Back to article: Regulation of anti-microbial autophagy by factors of the complement system
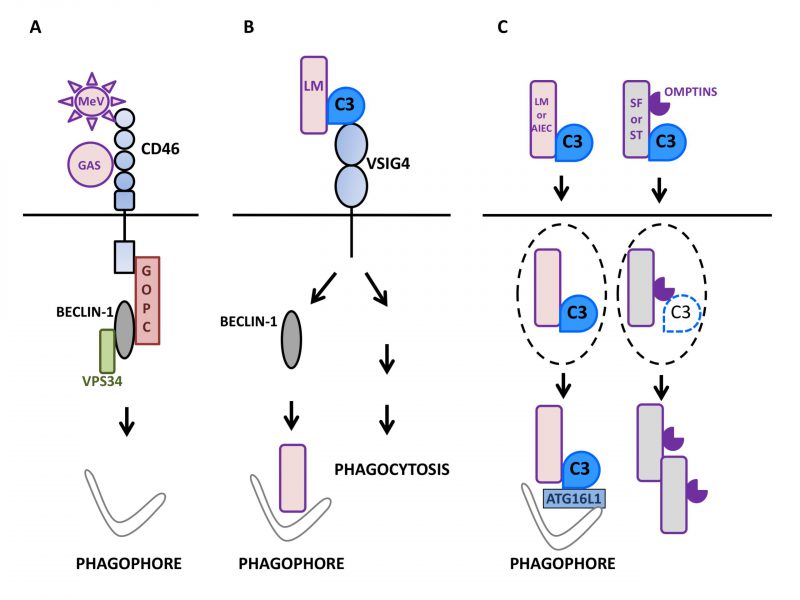
FIGURE 1: Autophagy induction by factors of the mammalian complement system in interaction with microbes.(A) Upon sensing of pathogens such as Measles virus (MeV) or group A Streptococcus (GAS), the regulator of complement activation CD46/MCP signals for autophagy induction in epithelial cells (depicted here by phagophore formation) by recruiting the BECLIN-1-VPS34 complex via the GOPC scaffold protein. (B) The macrophage surface receptor VSIG4/CRIg activates autophagy in a BECLIN-1-dependent manner in response to detection of C3 adsorbed onto L. monocytogenes (LM). Such an induction leads to the effective targeting of internalized bacteria to LC3-positive punctiform structures and then, lysosomal compartments. VSIG4/CRIg is also a positive regulator of phagocytosis in macrophages. (C) C3 deposited onto invading bacteria can induce autophagy by activating the autophagy machinery after reaching the cytosol of epithelial cells. Thus, after exiting their containing vacuole (dotted lines), C3-coated L. monocytogenes (LM) and adherent-invasive Escherichia Coli (AIEC) activate autophagy via the interaction of C3 with ATG16L1, one of the core autophagy factors. As a result, both bacteria are restricted by the induced autophagy. In contrast, other bacteria are able to escape such an antibacterial autophagy and multiply. For instance, C3-opsonized S. flexneri (SF) and S. Typhimurium (ST) are able to oppose autophagic restriction by lowering the level of the cytosolic C3-ATG16L1 interaction. This is due to their capacity to express proteolytic factors called Omptins that target C3 and reduce its amount deposited on the surface of bacterial cells.