-type counterpart (BY4741). The transcriptomic profiling of BY4741 and BY4741_com2? cells revealed dramatic changes in both strains especially after exposure to SO2 (0.5 mM), although this effect was markedly different in the two strains. A higher number of genes were differentially expressed in the wild type compared to the com2? strain, indicating that these SO2-responsive genes present in the wild type and without activation in the mutant strain could be considered as Com2 targets. No significant effect of Com2 was found in the absence of SO2. The functional categories obtained after analyzing these genes are old known from the transcriptomic analysis performed to date and are mainly related to nitrogen, sulfur and some aminoacids metabolism [16]. In this line, authors have shown that supplementation of the medium with arginine and lysine somehow alleviated SO2 toxicity. A plausible explanation could be that exposure to SO2 can lead to a depletion of intracellular lysine and arginine or, in the case of arginine, by contributing to the integrity of the cell wall and the plasma membrane [17].
–
However, as mentioned above, the exposition to SO2 during hundreds of years and thousands of generations have caused S. cerevisiae wine strains to be more tolerant to this compound than the laboratory strains. They have evolved to employ various anthropic niches or environments during the so-called “unaware domestication” process [1][3][18]. Their genomes present genetic polymorphisms with different evolutionary consequences all of which help wine yeast genomes to adapt [19][20]. Thus, the main limitation of this study is the use of a single laboratory strain genetic background to investigate a phenotype of industrial relevance in the wine industry. Thus, the remaining question is: How much is the contribution of the Com2 in a wine yeast background adapted to sulfite stress? The key role of this gene in the laboratory strains is undeniable but we have to be careful when transferring this information from one background to another and, therefore, more studies are needed to determine its possible application in the industry.
–
REFERENCES
- Legras J, Galeote V, Bigey F, Camarasa C, Marsit S, Nidelet T, Sanchez I, Couloux A, Guy J, Franco-duarte R, Marcet-houben M, Gabaldon T, Schuller D, and Sampaio JP (2018). Adaptation of S . cerevisiae to fermented food environments reveals remarkable genome plasticity and the footprints of domestication. Mol Biol Evol 35(May): 1712–1727. 10.1093/molbev/msy066
- Liti G, Carter DM, Moses AM, Warringer J, Parts L, James SA, Davey RP, Roberts IN, Burt A, Koufopanou V, Tsai IJ, Bergman CM, Bensasson D, O ‘kelly MJT, Van Oudenaarden A, Barton DBH, Bailes E, Nguyen Ba AN, Jones M, Quail MA, Goodhead I, Sims S, Smith F, Blomberg A, Durbin R, and Louis EJ (2009). Population genomics of domestic and wild yeasts. Nature 458: 337–341. 10.1038/nature07743
- Almeida P, Barbosa R, Zalar P, Imanishi Y, Shimizu K, Turchetti B, Legras JL, Serra M, Dequin S, Couloux A, Guy J, Bensasson D, Gonçalves P, and Sampaio JP (2015). A population genomics insight into the Mediterranean origins of wine yeast domestication. Mol Ecol 24(21): 5412–5427. 10.1111/mec.13341
- Gonçalves M, Pontes A, Almeida P, Barbosa R, Serra M, Libkind D, Hutzler M, Gonçalves P, and Sampaio JP (2016). Distinct Domestication Trajectories in Top-Fermenting Beer Yeasts and Wine Yeasts. Curr Biol 26(20): 2750–2761. 10.1016/j.cub.2016.08.040
- Borneman AR, Forgan AH, Kolouchova R, Fraser JA, and Schmidt SA (2016). Whole Genome Comparison Reveals High Levels of Inbreeding and Strain Redundancy Across the Spectrum of Commercial Wine Strains of Saccharomyces cerevisiae. G3 6(4): 957–971. 10.1534/g3.115.025692
- Gayevskiy V, Lee S, and Goddard MR (2016). European derived Saccharomyces cerevisiae colonisation of New Zealand vineyards aided by humans. FEMS Yeast Res 16(7): 1–12. 10.1093/femsyr/fow091
- Divol B, Du Toit M, and Duckitt E (2012). Surviving in the presence of sulphur dioxide: Strategies developed by wine yeasts. Appl Microbiol Biotechnol 95(3): 601–613. 10.1007/s00253-012-4186-x
- Casalone E, Colella CM, Daly S, Gallori E, Moriani L, and Polsinelli M (1992). Mechanism of resistance to sulphite in Saccharomyces cerevisiae. Curr Genet 22(6): 435–440. 10.1007/BF00326407
- Pérez-Ortín JE, Querol A, Puig S, and Barrio E (2002). Molecular characterization of a chromosomal rearrangement involved in the adaptive evolution of yeast strains. Genome Res 12(10): 1533–9. 10.1101/gr.436602
- Zimmer A, Durand C, Loira N, Durrens P, Sherman DJ, and Marullo P (2014). QTL dissection of lag phase in wine fermentation reveals a new translocation responsible for Saccharomyces cerevisiae adaptation to sulfite. PLoS One 9(1): e86298. 10.1371/journal.pone.0086298
- Yuasa N, Nakagawa Y, Hayakawa M, and Iimura Y (2004). Distribution of the sulfite resistance gene SSU1-R and the variation in its promoter region in wine yeasts. J Biosci Bioeng 98(5): 394–397. 10.1016/S1389-1723(04)00303-2
- García-Ríos E, Nuévalos M, Barrio E, Puig S, and Guillamón JM (2019). A new chromosomal rearrangement improves the adaptation of wine yeasts to sulfite. Environ Microbiol 21(5): 1771–1781. 10.1111/1462-2920.14586
- Nardi T, Corich V, Giacomini A, and Blondin B (2010). A sulphite-inducible form of the sulphite efflux gene SSU1 in a Saccharomyces cerevisiae wine yeast. Microbiology 156(6): 1686–1696. 10.1099/mic.0.036723-0
- Lage P, Sampaio-marques B, Ludovico P, Mira NP, and Mendes-ferreira A (2019). Transcriptomic and chemogenomic analyses unveil the essential role of Com2-regulon in response and tolerance of Saccharomyces cerevisiae to stress induced by sulfur dioxide. Microbial Cell 6(11): 509-523. 10.15698/mic2019.11.697
- Gasch AP, Spellman PT, Kao CM, Carmel-Harel O, Eisen MB, Storz G, Botstein D, and Brown PO (2000). Genomic expression programs in the response of yeast cells to environmental changes. Mol Biol Cell 11(12): 4241–4257. 10.1091/mbc.11.12.4241
- Nadai C, Treu L, Campanaro S, Giacomini A, and Corich V (2016). Different mechanisms of resistance modulate sulfite tolerance in wine yeasts. Appl Microbiol Biotechnol 100(2): 797–813. 10.1007/s00253-015-7169-x
- Cheng Y, Du Z, Zhu H, Guo X, and He X (2016). Protective Effects of Arginine on Saccharomyces cerevisiae Against Ethanol Stress. Sci Rep 6: 1–12. 10.1038/srep31311
- Peter J, De Chiara M, Friedrich A, Yue J-X, Pflieger D, Bergstrom A, Sigwalt A, Barré B, Freel K, Llored A, Cruaud C, Labadie K, Aury JM, Istace B, Lebrigand K, Barbry P, Engelen S, Lemainque A, Liti G, and Schacherer J (2018). Genome evolution across 1,011 Saccharomyces cerevisiae isolates. Nature 556: 339–344. 10.1038/s41586-018-0030-5
- Sicard D, and Legras JL (2011). Bread, beer and wine: yeast domestication in the Saccharomyces sensu stricto complex. C R Biol 334(3): 229–36. 10.1016/j.crvi.2010.12.016
- Bisson LF (2012). Geographic origin and diversity of wine strains of Saccharomyces. Am J Enol Vitic 63(2): 165–176. 10.5344/ajev.2012.11083
ACKNOWLEDGMENTS
This work was supported by the Span-ish Government through “Ministerio de Ciencia, Inno-vación y Universidades” (MICINN) and “Fondo Europeo de Desarrollo Regional” (FEDER) funds (grant number PCIN-2015-143, AGL2016-77503-C3-1-R) to JMG. This study has been carried out in the context of the Europe-an Project ERA-IB “YeastTempTation".
COPYRIGHT
© 2019

Sulfur dioxide resistance in Saccharomyces cerevisiae: beyond SSU1 by García-Ríos and Guillamón is licensed under a Creative Commons Attribution 4.0 International License.