Research Articles:
Microbial Cell, Vol. 10, No. 4, pp. 88 - 102; doi: 10.15698/mic2023.04.795
Acetate modulates the inhibitory effect of Lactobacillus gasseri against the pathogenic yeasts Candida albicans and Candida glabrata
1 iBB, Institute for Bioengineering and Biosciences, Instituto Superior Técnico – Department of Bioengineering, Universidade de Lisboa, 1049-001 Lisboa, Portugal.
2 Associate Laboratory i4HB—Institute for Health and Bioeconomy at Instituto Superior Técnico, Universidade de Lisboa, Av. Rovisco Pais, 1049-001 Lisboa, Portugal.
3 CQE-Centro Química Estrutural, Instituto Superior Técnico, Universidade de Lisboa, Av. Rovisco Pais, 1049-001 Lisboa, Portugal.
Keywords: lactobacilii-Candida interference, vaginal lactobacilii, vaginal candidiasis, probiotics, lactobacillus gasseri, candida phyiology.
Abbreviations:
NACS – non-albicans Candida species.
Received originally: 07/11/2022 Received in revised form: 10/03/2023
Accepted: 16/03/2023
Published: 21/03/2023
Correspondence:
Nuno P Mira, Instituto Superior Técnico, Department of Bioengineering, University of Lisbon, Portugal; nuno.mira@tecnico.ulisboa.pt
Conflict of interest statement: The authors declare no conflict of interest.
Please cite this article as: Nuno A. Pedro, Gabriela Fontebasso, Sandra N Pinto, Marta Alves and Nuno P Mira (2023). Acetate modulates the inhibi-tory effect of Lactobacillus gasseri against the pathogenic yeasts Candida albicans and Candida glabrata. Microbial Cell 10(4): 88-102. doi: 10.15698/mic2023.04.795
Abstract
The exploration of the interference prompted by commensal bacteria over fungal pathogens is an interesting alternative to develop new therapies. In this work we scrutinized how the presence of the poorly studied vaginal species Lactobacillus gasseri affects relevant pathophysiological traits of Candida albicans and Candida glabrata. L. gasseri was found to form mixed biofilms with C. albicans and C. glabrata resulting in pronounced death of the yeast cells, while bacterial viability was not affected. Reduced viability of the two yeasts was also observed upon co-cultivation with L. gasseri under planktonic conditions. Either in planktonic cultures or in biofilms, the anti-Candida effect of L. gasseri was augmented by acetate in a concentration-dependent manner. During planktonic co-cultivation the two Candida species counteracted the acidification prompted by L. gasseri thus impacting the balance between dissociated and undissociated organic acids. This feature couldn’t be phenocopied in single-cultures of L. gasseri resulting in a broth enriched in acetic acid, while in the co-culture the non-toxic acetate prevailed. Altogether the results herein described advance the design of new anti-Candida therapies based on probiotics, in particular, those based on vaginal lactobacilli species, helping to reduce the significant burden that infections caused by Candida have today in human health.
INTRODUCTION
Candidiasis are infections caused by pathogenic yeasts of the Candida genus and account around 50 to 70% of the reported fungal infections worldwide [1]. Infections caused by Candida are more frequent in the oral or in the vaginal mucosa, but in more serious cases, often life-threatening, these yeasts disseminate in the bloodstream and colonize major organs [2][3]. Due to their high mortality rates, aggressiveness and recurrence, infections caused by Candida have a very high societal impact [4]. The occurrence of systemic candidiasis is most frequently observed upon immunosuppression (e.g., in patients undergoing chemotherapy or in the elderly), but vaginal candidiasis is common among the healthy female population. Women are described to suffer two to three episodes of vaginal candidiasis during their life-time, with a significant proportion (5 to 10%) suffering from recurrent infections, a condition known as recurrent vulvovaginal candidiasis [5][6]. The more relevant species causative of candidiasis, both superficial and systemic, is Candida albicans but the incidence of infections caused by non-albicans Candida species (also called as NACS) is increasing, in some geographies prominently [7][8]. Infections caused by NACS are worrisome as these species are usually very resilient to commonly used antifungals and the underlying infections have poorer outcomes for patients, compared to infections caused by C. albicans [9][10]. Among NACS, Candida glabrata is usually the most prevalent species, in part due to its innately high tolerance to azoles and extreme genomic plasticity that, among other traits, prompts fast adaptive responses to the challenging environment of infection sites [9][11].
–
A distinguishing aspect of the pathophysiology of Candida, compared with other human-infecting fungi, is that these species are part of the microbiota of various niches, even in the absence of disease [12][13][14]. Indeed, C. albicans has been identified as a true gut symbiont based on its consistent identification in resident microbial populations of this niche [12][13]. C. glabrata has also been identified in the gut microbiome, however, not consistently, thus remaining to be elucidated whether it is a true symbiont or a transient passenger [15][16]. C. albicans and C. glabrata have also been identified in the vaginal mycobiome of “healthy” women, using both culture-dependent and independent methods [17][18][19]. However, these yeasts are not ubiquitously observed in all samples suggesting that the presumed “healthy” women could be asymptomatic carriers and leaving open whether or not the vaginal tract is a primary site of colonization of Candida. While in the past not much attention was given to these commensal populations of Candida, in the recent years it has been demonstrated that they can be reservoirs for dissemination, especially in the gastrointestinal (GI) tract [3][12].
–
Besides Candida a plethora of other species compose the vaginal microbiome, their identity and abundance differing with the anatomic site (for example, prominent differences were observed between the cervix or the uterus), with age, habits or race [20][21][22]. Despite the inter-individual variation, it is clear that the vaginal microbiome is dominated by lactobacilli with Lactobacillus iners, Lactobacillus crispatus, Lactobacillus jensenii and Lactobacillus gasseri being most abundant [23][24]. Perturbation of the vaginal microbiome was linked to adverse gynaecologic/obstetric outcomes including preterm birth [25], mucosal inflammation [26] or infections caused by HPV [27], HIV [28] or bacteria [22][29]. Studies examining the vaginal microbiome of asymptomatic women or of patients with diagnosed vaginal candidiasis obtained conflicting results: some reported decreased abundance of the lactobacilli population; others report only changes in the species profile (for example, unusual predominance of L. iners); and others don’t report any alterations to the habitual lactobacilli-enriched flora [17][19][30][31][32][33][34][35][36][37]. Although it is not totally clear whether vaginal lactobacilli provide protection against candidiasis in vivo, the potential of those species in inhibiting growth and relevant pathogenic traits of Candida in vitro has been well reported [38][39][40]. These observations opened the door to the use of probiotics based on lactobacilli as possible anti-Candida treatments with results pointing to a positive contribution in the prevention of relapse and avoidance of recolonization [41][42][43]. However, most of these probiotic cocktails were developed using lactobacilli species not indigenous to the vaginal tract, likely due to the poor knowledge available concerning their genetics and physiology [41][44]. Intestinal lactobacilli species (which differ from those found in the vaginal tract) have also been found to restrain growth and virulence of Candida in vitro and in infection models [45][46][47].
–
Resulting from this interest in the exploitation of the lactobacilli microflora as potential anti-Candida agents some studies examined this interference in more detail (as reviewed in [47][48][49]). Usually, the role of lactobacilli as drivers of vaginal health was attributed to lactic acid production which maintains an acidic vaginal pH and restrains bacterial growth [50][51][52]. However, the acidophilic nature of yeasts, including Candida spp., along with the demonstration that at pH of 4 (the usual pH of a vaginal fluid dominated by lactobacilli [53]) physiologically relevant concentrations of lactic acid don’t inhibit growth of C. albicans or C. glabrata [54], suggests that lactic acid may play a minor role in restraining Candida. Interestingly, it has been shown that lactate promotes the evasion of Candida cells from the immune system by inducing β-glucan masking and reducing macrophage recruitment [55], suggesting that these yeasts evolved adaptive responses to allow them to cope with the presence of this acid anion in the environment in a manner that favors colonization and, eventually, infection. Additionally, lactic acid was found to be a potent immunomodulator, reducing production of inflammatory cytokines by epithelial cells and thereby restraining the inflammatory response [56][57][58]. More recently, the anti-Candida potential of some lactobacilli species was attributed to the production of 1-acetyl-β-carboline [59], however, the only vaginal lactobacilli species studied was L. gasseri and the authors concluded that under their experimental setting supernatants obtained from culturing this bacterium inhibited filamentation of C. albicans, but not growth [59]. Other studies also demonstrated reduced filamentation and growth of C. albicans when cultivated in medium supplemented with supernatants obtained from L. gasseri, L. jensenii or L. crispatus cultures, although strain-to-strain variation was observed, especially for L. gasseri strains [40]. Other described anti-Candida effects attributed to lactobacilli involve the reduction in the ability of the yeasts to bind to epithelial cells due to a higher bacterial affinity for epithelial receptors, secretion of biosurfactants [60][61][62][63] or the production of bacteriocins [64][65]. However, no specific molecular players mediating these responses have been identified.
–
In this work we examined growth, physiology and virulence traits of C. albicans and C. glabrata when co-cultivated with the poorly studied vaginal species L. gasseri under planktonic or biofilm-forming conditions. This combined approach is a distinct aspect of our work since the majority of studies addressing this interaction used the cultivation of the yeasts in the presence of supernatants produced by vaginal bacterial cultures and not directly the cell-cell interaction. Besides assessing relevant physiological aspects of the interaction established between these two species, we have also uncovered the role of acetate as a positive modulator of the interference of L. gasseri over Candida even in concentrations similar to those found in a vaginal microflora dominated by lactobacilli.
RESULTS
Co-cultivation of C. glabrata and C. albicans with L. gasseri results in decreased viability of the yeasts, either under planktonic or biofilm-forming conditions
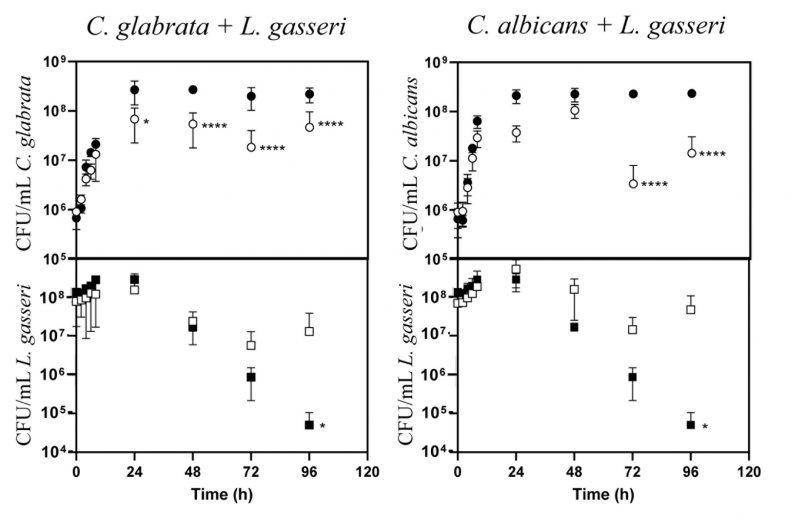
The majority of the studies that examined the interference established between the vaginal species L. gasseri and Candida focused on the inhibitory effect prompted by supernatants obtained from bacterial cultures [61][66][67], usually obtained in MRS, the canonical growth medium used for lactobacilli [68][69]. Considering that in vivo these two species are in close contact, we focused on their direct co-cultivation under planktonic and biofilm-forming conditions. To examine planktonic growth, the two Candida species (C. albicans and C. glabrata) and L. gasseri ATCC33323 were cultivated in liquid MRS for 96 h (Fig. 1). To simulate the higher abundance of lactobacilli in the vaginal microflora compared with the one of Candida [70][71][72][73], these co-cultivations were started using ∼100x more bacterial than yeast cells (∼108 CFUs/mL L. gasseri compared to 106 CFUs/mL of C. albicans or C. glabrata; Fig. 1). Under the experimental conditions used for the co-cultivation (100 rpm of agitation and 37°C) the yeast cells resumed growth immediately after re-inoculation and maintained it until 24 h after which they entered stationary phase (Fig. 1). The bacterial population also increased, although much less, likely due to a higher number of inoculated cells, compared with the one use for Candida (Fig. 1). Consistent with co-cultivation being a more competitive and challenging environment, the growth rate of C. glabrata and C. albicans in the presence of L. gasseri decreased 32% and 33% respectively, compared to the values observed in single culture (0.25 h-1 obtained in single culture of C. glabrata, compared to 0.17 h-1 in co-culture; 0.24 h-1 obtained in single culture of C. albicans, compared to 0.16 h-1 obtained in co-culture). Besides a reduction in the growth rate, it was also noticeable that co-cultivation with L. gasseri induced a prominent decrease (ranging between 53 and 98%) in the cellular viability of C. albicans and C. glabrata, this being considerably more prominent for the first species (Fig. 1). No significant reduction in viability was observed in the two single cultures of Candida indicating that the loss of viability observed in the co-culture setting is a direct effect of the presence of L. gasseri (Fig. 1). Differently, co-cultivation increased cellular viability of L. gasseri up to 107 CFUs/mL, compared to 105 CFUs/mL attained in single-culture (Fig. 1). In agreement with our results, reduced viability of L. gasseri cells upon short-medium term cultivation in MRS have been reported in other studies, presumably due to autolysis [74][75][76][77][78][79]. In order to exclude strain-dependent effects, we have repeated the co-cultivations using four vaginal C. glabrata and C. albicans strains and in both cases it was clear that co-cultivation with L. gasseri resulted in reduced viability of the yeast strains (see results in Supplementary Fig. S1). However, it was interesting to observe that the decrease in viability of the vaginal Candida strains imposed by co-cultivation with L. gasseri was smaller than the one obtained with the reference strains (compare results in Supplementary Fig. S1 and Fig. 1).
–
–
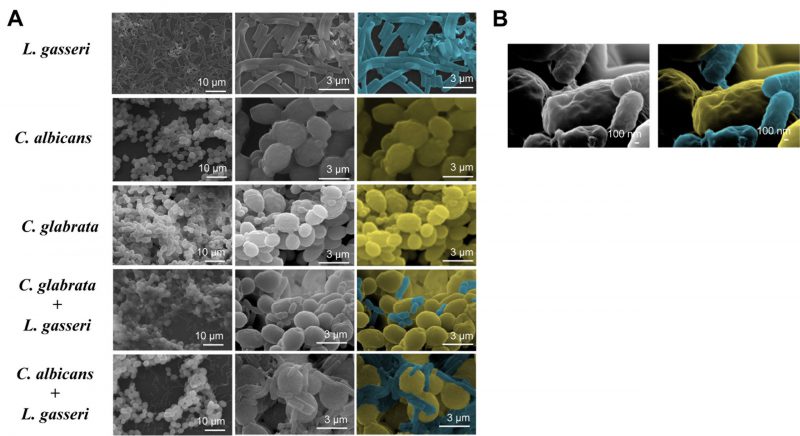
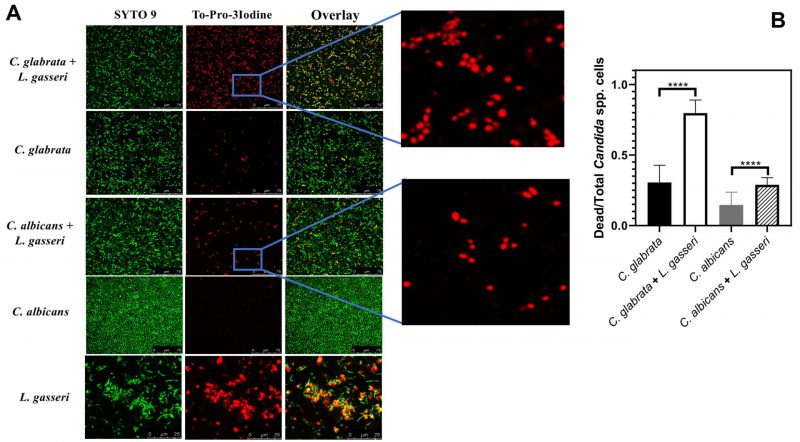
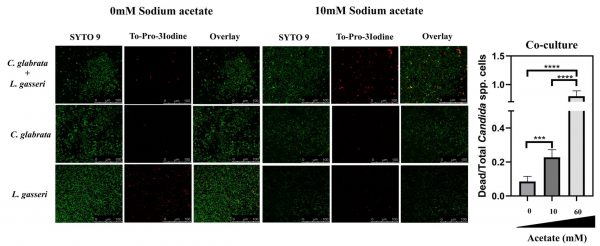
To test the effect of co-cultivation under sessile conditions the same experimental setting was used with the difference that C. albicans, C. glabrata and L. gasseri were cultivated in 8-well microplates. After 24 h of cultivation it was possible to observe that C. albicans and C. glabrata formed a mixed biofilm with L. gasseri involving very close cell-to-cell contacts, as shown by the microscopy SEM images depicted in Fig. 2. Single-species biofilms formed by C. albicans exhibited what appeared to be a multi-layered structure, while those formed by C. glabrata appeared more disperse (Fig. 3). These differences in structural organization of the biofilms formed by these two yeasts is consistent with results reported in other studies [80][81][82][83]. No significant differences were obtained concerning height of the single-species and multi-species biofilms, with the exception of the biofilms formed by L. gasseri alone that were considerably thinner (Supplementary Fig. S2). To understand what could be the outcome of the formation of these mixed biofilms in terms of viability, we took advantage of SYTO9 and TO-PRO3 iodide labelling that allowed us to differentiate, the yeast from the bacterial cells, while also distinguishing viable from non-viable cells directly in the biofilm [84]. In Fig. 3 we show the results of this labelling in single- and in multi-species biofilms. The results clearly demonstrate that the proportion of non-viable cells (labelled in red) was much higher in the mixed-biofilms than in the single-species ones, this effect being clearly more evident in the mixed biofilms formed between L. gasseri and C. glabrata (Fig. 3). Closer inspection of the images shows that these non-viable red-labelled cells correspond almost entirely to the yeasts while no significant loss of viability was observed for the L. gasseri cells (Fig. 3). To get a more quantitative view, we imaged in close detail a set of pictures (corresponding to more than 1000 yeast cells per condition) and found that the number of non-viable C. glabrata cells was about 2.6-fold higher in the mixed-biofilms than in the biofilms formed in the absence of the bacterium (Fig. 3). In the case of C. albicans the number of non-viable cells in the mixed-biofilm increased roughly 2-fold (Fig. 3). A striking labelling of TO-PRO-3 iodide was observed in the single-species L. gasseri biofilms visualizing at higher magnification that the labelling corresponds to what appears to be the extracellular matrix (Fig. 3 and a magnification shown in Supplementary Fig. S3). Upon entry into microbial cells, TO-PRO-3 iodide is described to bind nucleic acids [84][85] and thus it is possible that the observed labelling results from accumulated extracellular DNA (eDNA), as this was described to occur in biofilms formed by other lactobacilli species [86][87][88]. Notably, such labelling pattern was not detected in the single-species biofilms formed by the two Candida species, albeit eDNA has been reported in the extracellular matrix of C. albicans biofilms [89].
–
–
While in co-cultivation with L. gasseri, C. albicans and C. glabrata buffer the acidification prompted by the bacterium with consequences in the equilibrium of ionizable species like lactic and acetic acids
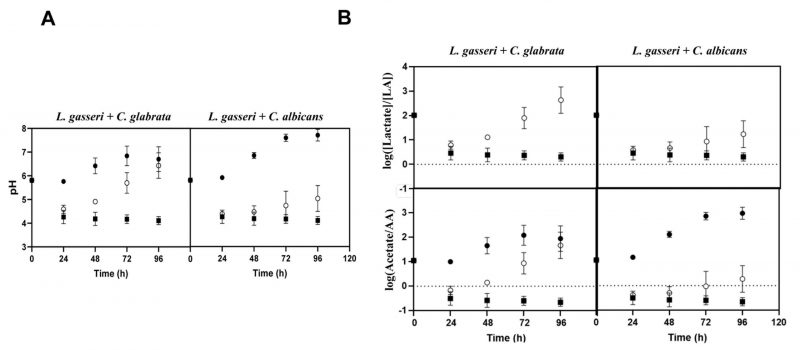
Taking into account that co-cultivation increased viability of L. gasseri, while it decreased viability of the two Candida species (Fig. 1), and that the high autolysis of L. gasseri in MRS was associated to the low pH of the medium (presumably due to the lactic acid formed) [77], we decided to monitor the pH achieved in single and in co-cultures. HPLC analyses of the broth confirmed the expected production of lactic acid by L. gasseri in amounts that ranged between 10 g/L in the single-culture to approximately 5 g/L in the co-culture (Supplementary Fig. S4). The lower amount of lactic acid produced in the co-cultures was consistent with a more rapid depletion of glucose in the fermentation medium, likely resulting from the yeast’s metabolic activity (Supplementary Fig. S4). As expected, no production of lactic acid was observed in the supernatants obtained from single cultures of the two Candida species, only an accumulation of ethanol was detectable (results not shown), which is compatible with the microaerophilic setting used. Concomitant with the production of lactic acid, a continuous acidification of the fermentation broth was observed, both in the single and in the co-cultivation settings (Fig. 4A). While in the single-cultures of L. gasseri the acidification persisted along the entire 96 h time frame and reaching a final pH of about 4, in the co-culture the pH started to increase after an initial drop (Fig. 4A). These observations suggest that the metabolic activity of the two Candida species buffers the acidification prompted by the accumulation of lactic acid, this buffering capacity being higher for C. glabrata than for C. albicans (final pH of the co-cultures was of 6.8 and 5, respectively; Fig. 4A). The same capacity of C. glabrata and C. albicans to alkalinize the broth when in co-cultivation with L. gasseri was observed for the tested vaginal strains (results not shown).
–
–
One of the factors imparted by the differences in pH of the broth registered during the single or co-cultivation of C. albicans/C. glabrata with L. gasseri is the distribution between the dissociated (RCOO–) and undissociated form (RCOOH) of lactic acid, as well as of other ionizable species accumulated in the broth, necessarily depending on their corresponding pKa values. Using the Handerson-Hasselbach equation to estimate the ratios between the amounts of lactate and lactic acid achieved during single or co-cultivation with Candida, it is clear that most of the lactic acid accumulated in the broth in the co-culture was dissociated (Fig. 4B). In the single-cultures of L. gasseri lactate also prevailed, however, the amount of undissociated lactic acid was considerably higher than the one present in the broth of co-cultures (Fig. 4B). Besides lactic acid, we also computed the ratio of dissociated and undissociated acetic acid since the MRS medium contains ∼60 mM of sodium acetate supplied as a sodium source (Fig. 4B). In this case, there was a marked difference between the result obtained in single-cultures of L. gasseri and in co-cultures, as undissociated acetic acid clearly predominated in the single-culture, while in the co-culture acetate prevailed (Fig. 4B). These observations show that the co-cultivation of C. glabrata or C. albicans with L. gasseri modulates important aspects of the composition of the supernatant including its pH and, consequently, the acid-base equilibrium of ionizable species like lactic and acetic acids.
–
–
Acetate augments anti-Candida activity prompted by L. gasseri cells
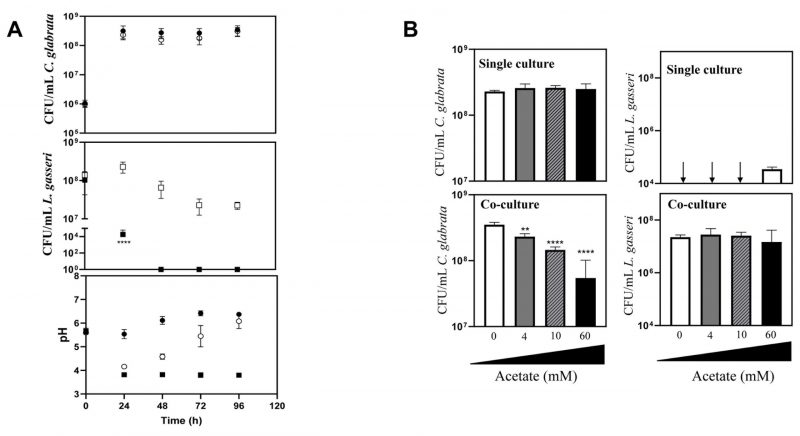
Our results shown in Fig. 4B demonstrate that in co-cultures of L. gasseri with C. albicans or C. glabrata the sodium acetate supplied in the MRS medium prevails in the acetate form. Considering that acetate has been demonstrated to induce the expression of bacteriocins in several Gram positive species [90][91], we hypothesized whether it could also modulate the anti-Candida effect exhibited by L. gasseri. For this we co-cultivated L. gasseri with C. glabrata in MRS medium under the same conditions used before but replacing sodium acetate by the same amount of sodium chloride. This replacement resulted in an accelerated loss of viability of L. gasseri cells when in single-culture, a phenotype that was rescued when the bacterial cells were cultivated in the presence of the yeasts (Fig. 5A). Remarkably, despite the maintenance in the viability of L. gasseri population, no significant loss of viability of C. glabrata could be detected in the co-cultures performed in MRS with NaCl (Fig. 5A). Notably, the capability of acetate to enhance the ability of L. gasseri cells to reduce viability of C. glabrata cells was concentration dependent and still detectable at concentrations as low as 4 mM (Fig. 5B, Supplementary Fig. S5). Consistent with the results observed for C. glabrata, co-cultivation of L. gasseri with C. albicans cells in MRS medium having NaCl (and not sodium acetate) as the sodium source also resulted in incapability of the bacterium to induce loss of viability in the yeast cell population (Supplementary Fig. S6). The replacement of sodium acetate by increasing amounts of sodium chloride led to a reduction in production of lactic acid (up to a maximum of 50%) prompted by the bacterium (Supplementary Fig. S4B and C). While in single-cultures, this decrease can be attributed to the lower viability of the bacterial cells when growing in MRS-NaCl, compared to MRS (Fig. 5A, Fig. 1), in the co-cultures performed in MRS-NaCl or in MRS the viability of L. gasseri cells was identical and therefore the lower production of lactic acid is likely to result from the lower availability of carbon (Fig. 5A, Fig. 1). The capability of acetate to increase the anti-Candida potential of L. gasseri was also detected when a vaginal clinical strain of the bacterium was used (L. gasseri ISTLg47) confirming that the effect is not exclusive for the used reference strain (Supplementary Fig. S7).
–
–
The effect of acetate in augmenting anti-Candida activity in L. gasseri cells is also observed under biofilm-forming conditions
The above reported effect of acetate in enhancing the capability of L. gasseri cells to induce loss of viability in C. albicans and in C. glabrata prompted us to examine whether the same effect could also be detected in biofilms. For that, we have used the same experimental setting as used above to detect the formation of mixed biofilms between L. gasseri and C. albicans or C. glabrata, including the live-dead confocal microscopy imaging. The modulation of acetate concentration in the MRS medium did not significantly alter the height of the biofilms formed (Supplementary Fig. S2). On the other hand, a marked increase in the viability of C. glabrata and C. albicans in mixed biofilms formed with L. gasseri during cultivation in MRS without acetate was observed, markedly contrasting with the high loss of viability that was observed in the normal composition of this medium including 60 mM acetate (Fig. 6 and Supplementary Fig. S8). Also as observed under planktonic conditions, this effect of acetate in augmenting the anti-Candida activity of L. gasseri cells was concentration-dependent (Fig. 6 and Supplementary Fig. S5). Strikingly, the unusual TO-PRO-3-labelling registered in the biofilms formed by L. gasseri cells were no longer detected when acetate was removed from the medium (results not shown).
–
DISCUSSION
In this work we focused on the interaction established between the poorly studied vaginal species L. gasseri and the pathogenic yeasts C. albicans and C. glabrata, both being frequent colonizers of the female vaginal tract. Other studies described the inhibitory potential of lactobacilli species, including L. gasseri, against Candida and other vaginal pathogens [39][67][92]. However, these studies don’t explore co-cultivation describing instead relevant alterations in pathophysiological traits of Candida when cultivated in media supplemented with various amounts of supernatants obtained from bacterial cultures [61][66][67][93]. The results shown herein demonstrate that the direct co-cultivation of L. gasseri with C. albicans and C. glabrata has outcomes that cannot be fully recapitulated by the bacterial supernatants themselves. For example, co-cultivation of C. albicans/C. glabrata with L. gasseri drastically altered the pH of the broth with consequences for the acid-base equilibrium of ionizable species like the organic acids acetic and lactic acid. In particular, we found that during co-cultivation, the two yeasts counteracted the prominent acidification of the medium promoted by the accumulation of lactic acid produced by L. gasseri cells. Consequently, the final pH of the co-cultures was ∼6 after 96 h, while in the single cultures of L. gasseri this pH was about ∼4.1. Prior studies have shown the capability of C. albicans to alkalinize the medium (via ammonium excretion) when using amino acids [94][95] or carboxylic acids as carbon sources [94][95]. C. glabrata has also been shown to alkalinize the medium while using amino acids as carbon sources but the identity of the buffering compound was not disclosed [96]. Notably, in these studies the alkalinization of the medium prompted by C. albicans or C. glabrata occurred under glucose limiting conditions, which is in line with the results we had obtained since the pH started to increase after 48 h, when no glucose was available in the broth (Supplementary Fig. S4). This capability of C. albicans and C. glabrata to induce alkalinization under acidic conditions has been linked with their increased ability to thrive in the highly acidic phagolysosome, favoring colonization and immune evasion [96][97]. In C. albicans the alkalinization has also been shown to auto-induce hyphal morphogenesis [95]. It is possible that in vivo, when present in the vaginal tract, Candida cells can also counteract the acidification prompted by lactobacilli. Among other outcomes (such as the promotion of yeast-hyphae transition) this buffering also avoids the accumulation of undissociated organic acids in the environment that have a potent antimicrobial effect, also against Candida [54][98][99]. Besides lactic acid, acetic acid, 4-hydroxyphenylacetic, succinic, butyric and formic acids are other organic acids present in the vaginal fluid [34][50][100] and whose chemical dissociation can be impacted by changes in pH.
–
The modulation of pH observed to occur along co-cultivation of Candida with L. gasseri shows that the use of supernatants from bacterial cells may not be a good proxy to study the interaction since the amount of undissociated acids is much higher than the one obtained in a co-culture supernatant creating confounding effects (is the inhibition caused by the lactobacilli or by the accumulation of undissociated organic acids?). The use of bacterial supernatants obtained from cultivation in MRS is particularly problematic since these will invariably contain toxic amounts of acetic acid. In this context, some of the inhibitory effects reported in growth and virulence traits of Candida upon exposure to lactobacilli supernatant cultures result, in fact, from the effects of acetic acid. In line with our results, the acetate present in MRS medium was described to have antifungal properties when used in synergy with Lactobacillus rhamnosus [101]. Collectively these findings increase the relevance of using experimental settings based on contact of the species as they appear to establish dynamic interactions that may go beyond the mere accumulation of metabolites in the broth and, therefore, are not fully phenocopied by bacterial supernatants.
–
Co-cultivation of C. albicans or C. glabrata with L. gasseri resulted in decreased viability of the two yeasts (and this is induced by the bacterium and not by accumulated organic acids since we demonstrated that in co-cultures the non-toxic acetate and lactate forms prevailed), either under planktonic biofilm-forming conditions. Previous studies showed that biofilm formation by C. albicans is impacted by co-cultivation with the vaginal species L. crispatus, L. jensenii and L. iners [33][63], however, as the biofilms formed were not microscopically observed the authors did not concluded about the capacity of these species to interact with one another. Both C. albicans and C. glabrata had been described to form mixed biofilms with other bacteria including Streptococcus mutans [102], Staphylococcus aureus [103] or Pseudomonas aeruginosa [104], but to the best of our knowledge this is the first report involving L. gasseri. In the biofilm, the C. glabrata cells were highly susceptible to the presence of L. gasseri, while the reduction of viability of C. albicans cells was considerably lower (opposite to what was observed under planktonic conditions). We cannot rule out that the mechanism of inhibition prompted by L. gasseri in a biofilm may differ from those imparted in planktonic growth as close cell-cell contacts (well demonstrated to occur in Fig. 2) may trigger specific responses. In this context, the previous demonstration that L. paracasei cells respond to direct contact with S. cerevisiae cells is interesting [105]. Another hypothesis is that the access of L. gasseri to C. glabrata cells in the mixed biofilm can be higher due to a stiffer structure of the C. albicans biofilms caused, among other aspects, by high amounts of extracellular matrix [106][107]. The higher sensitivity of C. glabrata to L. gasseri in the mixed biofilm is interesting since this species is much less frequently isolated from the vaginal tract than C. albicans [7][108][109], although it is known to be very high resilient to environmental stress [9][54][110].
–
The fact that the anti-Candida activity prompted by L. gasseri cells depends on the presence of acetate in a concentration-dependent manner is a novel finding of our work. In vivo acetate is present in the vaginal fluid due to metabolic activity of colonizing microbes [50][111][112] and thus it is possible that it contributes to maintain the interference of L. gasseri over Candida. Note that the potential of acetate in augmenting virulence of L. gasseri towards Candida cells was detectable at 4 mM acetate, a concentration within the range found in vaginal fluid in a lactobacilli-dominated vaginal microflora [50][113][114]. Little is known concerning the biology and physiology of L. gasseri and thus the effects of acetate remain to be studied. In some Gram positive species such as L. plantarum, Lactobacillus sakei, Lactobacillus plantarum and L. rhamnosus, acetate potentiated the expression of bacteriocin-encoding genes [91][115][116][117] and thus one possibility is that it may have a similar effect in L. gasseri. Due to their small size and significant inter-species variation, the annotation of bacteriocin-encoding genes is difficult [118][119]. Recent genomic analysis of L. gasseri strains (including ATCC33323) predicted that this species encodes acidocin A, gassericin and helveticin J [67][120][121]. However, in the reference strain only helveticin J, a class III bacteriocin that has been shown to have some activity against two clinical strains of C. albicans and C. glabrata [67], is represented in its genome. The fact that cellular density also affects the modulatory effect of acetate over bacteriocin-encoding genes [91][117][122] creates an important factor that has to be considered in a future study that may aim the study of the molecular mechanism underlying this acetate-induced virulence of L. gasseri against Candida.
MATERIALS AND METHODS
Strains and growth media
In this study we used the reference strains L. gasseri ATCC33323 (acquired from DSMZ); C. glabrata KUE100 (a wild-type strain derived from the CBS138 strain [123]); and Candida albicans SC5314 strains. We have also made use of five clinical strains: L. gasseri, ISTLg97, a vaginal isolate whose species identity was confirmed by Maldi-TOF and based on sequencing of the 16S RNA sequence; C. glabrata VG49, C. glabrata VG216, C. albicans VG217 and C. albicans VG485, all vaginal strains that had been recovered along epidemiological surveys undertaken in the Lisbon area [124]. The MRS medium used to co-cultivate yeasts and bacteria contains, per liter 10 g casein peptone (Gibco), 10 g meat extract (Panreac AppliChem), 5 g yeast extract (Gibco), 20 g glucose (Nzytech), 1 g Tween 80 (Sigma), 2 g K2HPO4 (Merck), 5 g sodium acetate (Merck), 3 g ammonium sulphate (Panreac AppliChem), 0.20 g MgSO4.7H2O (Labchem) and 0.05 g MnSO4.H2O (Sigma). After preparation, pH of MRS was adjusted to 6.2-6.5 using HCl or NaOH. In indicated experiments the sodium acetate used to prepare MRS was replaced by sodium chloride (Honeywell, FlukaTM). YPD medium, used for maintenance of the strains, contains, per liter 20 g glucose (Nzytech), 20 g peptone (Gibco) and 10 g yeast extract (Gibco). Solid YPD or MRS were prepared by supplementing the corresponding liquid medium with 2% and 1.5% agar (Nzytech), respectively. Media were prepared using deionized water and sterilized by autoclaving for 15 min at 121°C and 1 atm.
–
Co-cultivation in liquid MRS medium of L. gasseri with C. glabrata or C. albicans
To examine growth of L. gasseri in liquid MRS alone or in the presence of C. glabrata or C. albicans, a pre-inoculum of each individual species was prepared in MRS and the cells were cultivated, overnight, at 37°C with an orbital agitation of 100 rpm. On the next day, these cells were used to inoculate (at an OD600nm of 0.4 for L. gasseri and 0.1 for the two Candida species) fresh MRS medium. Growth in this co-culture system was accompanied for 4 days at 37°C and using an orbital agitation of 100 rpm, by following the increase in cellular viability of the two species based on the number of colony forming units (CFUs). For this, aliquots of co-cultures were taken, serially diluted and plated on MRS supplemented with 96 mg/L fluconazole (an antifungal concentration that fully prevented growth of Candida colonies and thus only L. gasseri colonies were visible) or in YPD supplemented with 300 mg/L tetracycline (an antibiotic concentration that fully prevented growth of L. gasseri colonies and therefore only Candida colonies were visible). The number of Candida colonies formed on the surface of YPD plates was counted after 2 days of incubation at 30°C, while the number of L. gasseri colonies formed onto the surface of MRS plates was counted after 2 days of plate incubation at 37°C in a Genbox (Biomerieux) with a candle inside to assure microaerophilia [125][126]. As controls we performed single-cultivations of L. gasseri, C. albicans and C. glabrata under the same conditions used for the co-cultivations. Quantification of the amounts of lactic acid, acetic acid or glucose present in the broth during single or multi-species cultivation was performed by HPLC (equipped with an UV detector, for quantification of lactic and acetic acids, and with an RI detector, for quantification of glucose) using an Aminex HPX87H (Biorad®) column and 0.005M H2SO4 (at a flow rate of 0.6 mL/min of) as eluent.
–
Co-cultivation of L. gasseri with C. glabrata or C. albicans under biofilm-forming conditions
To examine growth under biofilm-forming conditions of L. gasseri alone or in co-cultivation with C. albicans or C. glabrata, a pre-inoculum of each individual species was prepared in MRS (or in this same medium containing sodium chloride as a sodium source) and the cells were cultivated overnight at 37°C using an orbital agitation of 100 rpm. These pre-cultures were used to inoculate 200 µL of fresh MRS in plastic µ-slide 8 well plates (Ibidi) so that the initial cell densities (estimated based on OD600nm) were 106 CFU/mL for the two Candida species and 2×108 CFU/mL for L. gasseri. After 24 h of cultivation at 37°C with 25 rpm agitation, the supernatant of the single or co-cultures was removed and the biofilm formed washed with 200 µL of PBS. In order to assess cellular viability in the single- or multi-species biofilms formed, 3 μM of SYTO 9 Green Fluorescent Nucleic Acid Stain (Molecular Probes, Eugene, OR, USA) was added to the single or co-cultures and the cells were left in the dark for 30 minutes. After this time, 4 μM TO-PRO-3 iodide (Molecular Probes, Eugene, OR, USA) were added and the cultures were incubated under the same conditions for another 15 minutes. The gain adjustment in each channel was optimized (and kept during the experiments) taking into account the intensity of the fluorescence signal of live and dead single cells. Live single cells were stained directly after growth, while dead single cells were prepared by heating a cell sample at 65°C for 10 minutes in a dry bath. Then, single or multiple species biofilms were imaged by confocal laser scanning microscopy using a Leica TCS SP5 inverted microscope with a 63x water (1.2 numerical aperture) apochromatic objective. Cells were imaged with the 488 nm Ar+ laser line to detect cells stained with SYTO 9 (emission collected at 500 – 590 nm) and with the 633 nm He-Ne laser line to detect cells stained with TO-PRO-3-Iodide (emission collected at 645-795 nm), a setup that minimizes cross interference between the two channels as described in [127]. To measure the height of the biofilms, the same experimental setup was used with the difference that the cells were only labelled with SYTO9 and the confocal microscopy images used for height quantification.
REFERENCES
- Borjian Boroujeni Z, Shamsaei S, Yarahmadi M, Getso MI, Salimi Khorashad A, Haghighi L, Raissi V, Zareei M, Saleh Mohammadzade A, Moqarabzadeh V, Soleimani A, Raeisi F, Mohseni M, Mohseni MS, Raiesi O (2021). Distribution of invasive fungal infections: Molecular epidemiology, etiology, clinical conditions, diagnosis and risk factors: A 3-year experience with 490 patients under intensive care. Microb Pathog 152: 104616. 10.1016/j.micpath.2020.104616
- Mishra AA, Koh AY (2021). The microbial and host factors that govern Candida gastrointestinal colonization and dissemination. Curr Opin Microbiol 63: 29-35. 10.1016/j.mib.2021.05.012
- Pappas PG, Lionakis MS, Arendrup MC, Ostrosky-Zeichner L, Kullberg BJ (2018). Invasive candidiasis. Nat Rev Dis Primers 4(1): 1-20. 10.1038/nrdp.2018.26
- Denning DW, Bromley MJ (2015). Infectious Disease. How to bolster the antifungal pipeline. Science 347(6229): 1414-1416. 10.1126/science.aaa6097
- Sobel JD (2007). Vulvovaginal candidosis. Lancet 369(9577): 1961-1971. 10.1016/S0140-6736(07)60917-9
- Denning DW, Kneale M, Sobel JD, Rautemaa-Richardson R (2018). Global burden of recurrent vulvovaginal candidiasis: a systematic review. Lancet Infect Dis 18(11): e339-e347. 10.1016/S1473-3099(18)30103-8
- Goncalves B, Ferreira C, Alves CT, Henriques M, Azeredo J, Silva S (2016). Vulvovaginal candidiasis: Epidemiology, microbiology and risk factors. Crit Rev Microbiol 42(6): 905-927. 10.3109/1040841X.2015.1091805
- Boyd Tressler A, Markwei M, Fortin C, Yao M, Procop GW, Soper DE, Goje O (2021). Risks for Recurrent Vulvovaginal Candidiasis Caused by Non-Candida albicans Versus Candida albicans. J Womens Health 30(11): 1588-1596. 10.1089/jwh.2020.8811
- Pfaller MA, Diekema DJ, Turnidge JD, Castanheira M, Jones RN (2019). Twenty Years of the SENTRY Antifungal Surveillance Program: Results for Candida Species From 1997-2016. Open Forum Infect Dis 6(Suppl 1): S79-S94. 10.1093/ofid/ofy358
- Gangneux JP, Cornet M, Bailly S, Fradin C, Feger C, Timsit JF, Leroy O, Sendid B, Bougnoux ME (2018). Clinical Impact of Antifungal Susceptibility, Biofilm Formation and Mannoside Expression of Candida Yeasts on the Outcome of Invasive Candidiasis in ICU: An Ancillary Study on the Prospective AmarCAND2 Cohort. Front Microbiol 9: 2907. 10.3389/fmicb.2018.02907
- Salazar SB, Simoes RS, Pedro NA, Pinheiro MJ, Carvalho M, Mira NP (2020). An Overview on Conventional and Non-Conventional Therapeutic Approaches for the Treatment of Candidiasis and Underlying Resistance Mechanisms in Clinical Strains. J Fungi 6(1): 23. 10.3390/jof6010023
- Fiers WD, Gao IH, Iliev ID (2019). Gut mycobiota under scrutiny: fungal symbionts or environmental transients? Curr Opin Microbiol 50: 79-86. 10.1016/j.mib.2019.09.010
- Hallen-Adams HE, Suhr MJ (2017). Fungi in the healthy human gastrointestinal tract. Virulence 8(3): 352-358. 10.1080/21505594.2016.1247140
- Seed PC (2014). The human mycobiome. Cold Spring Harb Perspect Med 5(5): a019810. 10.1101/cshperspect.a019810
- Suhr MJ, Hallen-Adams HE (2015). The human gut mycobiome: pitfalls and potentials–a mycologist’s perspective. Mycologia 107(6): 1057-1073. 10.3852/15-147
- Huseyin CE, O’Toole PW, Cotter PD, Scanlan PD (2017). Forgotten fungi-the gut mycobiome in human health and disease. FEMS Microbiol Rev 41(4): 479-511. 10.1093/femsre/fuw047
- Sobel JD, Chaim W (1996). Vaginal microbiology of women with acute recurrent vulvovaginal candidiasis. J Clin Microbiol 34(10): 2497-2499. 10.1128/jcm.34.10.2497-2499.1996
- Kalia N, Singh J, Kaur M (2020). Microbiota in vaginal health and pathogenesis of recurrent vulvovaginal infections: a critical review. Ann Clin Microbiol Antimicrob 19(1): 1-19. 10.1186/s12941-020-0347-4
- Tortelli BA, Lewis WG, Allsworth JE, Member-Meneh N, Foster LR, Reno HE, Peipert JF, Fay JC, Lewis AL (2020). Associations between the vaginal microbiome and Candida colonization in women of reproductive age. Am J Obstet Gynecol 222(5): 471. e471-471. e479. 10.1016/j.ajog.2019.10.008
- Serrano MG, Parikh HI, Brooks JP, Edwards DJ, Arodz TJ, Edupuganti L, Huang B, Girerd PH, Bokhari YA, Bradley SP (2019). Racioethnic diversity in the dynamics of the vaginal microbiome during pregnancy. Nat Med 25(6): 1001-1011. 10.1038/s41591-019-0465-8
- Auriemma RS, Scairati R, Del Vecchio G, Liccardi A, Verde N, Pirchio R, Pivonello R, Ercolini D, Colao A (2021). The Vaginal Microbiome: A Long Urogenital Colonization Throughout Woman Life. Front Cell Infect Microbiol 11: 686167. 10.3389/fcimb.2021.686167
- Bommana S, Richards G, Kama M, Kodimerla R, Jijakli K, Read TD, Dean D (2022). Metagenomic Shotgun Sequencing of Endocervical, Vaginal, and Rectal Samples among Fijian Women with and without Chlamydia trachomatis Reveals Disparate Microbial Populations and Function across Anatomic Sites: a Pilot Study. Microbiol Spectr 10(3): e0010522. 10.1128/spectrum.00105-22
- Nunn KL, Forney LJ (2016). Unraveling the Dynamics of the Human Vaginal Microbiome. Yale J Biol Med 89(3): 331-337. 27698617
- Lamont RF, Sobel JD, Akins RA, Hassan SS, Chaiworapongsa T, Kusanovic JP, Romero R (2011). The vaginal microbiome: new information about genital tract flora using molecular based techniques. BJOG 118(5): 533-549. 10.1111/j.1471-0528.2010.02840.x
- Fettweis JM, Serrano MG, Brooks JP, Edwards DJ, Girerd PH, Parikh HI, Huang B, Arodz TJ, Edupuganti L, Glascock AL (2019). The vaginal microbiome and preterm birth. Nat Med 25(6): 1012-1021. 10.1038/s41591-019-0450-2
- Lennard K, Dabee S, Barnabas SL, Havyarimana E, Blakney A, Jaumdally SZ, Botha G, Mkhize NN, Bekker LG, Lewis DA, Gray G, Mulder N, Passmore JS, Jaspan HB (2018). Microbial Composition Predicts Genital Tract Inflammation and Persistent Bacterial Vaginosis in South African Adolescent Females. Infect Immun 86(1): e00410-17. 10.1128/IAI.00410-17
- Norenhag J, Du J, Olovsson M, Verstraelen H, Engstrand L, Brusselaers N (2020). The vaginal microbiota, human papillomavirus and cervical dysplasia: a systematic review and network meta-analysis. BJOG 127(2): 171-180. 10.1111/1471-0528.15854
- McClelland RS, Lingappa JR, Srinivasan S, Kinuthia J, John-Stewart GC, Jaoko W, Richardson BA, Yuhas K, Fiedler TL, Mandaliya KN (2018). Evaluation of the association between the concentrations of key vaginal bacteria and the increased risk of HIV acquisition in African women from five cohorts: a nested case-control study. Lancet Infect Dis 18(5): 554-564. 10.1016/S1473-3099(18)30058-6
- De Seta F, Lonnee-Hoffmann R, Campisciano G, Comar M, Verstraelen H, Vieira-Baptista P, Ventolini G, Lev-Sagie A (2022). The Vaginal Microbiome: III. The Vaginal Microbiome in Various Urogenital Disorders. J Low Genit Tract Dis 26(1): 85-92. 10.1097/LGT.0000000000000645
- Seelig MS (1966). Mechanisms by which antibiotics increase the incidence and severity of candidiasis and alter the immunological defenses. Bacteriol Rev 30(2): 442-459. 10.1128/br.30.2.442-459.1966
- Osset J, Garcia E, Bartolome R, Andreu A (2001). Role of Lactobacillus as protector against vaginal candidiasis. Med Clin 117(8): 285-288. 10.1016/s0025-7753(01)72089-1
- Jian S, SU M-j (2015). Features of vaginal bacteria community in women with recurrent vulvovaginal candidiasis. Reprod Contracept 26(4): 229-238. 10.7669/j.issn.1001-7844.2015.04.0229
- McKloud E, Delaney C, Sherry L, Kean R, Williams S, Metcalfe R, Thomas R, Richardson R, Gerasimidis K, Nile CJ, Williams C, Ramage G (2021). Recurrent Vulvovaginal Candidiasis: a Dynamic Interkingdom Biofilm Disease of Candida and Lactobacillus. mSystems 6(4): e0062221. 10.1128/mSystems.00622-21
- Ceccarani C, Foschi C, Parolin C, D’Antuono A, Gaspari V, Consolandi C, Laghi L, Camboni T, Vitali B, Severgnini M (2019). Diversity of vaginal microbiome and metabolome during genital infections. Sci Rep 9(1): 1-12. 10.1038/s41598-019-50410-x
- Liu MB, Xu SR, He Y, Deng GH, Sheng HF, Huang XM, Ouyang CY, Zhou HW (2013). Diverse vaginal microbiomes in reproductive-age women with vulvovaginal candidiasis. PLoS One 8(11): e79812. 10.1371/journal.pone.0079812
- Ilkit M, Guzel AB (2011). The epidemiology, pathogenesis, and diagnosis of vulvovaginal candidosis: a mycological perspective. Crit Rev Microbiol 37(3): 250-261. 10.3109/1040841X.2011.576332
- Zhou X, Westman R, Hickey R, Hansmann MA, Kennedy C, Osborn TW, Forney LJ (2009). Vaginal microbiota of women with frequent vulvovaginal candidiasis. Infect Immun 77(9): 4130-4135. 10.1128/IAI.00436-09
- Kang CH, Kim Y, Han SH, Kim JS, Paek NS, So JS (2018). In vitro probiotic properties of vaginal Lactobacillus fermentum MG901 and Lactobacillus plantarum MG989 against Candida albicans. Eur J Obstet Gynecol Reprod Biol 228: 232-237. 10.1016/j.ejogrb.2018.07.005
- Parolin C, Marangoni A, Laghi L, Foschi C, Nahui Palomino RA, Calonghi N, Cevenini R, Vitali B (2015). Isolation of Vaginal Lactobacilli and Characterization of Anti-Candida Activity. PLoS One 10(6): e0131220. 10.1371/journal.pone.0131220
- Wang S, Wang Q, Yang E, Yan L, Li T, Zhuang H (2017). Antimicrobial Compounds Produced by Vaginal Lactobacillus crispatus Are Able to Strongly Inhibit Candida albicans Growth, Hyphal Formation and Regulate Virulence-related Gene Expressions. Front Microbiol 8: 564. 10.3389/fmicb.2017.00564
- van de Wijgert J, Verwijs MC (2020). Lactobacilli-containing vaginal probiotics to cure or prevent bacterial or fungal vaginal dysbiosis: a systematic review and recommendations for future trial designs. BJOG 127(2): 287-299. 10.1111/1471-0528.15870
- Andrade JC, Kumar S, Kumar A, Cernakova L, Rodrigues CF (2021). Application of probiotics in candidiasis management. Crit Rev Food Sci Nutr 2(30):8249-8264. 10.1080/10408398.2021.1926905
- Vladareanu R, Mihu D, Mitran M, Mehedintu C, Boiangiu A, Manolache M, Vladareanu S (2018). New evidence on oral L. plantarum P17630 product in women with history of recurrent vulvovaginal candidiasis (RVVC): a randomized double-blind placebo-controlled study. Eur Rev Med Pharmacol Sci 22(1): 262-267. 10.26355/eurrev_201801_14128
- Shenoy A, Gottlieb A (2019). Probiotics for oral and vulvovaginal candidiasis: A review. Dermatol Ther 32(4): e12970. 10.1111/dth.12970
- Alonso-Roman R, Last A, Mirhakkak MH, Sprague JL, Moller L, Grossmann P, Graf K, Gratz R, Mogavero S, Vylkova S, Panagiotou G, Schauble S, Hube B, Gresnigt MS (2022). Lactobacillus rhamnosus colonisation antagonizes Candida albicans by forcing metabolic adaptations that compromise pathogenicity. Nat Commun 13(1): 3192. 10.1038/s41467-022-30661-5
- Zeise KD, Woods RJ, Huffnagle GB (2021). Interplay between Candida albicans and Lactic Acid Bacteria in the Gastrointestinal Tract: Impact on Colonization Resistance, Microbial Carriage, Opportunistic Infection, and Host Immunity. Clin Microbiol Rev 34(4): e0032320. 10.1128/CMR.00323-20
- Graf K, Last A, Gratz R, Allert S, Linde S, Westermann M, Groger M, Mosig AS, Gresnigt MS, Hube B (2019). Keeping Candida commensal: how lactobacilli antagonize pathogenicity of Candida albicans in an in vitro gut model. Dis Model Mech 12(9): dmm039719. 10.1242/dmm.039719
- Zangl I, Pap IJ, Aspock C, Schuller C (2019). The role of Lactobacillus species in the control of Candida via biotrophic interactions. Microbial Cell 7(1): 1-14. 10.15698/mic2020.01.702
- d’Enfert C, Kaune AK, Alaban LR, Chakraborty S, Cole N, Delavy M, Kosmala D, Marsaux B, Frois-Martins R, Morelli M, Rosati D, Valentine M, Xie Z, Emritloll Y, Warn PA, Bequet F, Bougnoux ME, Bornes S, Gresnigt MS, Hube B, Jacobsen ID, Legrand M, Leibundgut-Landmann S, Manichanh C, Munro CA, Netea MG, Queiroz K, Roget K, Thomas V, Thoral C, et al. (2021). The impact of the Fungus-Host-Microbiota interplay upon Candida albicans infections: current knowledge and new perspectives. FEMS Microbiol Rev 45(3): fuaa060. 10.1093/femsre/fuaa060
- Aldunate M, Srbinovski D, Hearps AC, Latham CF, Ramsland PA, Gugasyan R, Cone RA, Tachedjian G (2015). Antimicrobial and immune modulatory effects of lactic acid and short chain fatty acids produced by vaginal microbiota associated with eubiosis and bacterial vaginosis. Front Physiol 6 : 164. 10.3389/fphys.2015.00164
- O’Hanlon DE, Moench TR, Cone RA (2011). In vaginal fluid, bacteria associated with bacterial vaginosis can be suppressed with lactic acid but not hydrogen peroxide. BMC Infect Dis 11: 200. 10.1186/1471-2334-11-200
- Mendling W, Shazly MAE, Zhang L (2022). The Role of Lactic Acid in the Management of Bacterial Vaginosis: A Systematic Literature Review. Future Pharmacology 2(3): 198-213. 10.3390/futurepharmacol2030014
- O’Hanlon DE, Come RA, Moench TR (2019). Vaginal pH measured in vivo: lactobacilli determine pH and lactic acid concentration. BMC Microbiol 19(1): 13. 10.1186/s12866-019-1388-8
- Lourenco A, Pedro NA, Salazar SB, Mira NP (2018). Effect of Acetic Acid and Lactic Acid at Low pH in Growth and Azole Resistance of Candida albicans and Candida glabrata. Front Microbiol 9: 3265. 10.3389/fmicb.2018.03265
- Ballou ER, Avelar GM, Childers DS, Mackie J, Bain JM, Wagener J, Kastora SL, Panea MD, Hardison SE, Walker LA, Erwig LP, Munro CA, Gow NA, Brown GD, MacCallum DM, Brown AJ (2016). Lactate signalling regulates fungal beta-glucan masking and immune evasion. Nat Microbiol 2: 16238. 10.1038/nmicrobiol.2016.238
- Santos CMA, Pires MCV, Leao TL, Silva AKS, Miranda LS, Martins FS, Silva AM, Nicoli JR (2018). Anti-inflammatory effect of two Lactobacillus strains during infection with Gardnerella vaginalis and Candida albicans in a HeLa cell culture model. Microbiology 164(3): 349-358. 10.1099/mic.0.000608
- Tachedjian G, Aldunate M, Bradshaw CS, Cone RA (2017). The role of lactic acid production by probiotic Lactobacillus species in vaginal health. Res Microbiol 168(9-10): 782-792. 10.1016/j.resmic.2017.04.001
- Hearps A, Tyssen D, Srbinovski D, Bayigga L, Diaz D, Aldunate M, Cone R, Gugasyan R, Anderson D, Tachedjian G (2017). Vaginal lactic acid elicits an anti-inflammatory response from human cervicovaginal epithelial cells and inhibits production of pro-inflammatory mediators associated with HIV acquisition. Mucosal Immunol 10(6): 1480-1490. 10.1038/mi.2017.27
- MacAlpine J, Daniel-Ivad M, Liu Z, Yano J, Revie NM, Todd RT, Stogios PJ, Sanchez H, O’Meara TR, Tompkins TA, Savchenko A, Selmecki A, Veri AO, Andes DR, Fidel PL, Jr., Robbins N, Nodwell J, Whitesell L, Cowen LE (2021). A small molecule produced by Lactobacillus species blocks Candida albicans filamentation by inhibiting a DYRK1-family kinase. Nat Commun 12(1): 6151. 10.1038/s41467-021-26390-w
- Nelson J, El-Gendy AO, Mansy MS, Ramadan MA, Aziz RK (2020). The biosurfactants iturin, lichenysin and surfactin, from vaginally isolated lactobacilli, prevent biofilm formation by pathogenic Candida. FEMS Microbiol Lett 367(15): fnaa126. 10.1093/femsle/fnaa126
- Matsuda Y, Cho O, Sugita T, Ogishima D, Takeda S (2018). Culture Supernatants of Lactobacillus gasseri and L. crispatus Inhibit Candida albicans Biofilm Formation and Adhesion to HeLa Cells. Mycopathologia 183(4): 691-700. 10.1007/s11046-018-0259-4
- Ceresa C, Tessarolo F, Caola I, Nollo G, Cavallo M, Rinaldi M, Fracchia L (2015). Inhibition of Candida albicans adhesion on medical-grade silicone by a Lactobacillus-derived biosurfactant. J Appl Microbiol 118(5): 1116-1125. 10.1111/jam.12760
- Matsubara VH, Wang Y, Bandara H, Mayer MPA, Samaranayake LP (2016). Probiotic lactobacilli inhibit early stages of Candida albicans biofilm development by reducing their growth, cell adhesion, and filamentation. Appl Microbiol Biotechnol 100(14): 6415-6426. 10.1007/s00253-016-7527-3
- Hefzy EM, Khalil MAF, Amin AAI, Ashour HM, Abdelaliem YF (2021). Bacteriocin-Like Inhibitory Substances from Probiotics as Therapeutic Agents for Candida Vulvovaginitis. Antibiotics 10(3): 306. 10.3390/antibiotics10030306
- Sharma A, Srivastava S (2014). Anti-Candida activity of two-peptide bacteriocins, plantaricins (Pln E/F and J/K) and their mode of action. Fungal Biol 118(2): 264-275. 10.1016/j.funbio.2013.12.006
- Tan Y, Leonhard M, Moser D, Ma S, Schneider-Stickler B (2018). Inhibitory effect of probiotic lactobacilli supernatants on single and mixed non-albicans Candida species biofilm. Arch Oral Biol 85: 40-45. 10.1016/j.archoralbio.2017.10.002
- Scillato M, Spitale A, Mongelli G, Privitera GF, Mangano K, Cianci A, Stefani S, Santagati M (2021). Antimicrobial properties of Lactobacillus cell-free supernatants against multidrug-resistant urogenital pathogens. Microbiologyopen 10(2): e1173. 10.1002/mbo3.1173
- De Man JC, Rogosa, D., & Sharpe, M. E. (1960). A medium for the cultivation of lactobacilli. J Appl Bacteriol 23(1): 5. 10.1111/j.1365-2672.1960.tb00188.x
- Roy D (2001). Media for the isolation and enumeration of bifidobacteria in dairy products. Int J Food Microbiol 69(3): 167-182. 10.1016/s0168-1605(01)00496-2
- Drell T, Lillsaar T, Tummeleht L, Simm J, Aaspollu A, Vain E, Saarma I, Salumets A, Donders GG, Metsis M (2013). Characterization of the vaginal micro- and mycobiome in asymptomatic reproductive-age Estonian women. PLoS One 8(1): e54379. 10.1371/journal.pone.0054379
- Pramanick R, Mayadeo N, Warke H, Begum S, Aich P, Aranha C (2019). Vaginal microbiota of asymptomatic bacterial vaginosis and vulvovaginal candidiasis: Are they different from normal microbiota? Microb Pathog 134: 103599. 10.1016/j.micpath.2019.103599
- Ma B, France MT, Crabtree J, Holm JB, Humphrys MS, Brotman RM, Ravel J (2020). A comprehensive non-redundant gene catalog reveals extensive within-community intraspecies diversity in the human vagina. Nat Commun 11(1): 940. 10.1038/s41467-020-14677-3
- Chee WJY, Chew SY, Than LTL (2020). Vaginal microbiota and the potential of Lactobacillus derivatives in maintaining vaginal health. Microb Cell Fact 19(1): 203. 10.1186/s12934-020-01464-4
- Hood S, Zoitola E (1988). Effect of low pH on the ability of Lactobacillus acidophilus to survive and adhere to human intestinal cells. J Food Sci53(5): 1514-1516. 10.1111/j.1365-2621.1988.tb09312.x
- Zimmerman T, Gyawali R, Ibrahim S (2017). Autolyse the cell in order to save it? Inducing, then blocking, autolysis as a strategy for delaying cell death in the probiotic Lactobacillus reuteri. Biotechnol Lett 39(10): 1547-1551. 10.1007/s10529-017-2380-8
- Kang O, Vézinz L-P, Laberge S, Simard R (1998). Some factors influencing the autolysis of Lactobacillus bulgaricus and Lactobacillus casei. J Dairy Sci 81(3): 639-646. 10.3168/jds.S0022-0302(98)75618-8
- Masuda T, Hidaka A, Kondo N, Ura T, Itoh T (2005). Intracellular enzyme activities and autolytic properties of Lactobacillus acidophilus and Lactobacillus gasseri. Food Sci Technol Res 11(3): 328-331. 10.3136/fstr.11.328
- Yokoi KJ, Kawasaki K, Taketo A, Kodaira K (2004). Characterization of lytic enzyme activities of Lactobacillus gasseri with special reference to autolysis. Int J Food Microbiol 96(3): 273-279. 10.1016/j.ijfoodmicro.2004.03.021
- Ismail EA, Neve H, Geis A, Heller KJ (2009). Characterization of temperate Lactobacillus gasseri phage LgaI and its impact as prophage on autolysis of its lysogenic host strains. Curr Microbiol 58(6): 648-653. 10.1007/s00284-009-9384-0
- Goncalves B, Fernandes L, Henriques M, Silva S (2020). Environmental pH modulates biofilm formation and matrix composition in Candida albicans and Candida glabrata. Biofouling 36(5): 621-630. 10.1080/08927014.2020.1793963
- Silva S, Henriques M, Martins A, Oliveira R, Williams D, Azeredo J (2009). Biofilms of non-Candida albicans Candida species: quantification, structure and matrix composition. Med Mycol 47(7): 681-689. 10.3109/13693780802549594
- Ramage G, Walle KV, Wickes BL, Lopez-Ribot JL (2001). Characteristics of biofilm formation by Candida albicans. Rev Iberoam Micol 18(4): 163-170. 15496122
- Baillie GS, Douglas LJ (1999). Role of dimorphism in the development of Candida albicans biofilms. J Med Microbiol 48(7): 671-679. 10.1099/00222615-48-7-671
- Kerstens M, Boulet G, Tritsmans C, Horemans T, Hellings M, Delputte P, Maes L, Cos P (2014). Flow cytometric enumeration of bacteria using TO-PRO(R)-3 iodide as a single-stain viability dye. J Lab Autom 19(6): 555-561. 10.1177/2211068214546745
- Tavecchio M, Simone M, Bernasconi S, Tognon G, Mazzini G, Erba E (2008). Multi-parametric flow cytometric cell cycle analysis using TO-PRO-3 iodide (TP3): detailed protocols. Acta Histochem 110(3): 232-244. 10.1016/j.acthis.2007.10.007
- George J, Halami PM (2019). Presence of extracellular DNA & protein in biofilm formation by gentamicin-resistant Lactobacillus plantarum. Indian J Med Res 149(2): 257-262. 10.4103/ijmr.IJMR_2022_17
- Muscariello L, Marino C, Capri U, Vastano V, Marasco R, Sacco M (2013). CcpA and three newly identified proteins are involved in biofilm development in Lactobacillus plantarum. J Basic Microbiol 53(1): 62-71. 10.1002/jobm.201100456
- Fernandez Ramirez MD, Smid EJ, Abee T, Nierop Groot MN (2015). Characterisation of biofilms formed by Lactobacillus plantarum WCFS1 and food spoilage isolates. Int J Food Microbiol 207: 23-29. 10.1016/j.ijfoodmicro.2015.04.030
- Martins M, Uppuluri P, Thomas DP, Cleary IA, Henriques M, Lopez-Ribot JL, Oliveira R (2010). Presence of extracellular DNA in the Candida albicans biofilm matrix and its contribution to biofilms. Mycopathologia 169(5): 323-331. 10.1007/s11046-009-9264-y
- Ge J, Kang J, Ping W (2019). Effect of Acetic Acid on Bacteriocin Production by Gram-Positive Bacteria. J Microbiol Biotechnol 29(9): 1341-1348. 10.4014/jmb.1905.05060
- Meng F, Zhao H, Nie T, Lu F, Zhang C, Lu Y, Lu Z (2021). Acetate Activates Lactobacillus Bacteriocin Synthesis by Controlling Quorum Sensing. Appl Environ Microbiol 87(13): e0072021. 10.1128/AEM.00720-21
- Matsubara VH, Wang Y, Bandara HM, Mayer MP, Samaranayake LP (2016). Probiotic lactobacilli inhibit early stages of Candida albicans biofilm development by reducing their growth, cell adhesion, and filamentation. Appl Microbiol Biotechnol 100(14): 6415-6426. 10.1007/s00253-016-7527-3
- James KM, MacDonald KW, Chanyi RM, Cadieux PA, Burton JP (2016). Inhibition of Candida albicans biofilm formation and modulation of gene expression by probiotic cells and supernatant. J Med Microbiol 65(4): 328-336. 10.1099/jmm.0.000226
- Danhof HA, Vylkova S, Vesely EM, Ford AE, Gonzalez-Garay M, Lorenz MC (2016). Robust Extracellular pH Modulation by Candida albicans during Growth in Carboxylic Acids. mBio 7(6): e01646-16. 10.1128/mBio.01646-16
- Vylkova S, Carman AJ, Danhof HA, Collette JR, Zhou H, Lorenz MC (2011). The fungal pathogen Candida albicans autoinduces hyphal morphogenesis by raising extracellular pH. mBio 2(3): e00055-00011. 10.1128/mBio.00055-11
- Kasper L, Seider K, Gerwien F, Allert S, Brunke S, Schwarzmuller T, Ames L, Zubiria-Barrera C, Mansour MK, Becken U, Barz D, Vyas JM, Reiling N, Haas A, Haynes K, Kuchler K, Hube B (2014). Identification of Candida glabrata genes involved in pH modulation and modification of the phagosomal environment in macrophages. PLoS One 9(5): e96015. 10.1371/journal.pone.0096015
- Vylkova S, Lorenz MC (2014). Modulation of phagosomal pH by Candida albicans promotes hyphal morphogenesis and requires Stp2p, a regulator of amino acid transport. PLoS Pathog 10(3): e1003995. 10.1371/journal.ppat.1003995
- Moosa MY, Sobel JD, Elhalis H, Du W, Akins RA (2004). Fungicidal activity of fluconazole against Candida albicans in a synthetic vagina-simulative medium. Antimicrob Agents Chemother 48(1): 161-167. 10.1128/AAC.48.1.161-167.2004
- Kasper L, Miramon P, Jablonowski N, Wisgott S, Wilson D, Brunke S, Hube B (2015). Antifungal activity of clotrimazole against Candida albicans depends on carbon sources, growth phase and morphology. J Med Microbiol 64(7): 714-723. 10.1099/jmm.0.000082
- Vitali B, Cruciani F, Picone G, Parolin C, Donders G, Laghi L (2015). Vaginal microbiome and metabolome highlight specific signatures of bacterial vaginosis. Eur J Clin Microbiol Infect Dis 34(12): 2367-2376. 10.1007/s10096-015-2490-y
- Stiles J, Penkar S, Plockova M, Chumchalova J, Bullerman LB (2002). Antifungal activity of sodium acetate and Lactobacillus rhamnosus. J Food Prot 65(7): 1188-1191. 10.4315/0362-028x-65.7.1188
- Khoury ZH, Vila T, Puthran TR, Sultan AS, Montelongo-Jauregui D, Melo MAS, Jabra-Rizk MA (2020). The Role of Candida albicans Secreted Polysaccharides in Augmenting Streptococcus mutans Adherence and Mixed Biofilm Formation: In vitro and in vivo Studies. Front Microbiol 11(307. 10.3389/fmicb.2020.00307
- Kong EF, Tsui C, Kucharikova S, Andes D, Van Dijck P, Jabra-Rizk MA (2016). Commensal Protection of Staphylococcus aureus against Antimicrobials by Candida albicans Biofilm Matrix. mBio 7(5): e01365-16. 10.1128/mBio.01365-16
- Bandara HM, Yau JY, Watt RM, Jin LJ, Samaranayake LP (2010). Pseudomonas aeruginosa inhibits in-vitro Candida biofilm development. BMC Microbiol 10: 125. 10.1186/1471-2180-10-125
- Yamasaki-Yashiki S, Sawada H, Kino-Oka M, Katakura Y (2017). Analysis of gene expression profiles of Lactobacillus paracasei induced by direct contact with Saccharomyces cerevisiae through recognition of yeast mannan. Biosci Microbiota Food Health 36(1): 17-25. 10.12938/bmfh.BMFH-2016-015
- Kucharikova S, Tournu H, Lagrou K, Van Dijck P, Bujdakova H (2011). Detailed comparison of Candida albicans and Candida glabrata biofilms under different conditions and their susceptibility to caspofungin and anidulafungin. J Med Microbiol 60(Pt 9): 1261-1269. 10.1099/jmm.0.032037-0
- Pierce CG, Vila T, Romo JA, Montelongo-Jauregui D, Wall G, Ramasubramanian A, Lopez-Ribot JL (2017). The Candida albicans Biofilm Matrix: Composition, Structure and Function. J Fungi 3(1): 14. 10.3390/jof3010014
- Salazar SB, Pinheiro MJF, Sotti-Novais D, Soares AR, Lopes MM, Ferreira T, Rodrigues V, Fernandes F, Mira NP (2022). Disclosing azole resistance mechanisms in resistant Candida glabrata strains encoding wild-type or gain-of-function CgPDR1 alleles through comparative genomics and transcriptomics. G3 12(7). 10.1093/g3journal/jkac110
- Pfaller MA, Diekema DJ, Gibbs DL, Newell VA, Ellis D, Tullio V, Rodloff A, Fu W, Ling TA, Global Antifungal Surveillance G (2010). Results from the ARTEMIS DISK Global Antifungal Surveillance Study, 1997 to 2007: a 10.5-year analysis of susceptibilities of Candida Species to fluconazole and voriconazole as determined by CLSI standardized disk diffusion. J Clin Microbiol 48(4): 1366-1377. 10.1128/JCM.02117-09
- Hassan Y, Chew SY, Than LTL (2021). Candida glabrata: Pathogenicity and Resistance Mechanisms for Adaptation and Survival. J Fungi 7(8): 667. 10.3390/jof7080667
- Gajer P, Brotman RM, Bai G, Sakamoto J, Schutte UM, Zhong X, Koenig SS, Fu L, Ma ZS, Zhou X, Abdo Z, Forney LJ, Ravel J (2012). Temporal dynamics of the human vaginal microbiota. Sci Transl Med 4(132): 132ra152. 10.1126/scitranslmed.3003605
- Nicolo S, Tanturli M, Mattiuz G, Antonelli A, Baccani I, Bonaiuto C, Baldi S, Nannini G, Menicatti M, Bartolucci G, Rossolini GM, Amedei A, Torcia MG (2021). Vaginal Lactobacilli and Vaginal Dysbiosis-Associated Bacteria Differently Affect Cervical Epithelial and Immune Homeostasis and Anti-Viral Defenses. Int J Mol Sci 22(12): 6487. 10.3390/ijms22126487
- Boskey ER, Cone RA, Whaley KJ, Moench TR (2001). Origins of vaginal acidity: high D/L lactate ratio is consistent with bacteria being the primary source. Hum Reprod 16(9): 1809-1813. 10.1093/humrep/16.9.1809
- O’Hanlon DE, Moench TR, Cone RA (2013). Vaginal pH and microbicidal lactic acid when lactobacilli dominate the microbiota. PLoS One 8(11): e80074. 10.1371/journal.pone.0080074
- Nilsson L, Nielsen MK, Ng Y, Gram L (2002). Role of acetate in production of an autoinducible class IIa bacteriocin in Carnobacterium piscicola A9b. Appl Environ Microbiol 68(5): 2251-2260. 10.1128/AEM.68.5.2251-2260.2002
- Kelly W, Asmundson R, Huang C (1996). Characterization of plantaricin KW30, a bacteriocin produced by Lactobacillus plantarum. J Appl Bacteriol 81(6): 657-662. 10.1111/j.1365-2672.1996.tb03561.x
- Meng F, Lu F, Du H, Nie T, Zhu X, Connerton IF, Zhao H, Bie X, Zhang C, Lu Z, Lu Y (2021). Acetate and auto-inducing peptide are independent triggers of quorum sensing in Lactobacillus plantarum. Mol Microbiol 116(1): 298-310. 10.1111/mmi.14709
- Zhou X, Yang B, Stanton C, Ross RP, Zhao J, Zhang H, Chen W (2020). Comparative analysis of Lactobacillus gasseri from Chinese subjects reveals a new species-level taxa. BMC Genomics 21(1): 119. 10.1186/s12864-020-6527-y
- de Jong A, van Heel AJ, Kok J, Kuipers OP (2010). BAGEL2: mining for bacteriocins in genomic data. Nucleic Acids Res 38(Web Server issue): W647-651. 10.1093/nar/gkq365
- Ito Y, Kawai Y, Arakawa K, Honme Y, Sasaki T, Saito T (2009). Conjugative plasmid from Lactobacillus gasseri LA39 that carries genes for production of and immunity to the circular bacteriocin gassericin A. Appl Environ Microbiol 75(19): 6340-6351. 10.1128/AEM.00195-09
- Garcia-Gutierrez E, O’Connor PM, Colquhoun IJ, Vior NM, Rodriguez JM, Mayer MJ, Cotter PD, Narbad A (2020). Production of multiple bacteriocins, including the novel bacteriocin gassericin M, by Lactobacillus gasseri LM19, a strain isolated from human milk. Appl Microbiol Biotechnol 104(9): 3869-3884. 10.1007/s00253-020-10493-3
- Maldonado-Barragán A, West SA (2020). The cost and benefit of quorum sensing-controlled bacteriocin production in Lactobacillus plantarum. J Evol Biol 33(1): 101-111. 10.1111/jeb.13551
- Ueno K, Matsumoto Y, Uno J, Sasamoto K, Sekimizu K, Kinjo Y, Chibana H (2011). Intestinal resident yeast Candida glabrata requires Cyb2p-mediated lactate assimilation to adapt in mouse intestine. PLoS One 6(9): e24759. 10.1371/journal.pone.0024759
- Cunha DV, Salazar SB, Lopes MM, Mira NP (2017). Mechanistic Insights Underlying Tolerance to Acetic Acid Stress in Vaginal Candida glabrata Clinical Isolates. Front Microbiol 8: 259. 10.3389/fmicb.2017.00259
- Biswas S, Keightley A, Biswas I (2019). Characterization of a stress tolerance-defective mutant of Lactobacillus rhamnosus LRB. Mol Oral Microbiol 34(4): 153-167. 10.1111/omi.12262
- Bhatia SJ, Kochar N, Abraham P, Nair NG, Mehta AP (1989). Lactobacillus acidophilus inhibits growth of Campylobacter pylori in vitro. J Clin Microbiol 27(10): 2328-2330. 10.1128/jcm.27.10.2328-2330.1989
- Pinto SN, Dias SA, Cruz AF, Mil-Homens D, Fernandes F, Valle J, Andreu D, Prieto M, Castanho M, Coutinho A, Veiga AS (2019). The mechanism of action of pepR, a viral-derived peptide, against Staphylococcus aureus biofilms. J Antimicrob Chemother 74(9): 2617-2625. 10.1093/jac/dkz223
–
SUPPLEMENTAL INFORMATION
![]() Download Supplemental Information
Download Supplemental Information
ACKNOWLEDGMENTS
This study was supported by funds received from FCT - Fundação para a Ciência e a Tecno-logia, I.P., in the scope of the project LactoCan (PTDC/BIA-MIC/31515/2017); the project supporting the Research Unit Institute for Bioengineering and Biosci-ences (UIDB/04565/2020) and the project supporting Institute for Health and Bioeconomy, i4HB (LA/P/0140/2020). FCT is also acknowledged for funding a PhD grant (PD/BD/143026/2018) to NAP through the doctoral program DP_AEM.
COPYRIGHT
© 2023

Acetate modulates the inhibitory effect of Lactobacillus gasseri against the pathogenic yeasts Candida albicans and Candida glabrata by Pedro et al. is licensed under a Creative Commons Attribution 4.0 International License.