Reviews:
Microbial Cell, Vol. 2, No. 6, pp. 182 - 196; doi: 10.15698/mic2015.06.205
Coordinate responses to alkaline pH stress in budding yeast
Departament de Bioquímica i Biologia Molecular & Institut de Biotecnologia i Biomedicina, Universitat Autònoma de Barcelona, Bellaterra 08193, Barcelona, Spain.
Keywords: calcineurin, Snf1, PKA, transcriptomics, S. cerevisiae.
Abbreviation:
CDRE - calcineurin dependent response element,
ESCRT - endosomal sorting complex required for transport,
MAPK - mitogen activated protein kinase,
STRE - stress response element.
Received originally: 09/02/2015 Received in revised form: 05/05/2015
Accepted: 08/05/2015
Published: 22/05/2015
Correspondence:
Dr. Joaquín Ariño, Institut de Biotecnologia i Biomedicina, Universitat Autònoma de Barcelona; Bellaterra 08193, Barcelona, Spain Joaquin.Arino@uab.es
Conflict of interest statement: The authors declare no conflict of interest.
Please cite this article as: Albert Serra-Cardona, David Canadell and Joaquín Ariño (2015). Coordinate responses to alkaline pH stress in budding yeast. Microbial Cell 2(6): 182-196.
Abstract
Alkalinization of the medium represents a stress condition for the budding yeast Saccharomyces cerevisiae to which this organism responds with profound remodeling of gene expression. This is the result of the modulation of a substantial number of signaling pathways whose participation in the alkaline response has been elucidated within the last ten years. These regulatory inputs involve not only the conserved Rim101/PacC pathway, but also the calcium-activated phosphatase calcineurin, the Wsc1-Pkc1-Slt2 MAP kinase, the Snf1 and PKA kinases and oxidative stress-response pathways. The uptake of many nutrients is perturbed by alkalinization of the environment and, consequently, an impact on phosphate, iron/copper and glucose homeostatic mechanisms can also be observed. The analysis of available data highlights cases in which diverse signaling pathways are integrated in the gene promoter to shape the appropriate response pattern. Thus, the expression of different genes sharing the same signaling network can be coordinated, allowing functional coupling of their gene products.
INTRODUCTION
The ability of living cells for adaptation to changes in the environment is crucial for survival. Extracellular pH is a key parameter that influences many aspects of the biology of cells. The budding yeast Saccharomyces cerevisiae grows under a relatively broad range of external pH, but proliferates far better at acidic than at neutral or alkaline pH. The maintenance of a suitable acidic environment is achieved by active proton extrusion mediated by the plasma-membrane H+-ATPase, encoded by the essential PMA1 gene [1]. In collaboration with the vacuolar H+-ATPase, Pma1 is also crucial for maintenance of cytosolic pH homeostasis [2]. The electrochemical gradient of protons across the plasma membrane is required for all secondary symporters and antiporters critical for the uptake of different nutrients, including diverse cations, mainly potassium [3][4]. In addition, external alkaline pH prompts differentiation programs, such as haploid invasive growth and sporulation, which are both inhibited under acidic growth [5][6]. Most S. cerevisiae laboratory strains can hardly grow in a medium buffered above pH 8.0-8.2, and alkalinization of the medium has widespread effects on the yeast physiology, as inferred from the very diverse nature of the near 300 mutants known to display growth defects at neutral or moderately alkaline pH [7][8][9]. Many of these mutants show vacuolar defects, particularly those lacking structural components of the vacuolar ATPase or those required for its assembly, which are extremely sensitive to even moderate alkalinization of the medium [8][10]. In addition, it has been shown that under some circumstances V-ATPase complexes are stabilized when the extracellular pH increases, becoming less susceptible to dissociation, and increasing their activity [11][12]. All these evidence clearly indicate that a functional vacuole is required for normal alkaline pH tolerance. Although a vacuolar role in maintaining proper cytosolic pH has been proposed, the necessity for a functional vacuole is not completely understood [13], and has probably a multifactorial nature that will be discussed further in other sections of this review.
–
The remodeling of gene expression is at the basis of all stress responses in S. cerevisiae [14]. The influence of alkaline pH on gene expression was first documented in Aspergillus nidulans, and later in Candida albicans, and the mechanisms behind this response have been characterized in some detail in these organisms [15][16][17]. Comparatively, knowledge on the adaptive responses of S. cerevisiae to alkaline pH was almost inexistent until early in the last decade [9]. However, a wealth of information has been generated since then, allowing the dissection of diverse pathways that are transiently modulated when budding yeast is exposed to environmental alkalinization, which – as expected – entails extensive regulation at the level of gene expression. In this work we describe these pathways and how they interact to shape a transient response that allows adaptation and, eventually, survival of the yeast cell. The knowledge of the adaptive mechanisms to alkaline pH will be useful, not only from the point of view of fundamental research, helping to dissect signaling pathways and their downstream targets, but also for industrial biotechnology, guiding the design of high-pH tolerant strains able to survive in demanding fermentation conditions. In addition, the signals that trigger the morphological change from the Candida albicans non-pathogenic yeast form to the pathogenic hyphal form are produced in response to a shift to neutral/alkaline pH. Thus, these pathways are also related to fungal virulence.
THE CALCINEURIN PATHWAY AND THE ALKALINE pH STRESS RESPONSE
Exposure of yeast cells to alkaline stress promotes a very fast entry of calcium from the environment that requires the Mid1/Cch1 high-affinity calcium transport system [18]. Although the precise mechanism is not known, it has been proposed that Mid1 is an stretch-activated cation channel [19][20], whereas Cch1 would be a voltage-gated calcium channel [20][21]. Interestingly, alkaline pH activates the Slt2 MAP kinase pathway through the Wsc1 surface sensor [22] (see below, section “OTHER STRESS SIGNALING PATHWAYS ACTIVATED BY ALKALINE pH STRESS”), which subsequently was demonstrated to be a mechanosensor [23]. Therefore, it is conceivable that alkalinization of the medium imposes a mechanic stress to the yeast cell surface (see below, section “OTHER STRESS SIGNALING PATHWAYS ACTIVATED BY ALKALINE pH STRESS”). In the case of Mid1, this stress would increase its ability to mediate calcium uptake, whereas depolarization associated to alkalinization of the cell surface would activate Cch1. This concerted effect would promote the fast calcium entry and calcineurin activation characteristics of the response to high pH stress.
–
In budding yeast the calcium-activated Ser/Thr protein phosphatase calcineurin is a heterodimer, composed of one of two redundant catalytic subunits, encoded by CNA1 and CNA2, plus a sole regulatory subunit, encoded by the gene CNB1 [24]. The importance of calcineurin in high pH tolerance in S. cerevisiae was illustrated by the early report that cnb1 mutants or wild-type cells treated with the calcineurin inhibitor FK-506 were sensitive to alkalinization of the medium [25][26] and is highlighted by the observation that more than 40% of the deletion mutants found to be sensitive to chemical inhibition of calcineurin by the drug FK-506 were also sensitive to alkaline pH [18]. The calcineurin pathway appears necessary for adaptation to alkaline environments not only in S. cerevisiae, but also in pathogenic fungi, such as Candida albicans [27] or Cryptococcus neoformans [28].
–
A major target of calcineurin is the zinc-finger transcription factor Crz1 (also known as Tcn1 and Hal8). Calcineurin dephosphorylates Crz1 [29][30][31], which then enters the nucleus and binds to specific sequences (5′-GNGGC(G/T)CA-3′), known as Calcineurin Dependent Response Elements (CDRE), present in the promoters of calcineurin-responsive genes, such as the Na+-ATPase ENA1 [29][31][32]. Yoshimoto and coworkers showed that among the genes whose expression is affected upon addition of calcium or sodium cations to the medium, around 160 genes are largely dependent on calcineurin and, in most cases, also on the presence of Crz1 [33]. The calcineurin/Crz1 tandem is conserved in fungi and other lower eukaryotes (see [34] for a recent review).
–
The importance of calcium signaling in the transcriptional response to high pH stress was pointed out by Serrano and coworkers [35], who showed that the calcineurin/Crz1 pathway significantly contributed to the rapid induction of ENA1 and PHO89 upon alkalinization of the medium. Subsequent work demonstrated that calcineurin is responsible for the induction of a significant subset of alkali-responsive genes [18]. Activation of calcineurin upon alkali stress can be explained by a very fast (seconds) entry of calcium from the extracellular medium into the cell through the Cch1-Mid1 high-affinity calcium channel. The transient rise in intracellular calcium activates calcineurin and promotes entry of Crz1 into the nucleus and binding to its targets in a matter of a few minutes after pH increase [36]. ChIP-Seq data generated in our laboratory (S. Petrezsélyová and J. Ariño, unpublished results) revealed an enrichment of Crz1 at around 190 gene promoters in response to alkaline pH stress, of which 24 were previously identified as Crz1-dependent upon alkaline conditions [18][36]. Crz1 has been identified as a direct target for several protein kinases. Phosphorylation of Crz1 by Hrr25 (a homolog of human casein kinase 1δ) or the Pho80/Pho85 kinase stimulates nuclear exclusion of the transcription factor [37][38], whereas phosphorylation by PKA would inhibit its nuclear import [39]. Since the activity of PKA and Pho85 is affected by alkalinization of the medium (see below), regulation of Crz1 appears at the crossroad of different signaling pathways (Figure 1).
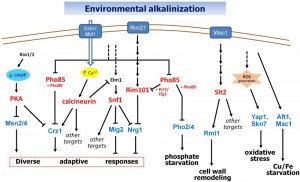 | FIGURE 1: Known signaling pathways in alkaline pH response. Discontinuous lines denote still uncharacterized links. See text for details. |
In contrast with the remarkable sensitivity of calcineurin-deficient strains to high pH, crz1 mutants have been reported to be insensitive or only marginally affected by alkaline pH [29]. This suggests that the Crz1-mediated transcriptional response has a relatively minor role for survival at high pH, and that additional targets for calcineurin, with a relevant role in high pH adaptation, must exist. Possible examples of these targets could be the tail-anchored ER membrane proteins Hph1 and Hph2 (renamed as Frt1 and Frt2). hph1 hph2 cells show impaired growth under alkaline pH, salt stress, and cell wall-damaging agents. Hph1, but not Hph2, interacts with and is a substrate of calcineurin. Thus, calcineurin positively modulates Hph1 in a Crz1-independent fashion [40]. More recent work has revealed that Hph1 and Hph2 are novel components of the Sec63/Sec62 posttranslational translocation complex that aid in vacuolar proton ATPase biogenesis [41]. This function could be at the basis of the alkali-sensitive phenotype of the hph1 hph2 strain, since cells with impaired V-ATPase function cannot properly acidify the vacuole and, consequently, fail to grow at alkaline pH. In this regard, it is known that the electrochemical gradient between the cytosol and vacuolar lumen created by the V-ATPase is required for transport of diverse metabolites and ions, among them, calcium. As mentioned above, exposure to high pH triggers a potent and almost immediate entry of extracellular calcium. It has been shown that cells lacking V-ATPase activity (vma– mutants) lose viability after a brief exposure to elevated extracellular Ca2+ and that under these circumstances, the V-ATPase-driven Vcx1 H+/Ca2+ antiporter is the main activity restoring cytosolic Ca2+ concentrations [42]. Therefore, high pH-induced rise in cytosolic calcium is likely to be particularly detrimental in vma– cells and proper sequestration of calcium could be one of the roles of the vacuole upon high pH stress.
THE PARTICIPATION OF THE Rim101 PATHWAY IN THE ALKALINE RESPONSE
Although initially identified in S. cerevisiae as a positive regulator of meiotic gene expression and sporulation [43], the role of Rim101 in mediating alkaline pH adaptation and signaling in budding yeast became evident by comparison with the well-established pH-responsive pathway of its homolog in Aspergillus nidulans, PacC [16][44]. In fact, recent evidence suggest that, in budding yeast cells forming a colony, alkali signaling between cells likely promotes the expansion of the region of sporulation in a way that depends on Rim101-mediated signaling [45].
–
Rim101 is a C2H2 zinc-finger transcription factor that binds to TGCCAAG sequences in the promoters of responsive genes, and its activation pathway resembles in many aspects that of A. nidulans PacC. The system includes a plasma-membrane located putative sensor system, composed by the 7 transmembrane proteins Rim21 (equivalent to A. nidulans PalH) and Dfg16, the 3 transmembrane Rim9 (PalI) protein, and the α-arrestin Rim8 (PalF). It has been proposed that a depolarization of the plasma membrane, caused either by external alkalinization or other mechanisms, can activate the signal transduction in a Rim21-dependent manner (whereas Dfg16 and Rim9 would only have auxiliary functions) [46]. The mechanism also involves the activation of Rim101 by alkaline pH-stimulated C-terminal proteolytic cleavage, and it was proposed that the proteolytic complex containing the Rim13 protease is formed and activated by an endocytic mechanism on a ESCRT-III endosomal membrane complex (see [47] for a review). However, evidence has been accumulating that PacC and Rim101 signaling occurs not at the late endosome via endocytosis, but at the plasma membrane by recruitment of downstream components of the pathway [48][49][50], (see [51] for a recent review), and that in S. cerevisiae such recruitment involves Rsp5-mediated ubiquinitation events of unknown components other than Rim8 [48].
–
In contrast to PacC, which acts directly as transcriptional activator of “alkaline” genes and as a repressor of “acid” genes, in S. cerevisiae Rim101 exerts its role as a repressor, thus controlling the expression of two other transcription factors, NRG1 and SMP1 [52]. A rim101 mutant shows high pH sensitivity and a decreased expression of the Na+-ATPase encoding gene ENA1, and both phenotypes are abolished by deletion of NRG1, encoding a transcriptional repressor, thus suggesting that the role of Rim101 in alkaline stress (and salt stress) would be mediated by its effect on the expression of NRG1 [52]. Since Nrg1 function is known to be inhibited by the Snf1 protein kinase [53][54], and Snf1 is activated by alkaline stress [55][56], Nrg1 appears to be at the convergence of two high pH stress-activated pathways (see Figure 1 and below, section “THE CONTRIBUTION OF THE Snf1 PATHWAY TO THE RESPONSE TO ALKALINE pH”).
–
It is worth noting that Rim101 is also involved in weak acid stress resistance [57], which suggests that Rim101 may retain some functionality at acidic conditions. This raises the question of whether cells might share overlapping mechanisms of resistance to both kinds of stresses. We have assembled a list of 292 alkaline pH sensitive mutations from existing literature [7][8][22] and crossed it with the set 629 mutants sensitive to acetic described in Mira et al. [58], obtaining a subset of 95 deletion strains sensitive to both stresses. The overlap, a 2.5-fold excess from what could be expected by selecting genes randomly, was enriched only in genes related to vacuolar acidification, which could be anticipated since this process has been proposed to support recovery of cytosolic pH upon treatment of the cells with weak acids, and mutants in the V-ATPase are highly sensitive to alkaline pH. On the other hand, analysis of the set of genes induced by both alkaline stress and acetic acid exposure yielded 33 genes. Gene Ontology analysis revealed an enrichment only in genes related to the broad category “response to stress” (p-value 2.20E-04). Therefore, it seems that the overlap in the response mechanisms to both kinds of stresses is very limited. This is not surprising, as it has been demonstrated that the transcription factor Haa1 is a major determinant in the transcriptional response to acetic acid stress [59], whereas this factor seems to play little role in the transcriptional response to high pH stress, nor its mutation renders cells sensitive to this condition.
–
Among other functions, the cyclin-dependent kinase Pho85 is involved in the transcriptional response to phosphate starvation and alkaline stress (see below). A strain lacking Pho85 is moderately sensitive to high pH [8], and mutation of rim101 suppresses the growth defect of pho85 cells under this condition [60]. Interestingly, Rim101-dependent genes, such as NRG1, are strongly repressed in a pho85 strain, and these cells accumulate the repressor in the nucleus. The same authors showed that phosphorylation of Rim101 plays an important role in determining its cellular location and that Pho85 could phosphorylate Rim101 in vitro, although they could not obtain evidence for in vivo phosphorylation [60]. Therefore, although the mechanism is still unclear, the evidence suggests that Pho85 exerts a negative effect on Rim101 function that could be relieved upon inhibition of Pho85 by alkalinization of the medium.
–
Alkaline pH is likely to cause cell wall damage and results in fast activation of the Slt2 pathway (see below, section “OTHER STRESS SIGNALING PATHWAYS ACTIVATED BY ALKALINE pH STRESS”). It has been reported that the Rim101 pathway contributes to cell wall remodeling in parallel with the protein kinase C (Pkc1)-Slt2 pathway and that it becomes essential in the absence of the Slt2 MAP kinase [61]. Interestingly, this role seems to be conserved for the PacC/Rim101 pathway in pathogenic fungi such as C. albicans or C. neoformans [62][63][64].
THE PKA PATHWAY IS TRANSIENTLY INHIBITED BY ALKALINIZATION OF THE ENVIRONMENT
The PKA pathway has great relevance in nutrient sensing and stress responses [65]. It has been reported that sudden acidification of the cytosol results in transitory increases in cAMP levels, thus activating the PKA pathway [66][67]. More recent work has demonstrated that mutations that activate the PKA pathway, such as ira1 ira2 or bcy1 cause sensitivity to alkaline pH, whereas deactivation of the pathway increases high pH stress tolerance [68]. In the same work it is shown that alkalinization causes a transient decrease in cAMP and the rapid (2-5 min) nuclear localization of Msn2, a transcription factor that, together with Mns4, contributes to PKA-dependent regulation of the expression of STRE-controlled genes [69][70]. Indeed, genome-wide transcriptomic analysis revealed that Msn2/Msn4 contributes to the short-term (10 min) up-regulation of nearly 50% of the genes induced by alkaline pH stress, whereas at longer times (30 min) the relevance of Msn2/Msn4 in the response substantially decreases [68]. Therefore, inhibition of the PKA pathway by alkaline pH represents a substantial component of the adaptive response to alkalinization of the environment.
–
Casado and coworkers also reported that only mutations in the Ras1/Ras2 branch, upstream adenylate cyclase (Cyr1) and PKA, affected alkaline pH tolerance, whereas mutations in diverse genes involved in the Gpr1/Gpa2 glucose sensor system do not produce phenotypic effects [68]. These observations, and the fact that the uptake of glucose is not impaired by alkalinization of the medium [56], could indicate that the impact of alkaline pH on the PKA pathway occurs in response to alteration of intracellular glucose metabolism and not to defects in the transport process and/or extracellular glucose detection. Recently, Broggi and coworkers [71] showed that high pH stress affects localization of Ras2, which becomes transiently excluded from the nucleus and translocates to the plasma membrane, and that the treatment triggers a fast increase in the Ras2-GTP/total Ras2 ratio. The authors postulate that alkalinization down-regulates the cAMP/PKA pathway acting on element(s) downstream Ras and that the increase of Ras2-GTP might be explained by a decrease of the reported feedback inhibition operated by PKA on Ras2 [72].
–
The activation state of the PKA pathway is likely to influence on other signaling pathways affected by high pH stress. For instance, as mentioned above (section “THE CALCINEURIN PATHWAY AND THE ALKALINE pH STRESS RESPONSE”), PKA can phosphorylate Crz1, thus opposing the action of calcineurin [39]. Therefore, inhibition of PKA may result in further enhancement of calcineurin/Crz1-mediated responses. In addition, it was reported that, upon calcium treatment, Crz1 may exert a destabilization effect on Msn2 levels [73]. Since alkalinization of the medium triggers a rapid burst of intracellular calcium, it is conceivable that high-pH induced entry of Crz1 in the nucleus may serve, in addition to activate calcineurin/Crz1-responsive promoters, as a negative-feedback system to avoid persistent activation of Msn2/4-responsive genes, thus explaining the transient Msn2/4-dependent transcriptional effect observed by Casado and coworkers (see above).
THE CONTRIBUTION OF THE Snf1 PATHWAY TO THE RESPONSE TO ALKALINE pH
Exposure to severe alkaline stress (pH 8.0-8.2) results in rapid induction (10-15 minutes) of a substantial number of genes involved in hexose transport and carbohydrate metabolism, such as high-affinity glucose transporters and TCA cycle enzymes-encoding genes [18][22] that are normally subjected to glucose repression. This set of genes largely overlaps with that of induced when cells are shifted to low glucose-containing medium or reach the diauxic shift [74], pointing that alkaline stress may perturb glucose utilization. This notion was confirmed by the analysis of TCA cycle flux changes after switching cells to pH 7.5, which revealed a substantial increase in TCA cycle activity and respiration rate [75].
–
The protein kinase Snf1 is the yeast homolog of the mammalian AMP-activated protein kinase and it is required for adaptation of yeast cells to glucose limitation and for utilization of alternative, less preferred carbon sources. In response to glucose limitation, Snf1 is phosphorylated at Thr210 by three upstream Snf1 kinases (Sak1, Elm1 and Tos3), leading to Snf1 activation (see [76][77] for reviews).
–
In addition to glucose scarcity, Snf1 has been shown to be activated by other unfavorable conditions, such as salt or oxidative stress. Snf1 is also related to alkaline pH stress. Cells lacking Snf1 are markedly sensitive to alkaline pH [55][78][79][80][81] and exposure to high pH results in increased Snf1 phosphorylation and activation [55][82]. Increasing the amount of glucose in the medium fully abolishes the snf1 alkali-sensitive phenotype and attenuates high pH-induced Snf1 phosphorylation at Thr210 [56], suggesting that the role of Snf1 on glucose metabolism appears to be important for its function on high pH tolerance. The same authors demonstrated that most likely the positive input on Snf1 elicited by alkaline pH is generated by the Elm1 kinase, since lack of Elm1 markedly increases alkali sensitivity and this phenotype is abolished by high amounts of glucose in the medium. The sak1 tos3 double mutant showed no alkali-sensitive phenotype, but sensitivity of the tos3 elm1 sak1 triple mutant was even more pronounced than that of snf1 cells and poorly rescued by glucose supplementation, suggesting an important function of these kinases in high pH tolerance besides its regulatory role on the Snf1 kinase. The specific signal(s) activating these kinases in response to high pH stress have not been explored.
–
Transcriptomic analysis of snf1 mutants revealed that about 75% of the genes induced in the short-term by high pH are also induced by glucose scarcity, pointing again the relevance of alkaline pH stress on carbohydrate metabolism. It was found that Snf1 mediates, in full or in part, the activation of a significant subset (38%) of short-term alkali-induced genes, including those encoding high-affinity hexose transporters and phosphorylating enzymes, and glycogen (but not trehalose) metabolism [56]. As depicted in Figure 2, the expression profile of diverse genes metabolically linked to the generation of Acetyl CoA shows a tendency to favor the synthesis of this compound and to avoid its transformation in malonyl CoA. In this regard it is worth noting that Snf1 phosphorylates and inhibits the acetyl-CoA carboxylase Acc1, the enzyme that catalyzes carboxylation of cytosolic acetyl-CoA to generate malonyl-CoA. Recently, Zhang and coworkers reported [83] that decreased expression of ACC1 results in increased acetyl-CoA levels and suppresses diverse phenotypes, including alkaline pH sensitivity, in the snf1 mutant. In addition, the low levels of acetyl-CoA of snf1 mutant cells and the alkaline pH sensitivity can be suppressed by deletion of MIG1, which is a repressor of the acetyl-CoA synthetase, encoded by the ACS1 gene. Therefore, the authors conclude that some of the phenotypes and transcriptional defects of snf1 cells are due to their low acetyl-CoA level, which results in global histone hypoacetylation [83]. These findings suggest that a possible role for Snf1 in response to alkaline stress could be to promote the expression of ACS1 and to limit the activity of ACC1, thus ensuring appropriate acetyl-CoA availability. This notion fits with transcriptional data showing that ACS1 is induced by high pH stress [22][56][68] and that this effect is partially dependent on the presence of Snf1 [56], whereas the same treatment decreases ACC1 mRNA levels [14][18] (see Figure 2).
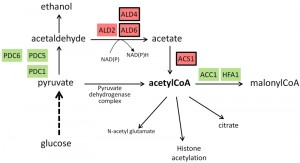 | FIGURE 2: Schematic depiction of reactions involved in generation and use of Acetyl CoA. Genes induced in response to alkaline pH stress are in red background whereas those repressed are in green background. Expression profiles have been collected from published transcriptomic data [14][18][22][56]][68][84]. Genes whose expression has been found to be dependent of Snf1 according to [56] are framed. |
Interestingly, it has been reported that many genes that are both repressed by glucose and rapidly induced by alkaline stress are also under the direct positive regulation of the calcineurin/Crz1 pathway [36]. In fact, activation of calcineurin (i.e. by addition of calcium to the medium) was sufficient to allow growth of a snf1 mutant under limiting glucose availability. Therefore, alkalinization of the medium would generate two independent signals, activation of calcineurin (through the influx of external calcium) and Snf1 (by activation of the Elm1 kinase) that, in the short-term, would represent independent but combined strategies to cope with alkaline pH stress. Very recently it has been reported that calcineurin can dephosphorylate and inactivate the Elm1 kinase during alkaline stress [84], perhaps providing a negative feedback mechanism to avoid excessive activation of the system. This feedback mechanism might explain the transient nature of Snf1 phosphorylation upon high pH stress [82].
OTHER STRESS SIGNALING PATHWAYS ACTIVATED BY ALKALINE pH STRESS
The cell wall integrity (CWI) signaling pathway is essential for growth and required for tolerance to cell wall damaging agents. Briefly, the pathway is composed of several, partially overlapping membrane sensors (Wsc1–4, Mid2, and Mtl1) which in response to cell wall stress interact with the GDP/GTP exchange factors Rom2 or Rom1 leading to activation of the small GTPase Rho1. GTP-Rho1 interacts with and activates the Pkc1, the only protein kinase C in budding yeast. In turn, Pkc1 activates a typical MAP kinase module, consisting of Bck1, the apparently redundant pair of MAPK kinases Mkk1 and Mkk2, and, finally, the Slt2 MAPK (see [85] for a review). Activation of Slt2 results in the phosphorylation of several cytosolic and nuclear targets, such as the transcription factors Rlm1 and the SCB-binding factor (Swi4 and Swi6). Strains defective for key components of this pathway display a lytic phenotype when exposed to cell wall damaging agents (i.e. Calcofluor white or Congo Red) or increased temperature, but this phenotype can be suppressed by addition of sorbitol to the medium.
–
As far as we know there is no available information about the chemical changes in cell-wall composition of yeast cells subjected to alkaline stress. However, it has been shown that the sensitivity to zymolyase (composed mainly of β-1,3-glucanase activity) of cells grown at pH 6.0 is higher than those grown at pH 4.0, while the relative proportion of β-1,6-glucan decreases around 40% [86]. In any case, a number of evidences suggest that high pH stress causes cell wall damage. The link between alkaline pH and cell wall stress was established years ago, on the basis of two independent genome-wide screens that identified bck1 and slt2 mutants as sensitive to high pH [7][8]. In fact, Gene Ontology analysis of a list of 289 mutants sensitive to high pH that we have compiled from the literature [7][8][22] shows a substantial enrichment in cells related to cell wall organization or biogenesis (7.2E-6). In addition, there is a remarkable overlap between the set of genes induced in response to severe alkaline stress [18] and those found induced after exposure to cell wall damaging agents, such as zymolyase or caspofungin, or in mutants for certain genes encoding cell wall components [87][88]. Serrano and coworkers reported that Slt2 resulted rapidly (5 min) and transiently activated after alkalinization of the medium, and this response was depended on the integrity of the MAP kinase module [22]. These authors also found that either lack of Wsc1 or specific mutations in this sensor impairing sensing or transduction of the signal, rendered cells alkali-sensitive and reduced the activation of Slt2 by high pH, whereas constitutive activation of Slt2 by the bck1-20 allele increased pH tolerance in the wsc1 mutant. These results support the notion that Wsc1 might act as sensor for high pH stress and that activation of the Slt2 MAP kinase module is required for normal high pH tolerance. The fact that Wsc1 has been identified as a cell surface mechanosensor [23] further suggests that alkaline stress causes alterations at the cell surface.
–
It has become evident in the last years that, in addition to Slt2, other signaling pathways could be relevant in the response to cell wall stress, such as the one mediated by Rim101 [61] or by the Hog1 MAP kinase [89][90]. However, whereas alkaline pH certainly activates the Rim101 pathway, this is not the case for the Hog1 pathway [22][91]. In contrast, activation of the Snf1 pathway by high pH stress might also contribute to preserve cell wall integrity, as it has recently shown that a snf1 mutant is hypersensitive to different compounds or treatments that cause cell wall stress, and that phosphorylation of Snf1 at Thr210 in response to treatment with the glucan synthase inhibitor caspofungin is important for the function of Snf1 in cell wall integrity [92]. Genetic evidence suggests that the role of Snf1 would be independent of the Slt2 cell wall integrity pathway.
–
Yeast cells exposed to high pH stress accumulate in relatively short time reactive oxygen species, suggesting they are also confronted with a certain degree of oxidative stress [18]. This explains the observation that cells lacking SOD1 or SOD2, encoding superoxide dismutases, are sensitive to high pH [8] and that a substantial number of genes that are induced in response to oxidative stress, such as PRX1, GPX2, GRX1, TRR1 or TRX2, are also induced by alkalinization of the medium [18][35]. This transcriptional response involves the transcription factors Yap1 and, probably in lesser extent, Skn7. A role for Snf1 in signaling oxidative stress has been proposed, since oxidative stress provokes a modest activation of the kinase. The observation that the snf1 mutant is not particularly sensitive to hydrogen peroxide [55][93], casts some doubts about its role in anti-oxidative defense. However, more recent work has proved that yeast cells require Snf1 (in an Elm1-dependent manner) for normal growth in the presence of diethyl maleate, diamide, or the pro-oxidant selenite, all of which cause alterations in glutathione redox homeostasis by increasing the levels of oxidized glutathione [94]. Therefore, the possibility that Snf1 could participate in the response to oxidative stress caused by alkalinization of the medium still remains open to speculation. Very recently, a protective role for Rim101 in cells treated with diverse oxidants, including diamide and selenite, has been reported by Pérez-Sampietro and coworkers [95]. This effect, similarly to the alkaline pH response, would require Rim8, the ESCRT-I/II/III complexes and the Rim13 protease involved in proteolytic activation of Rim101, as well as Nrg1, the transcriptional repressor downstream Rim101. These authors postulate that the protective effect of Rim101 in cells confronted with selenite is based on the regulatory role of Rim101 in vacuolar homeostasis and its requirement for normal vacuolar acidification. Therefore, the function of Rim101 in adaptation to alkaline stress might also include a role in confronting the associated oxidative stress.
ALKALINE STRESS AND NUTRIENT HOMEOSTASIS
In yeast cells uptake of most nutrients rely on active transport processes that are based in symport with protons and thus the majority of nutrients transporters are dependent on the proton gradient. Among others, these include some sugar transporters, all amino acids transporters, organic acid transporters, anion transporters (such as phosphate and sulfate), some monovalent (i.e. potassium) or divalent (i.e. magnesium, zinc) cations. Therefore, it can be anticipated that alkalinization of the medium would have a notorious impact on nutrient homeostasis. These effects were highlighted by early works describing the genome-wide transcriptional response to high pH stress.
–
As mentioned above, high pH stress provokes a rapid induction of genes that are normally subjected to glucose repression. These includes high-affinity glucose transporters, as well as glycolytic and TCA cycle enzymes-encoding genes [18][22][36], generating a pattern that is reminiscent of that observed when cells are shifted to low glucose-containing medium or they reach the diauxic shift [74]. However, although increasing the amount of glucose in the medium from the standard 2% up to 6% significantly improves growth at pH 8.0 (but not at pH 5.5) and normalizes high pH tolerance of the hypersensitive snf1 strain [56], alkalinization of the medium does not impairs glucose uptake [56], so it is likely that in this case the signal is generated inside the cell possibly due to the perturbation of glucose utilization. It is well known that the PKA, Snf1 and TOR pathways have key roles in nutrient sensing [96]. However, whereas the first two pathways play important roles in adaptation to alkalinization of the medium, the role of the TOR pathway is questionable, since mutants in most genes involved in the pathway, with the exception of the pleiotropic ure2 mutant, do not show a high pH-sensitive phenotype [8].
–
Shifting cells to high pH also results in an important alteration in copper and iron homeostasis. This is likely caused by the decrease in solubility of these metals in alkaline conditions and is reflected in that:
–
i) alkaline stress gives rise to increased mRNAs levels for genes encoding iron (FET3, FRE1, FET4), siderophores (ARN1, ARN2, ARN3, ARN4, FIT2) or copper (CTR1, CTR3) transporters [35][97],
–
ii) mutant strains lacking CTR1 (encoding the main high-affinity copper transporter) or AFT1 (the transcription factor required for activation of target genes in response to changes in iron availability), as well as other mutants related to copper and iron homeostasis, such as ccs1, ccc2 or fet3, are also alkali-sensitive [8], and
–
iii) only two genes, FET4 and CTR1, were able to increase high pH tolerance when overexpressed in a wild type strain [8]. Further work demonstrated that the beneficial effect of overexpression of CTR1 was largely due to the improvement in iron uptake, highlighting the tight connection between copper and iron metabolisms [98]. Addition to the medium of micromolar concentrations of these metals drastically improves growth at high pH [8].
–
Collectively, these results emphasize the fact that copper and iron availability are important limiting factors for growth under alkaline pH conditions. It should be noted that diverse genes involved in vacuolar acidification have been shown to be necessary for growth under limiting iron conditions [99]. Since an equivalent situation is triggered by high pH, this might point to a possible role of vacuole acidification in normal alkaline stress tolerance. In addition, several enzymes involved in antioxidant defenses are metalloenzymes requiring iron or copper. Thus, catalases (such as yeast Ctt1) possess iron-containing heme groups [100], whereas superoxide dismutases (such as Sod1) are copper/zinc containing enzymes. Therefore, it cannot be excluded that high pH is capable of negatively affecting cellular antioxidant defenses.
–
Expression of CTR1 and FRE1 is regulated by the Mac1 transcription factor, and FET3 is under the control of Aft1. It is worth noting that Pho85 is required for proper expression of MAC1 and AFT1 [101]. Given the various links between Pho85 and alkaline pH signaling (see above and below), the sensitivity of a pho85 strain under alkaline conditions [8][9][60] might have diverse origins. In addition, expression of FET4 has been reported to be, in part, Rim101-dependent [52].
–
Alkaline stress results in the activation of many genes involved in phosphate (Pi) acquisition or metabolism, such as PHO84, PHO89, PHO11, PHO12, VTC1, or VTC4, which are also up-regulated during Pi starvation, suggesting that alkalinization of the medium mimics a situation of Pi scarcity. This is reasonable if it is considered that low-affinity Pi transport is based on H+/Pi symport [102][103][104]. It has been postulated that increasing the pH of the medium decreases the ionic (monovalent) form of phosphate, which is the proposed uptake form for a yeast cell [105]. However, this effect could be also explained by the fact that the major high-affinity Pi transporter, Pho84, is also a H+/Pi transporter, so that at high pH its uptake capacity is far from optimal. In that case, the capacity of Pho89, able to work at alkaline pH, would be clearly lower than that of low-affinity and Pho84 transporters [106].
–
The response of PHO84 (a high-affinity H+/Pi co-transporter) and PHO12 (a secretable phosphatase) to alkaline pH was shown to be fully dependent of Pho2/Pho4 [35], the transcription factors that mediate the transcriptional responses to phosphate starvation, and of the cyclin-dependent kinase inhibitor Pho81, which was considered an intracellular phosphate sensor [107]. Alkalinization of the medium was shown to trigger degradation of the polyPs stock [108], and our results (Figure 3A) indicate that this is a very fast process. In fact, the decrease in polyP reserves induced by alkaline stress was faster than that observed upon shifting cells to phosphate limitation. This decrease was only partially blocked in cells lacking PPN1 and PPX1 (Figure 3A), the genes encoding the presumed major polyP phosphatase activities [109], thus supporting the notion that additional polyP phosphatase activities should exist. In any case, the yeast vacuole is commonly considered the major storage site for polyP, which is regarded as a cellular phosphate buffer that can be mobilized during phosphate limitation [110]. Since alkaline stress subjects the cell to a situation similar to that of phosphate starvation, and it is known that most V-ATPase mutants fail to accumulate significant levels of polyP [111], it is conceivable that maintenance of normal levels of polyP could be one of the contributions of normal vacuolar acidification to high pH stress tolerance.
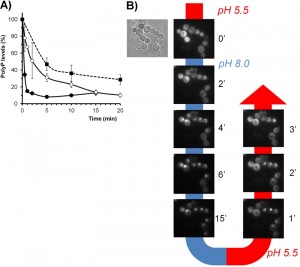 | FIGURE 3: Effects of alkaline pH stress on phosphate metabolism. (A) Depletion of intracellular PolyP stocks upon alkalinization of the medium. Strains BY4741 (wild type, closed circles) and its isogenic derivative DCS30 (ppn1 ppx1, open circles) were grown up to OD600 0.6 and subjected to alkaline stress (pH 8.0) as in [82]. For comparison, in parallel experiments the wild type strain was subjected to phosphate limitation (0.1 mM Pi) as in [112]. Samples of the culture were taken at the indicated times and polyP content determined as in [112]. Data is expressed as percentage relative to the levels just prior initiation of the stress and are mean ± SEM from 3 independent experiments. (B) Reversible cytosolic-nuclear transfer of Pho4 upon modification of the extracellular pH. Wild type strain DBY746 was transformed with plasmid Pho84-GFP, carrying a C-terminally GFP-tagged version of PHO4, and grown until OD660 0.6 in Verduyn medium [113]. Aliquots were injected into a CellASIC® ONIX Y04C-02 microfluidics chamber and the pH of the medium changed with time as indicated in the figure. Images were collected with a Nikon Eclipse TE2000-E microscope using a FITC filter and a Hamamatsu ORCA-ER camera. |
Alkalinization of the medium also provoked the fast entry of Pho4 into the nucleus, which was rapidly reversed upon acidification of the medium (Figure 3B). Induction of PHO84 and PHO12 after high pH shock is a relatively late event (45-120 min) [35][97], and the kinetics of accumulation of Pho84 upon high pH stress are very similar to that observed upon phosphate starvation [82]. All these results indicate that, in this case, the signaling pathway activated by alkaline pH is the same that is triggered by phosphate starvation. Consistently, mutation of PHO2, PHO4 or PHO81 conferred an alkali-sensitive phenotype, which was particularly dramatic for the pho84 pho89 mutant [8][9]. In contrast, induction of PHO89 is an early (10-15 min) and transient event. PHO89 encodes a Na+/Pi co-transporter that, in contrary to Pho84, is mainly active at alkaline pH [112]. In addition to regulation by Pho2/Pho4, our earlier work demonstrated that expression of PHO89 upon alkaline stress is strongly regulated by calcineurin/Crz1 [35]. The complex response of PHO89 and its physiological implications will be discussed in the next section.
INTEGRATION OF DIFFERENT SIGNALING PATHWAYS IN HIGH pH-RESPONSIVE GENE PROMOTERS
The great diversity of signaling pathways with an impact on gene expression that are modulated by alkaline pH stress likely reflects the very broad impact that environmental alkalinization has on the biology of the yeast cell. Therefore, it is not surprising that several high-pH activated pathways may impinge on a given gene promoter, thus finely tuning the response. An example is represented by genes such as HXT2, HXT7, ALD4 or ADH3, which are normally induced in response to the scarcity of glucose through activation of Snf1 and, upon alkalinization of the medium, becoming also sensitive to activation mediated by calcineurin/Crz1. This dual mechanism has physiological significance since, as mentioned above in the section “THE CONTRIBUTION OF THE Snf1 PATHWAY TO THE RESPONSE TO ALKALINE pH”, activation of calcineurin by external calcium was shown to override the growth defect of a snf1 mutant in limiting glucose conditions [36]. Similarly, a recent publication reported that under alkaline conditions the inhibition of Pho85 and the subsequent activation of Pho4, Crz1 and Rim101 contributed to the downregulation of CLN2, probably leading to an alteration of the cell cycle progression [113].
–
We have analyzed existing transcriptomic data to evaluate, at the genome-wide level, how frequent the incidence of two or more regulatory pathways is in the control of the expression of a given gene in response to high pH stress. The dataset encompasses comparisons between the profile of the wild type strain with those of msn2 msn4, snf1, cnb1/crz1, slt2 or rim101 mutants. The entire dataset shows nearly 1000 genes that are induced in at least one of the experiments at a given time-point (it must be noted that individual datasets were obtained in different experimental conditions, i.e. pH of the medium ranging from 7.6 to 8.2). As illustrated in Figure 4, induction of 349 genes depends on the presence of at least one of the factors described above, and the most prevalent regulatory components appears to be the PKA pathway, through the Msn2/Msn4 transcription factors, and the Snf1 pathway. Sixty genes (17%) appear to be multiregulated. Among them, 38 genes appear to be induced by the contribution of the Snf1 and Msn2/Msn4 pathways, and this set is enriched in genes related to glycogen metabolism and oxido-reduction processes. Eighteen genes are under the control of Snf1 and calcineurin, with significant presence of acetate biosynthetic genes, whereas 15 genes are regulated by calcineurin and Msn2/Msn4, among them several related to trehalose and β-alanine biosynthesis (Figure 4). Remarkably, up to 7 genes, all of them related to carbohydrate metabolism (GSY2, PGM2, MDH2, HXT7, GSY1, HXK1, and HXT5) are regulated by 3 different pathways: Snf1, Msn2/Msn4 and calcineurin. Finally, the genes regulated by Slt2 and Rim101 are less interconnected with the other pathways, perhaps due to the specificity of the target genes controlled by these pathways.
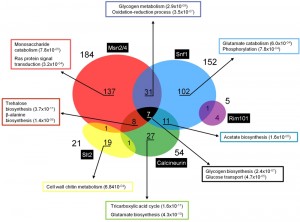 | FIGURE 4: Genes regulated by multiple signaling pathways in response to alkaline pH stress. Expression profiles have been collected from published transcriptomic data [18][22][36][56][68][84]. Gene Ontology analyses were based on FuncAssociate 2.0 [116] and YeastMine [117]. Numbers in parentheses denote the p-value for a given category. |
Regulatory inputs are particularly complex in the promoter of the ENA1 Na+-ATPase gene, a key determinant for saline stress tolerance (see [91] for a review). Under alkaline pH stress, the ENA1 promoter integrates signals from at least 3 major pathways: calcineurin (via Crz1 at two CDRE) [32], Snf1 (via both Mig1/Mig2 and Nrg1), and the Rim101 pathway, via Nrg1 [32][81]. Although the ENA1 promoter contains several putative GATA sequences [114], which would suggest it is under the control of the TOR pathway, they are likely irrelevant in response to high pH stress. Similarly, under pH stress, ENA1 does not seem to be regulated by the Msn2/Msn4 transcription factors [35], although the inhibition of the PKA pathway might led to hypophosphorylation of Crz1, thus reducing the chances for inactivation of the transcription factor (see above, section “THE PKA PATHWAY IS TRANSIENTLY INHIBITED BY ALKALINIZATION OF THE ENVIRONMENT”). Remarkably, a very recent report highlighted that, in response to high pH stress, ENA1 and PHO89 share the same regulatory network [82] and that both proteins accumulate roughly at the same time at the plasma membrane. The fact that PHO89 promoter is responsive to calcineurin, Snf1 and Rim101 explains its fast activation upon alkaline stress. In addition, it allows establishing a physiological link between Pho89 and Ena1 functions and justifies the puzzling observation that Ena1 is induced not only by salt stress (very reasonable for a Na+-ATPase) but also by alkaline pH even in the absence of salt stress. At high pH, Pho89 becomes an important contributor to Pi uptake [106]. However, this has a trade-off: the co-transport of Na+ cations. Therefore, rapid induction of ENA1 would be a safety measure not only to avoid accumulation of Na+ up to toxic levels, but also because increasing intracellular Na+ would eliminate the extracellular/intracellular cation gradient necessary to sustain uptake of Pi through Pho89. Consequently, even under mild alkaline conditions, a pho84 ena1 mutant phenocopies the extreme growth defect of a pho84 pho89 strain, a mutant devoid of high-affinity Pi transport [82]. The widespread presence of Pho89-like transporters among eukaryotes and the requirement for Ena1 to grow at alkaline pH in the absence of salt stress, observed in diverse fungi and in bryophytes (see [115] and references therein) suggest that the regulatory coupling between Pho89 and Ena1 transporters might be evolutionarily conserved.
CONCLUSIONS
Shifting yeast cells from the standard acidic environment to a moderately alkaline medium transiently alters the mRNA level of hundreds of genes. Whereas this review essentially focuses on the signaling pathways impinging on the transcriptional regulation of high pH responsive genes, it must be kept in mind that the mRNA level of a given gene is the result of two opposing forces: the activity of the promoter and the stability of the generated mRNA. The relevance of changes in mRNA stability in response to stress conditions has been recognized in the last few years and investigated in some cases, such as osmotic, oxidative or heat-shock stresses [116][117]. In collaboration with the laboratory of J.E. Pérez-Ortín (University of Valencia), our group has recently characterized with the aid of the Genomic Run-On technology (GRO) the effect of environmental alkalinization on transcription rate and mRNA stability [118]. It was found that alkalinization of the media to pH 8.0 causes an overall decrease in transcription rate and a fast destabilization of mRNAs, followed by a more prolonged stabilization phase. In some cases, as in genes responsive to oxidative stress, the increment on mRNA levels occur without a relevant increase in the transcription rate, and is probably attributable to mRNA stabilization of these genes. In comparison to other forms of stress previously analyzed such as oxidative, osmotic and heat-shock stresses, alkaline stress response is atypical because of the limited contribution of transcription rate to the increase in mRNA levels. In contrast, the decrease in the amount of mRNAs is contributed by a decline in both the transcription rate and mRNA stability. Since there is growing evidence that mRNA decay is also a subject for regulation, it seems necessary to focus future work in this direction to gain full understanding of the adaptive response to high pH stress.
–
Expanding our knowledge of the adaptive mechanisms of yeasts to alkalinization may have a relevant impact on technological or health-related applications. We have mentioned above that supplementing the medium with minimal amounts of copper and iron cations markedly increases tolerance to high pH, an effect that is also achieved by overexpression of the CTR1 and FET4 metal transporters [8]. More recently, it has been shown that expression of Kluyveromyces lactis FAD2 and FAD3 genes in S. cerevisiae leads to accumulation of linoleic and alpha-linolenic acids and results in increased alkaline pH tolerance, as well as to Li+ and Na+ cations [119]. This effect was related to the activation of the Rim101 pathway. Improving tolerance to high pH in yeast by a combination of genetic manipulation and conditioning of the growth medium could be useful for industrial applications requiring growth of yeast at pH higher than normal. On the other hand, adaptation to the wide range of extracellular pH found in the different hosts’ environments is required for pathogenic fungi for survival.
–
Interestingly, evidence has been accumulating in support of a role of several signaling pathways described in this review in drug resistance. A relationship between tolerance to antifungal drugs and calcineurin was described in C. albicans, A. fumigatus and C. neoformans, and the combination of drugs such as azoles or caspofungin with calcineurin inhibitors leads to an increased susceptibility to the drugs [120][121][122]. The Pac/Rim pathway was also found to be involved in tolerance to azoles (see [123] and references therein), and similar results were observed in C. neoformans [124]. The relevance of pH signaling in fungal pathogenesis in humans can be probably extended to fungal plant pathogens. Thus, in the rice pathogen Magnaporthe oryzae, deletion of the RIM101/PacC homolog PACC, caused a progressive loss in growth rate from pH 5 to pH 8, a loss in conidia production at pH 8 in vitro, and a partial loss in virulence towards barley and rice, suggesting that PacC plays an important role during interaction between the M. oryzae and its host plants [125].
References
- R. Serrano, M.C. Kielland-Brandt, and G.R. Fink, "Yeast plasma membrane ATPase is essential for growth and has homology with (Na+ + K+), K+- and Ca2+-ATPases.", Naturehttp://www.ncbi.nlm.nih.gov/pubmed/3005867
- G.A. Martínez-Muñoz, and P. Kane, "Vacuolar and Plasma Membrane Proton Pumps Collaborate to Achieve Cytosolic pH Homeostasis in Yeast", Journal of Biological Chemistry, vol. 283, pp. 20309-20319, 2008. http://dx.doi.org/10.1074/jbc.M710470200
- R. Serrano, "Salt tolerance in plants and microorganisms: toxicity targets and defense responses.", International review of cytology, 1996. http://www.ncbi.nlm.nih.gov/pubmed/8900956
- J. Ariño, J. Ramos, and H. Sychrová, "Alkali Metal Cation Transport and Homeostasis in Yeasts", Microbiology and Molecular Biology Reviews, vol. 74, pp. 95-120, 2010. http://dx.doi.org/10.1128/MMBR.00042-09
- W. Li, and A.P. Mitchell, "Proteolytic activation of Rim1p, a positive regulator of yeast sporulation and invasive growth.", Genetics, 1997. http://www.ncbi.nlm.nih.gov/pubmed/9017390
- M. Hayashi, K. Ohkuni, and I. Yamashita, "Control of division arrest and entry into meiosis by extracellular alkalisation in Saccharomyces cerevisiae.", Yeast (Chichester, England), 1998. http://www.ncbi.nlm.nih.gov/pubmed/9717236
- G. Giaever, A.M. Chu, L. Ni, C. Connelly, L. Riles, S. Véronneau, S. Dow, A. Lucau-Danila, K. Anderson, B. André, A.P. Arkin, A. Astromoff, M. El-Bakkoury, R. Bangham, R. Benito, S. Brachat, S. Campanaro, M. Curtiss, K. Davis, A. Deutschbauer, K. Entian, P. Flaherty, F. Foury, D.J. Garfinkel, M. Gerstein, D. Gotte, U. Güldener, J.H. Hegemann, S. Hempel, Z. Herman, D.F. Jaramillo, D.E. Kelly, S.L. Kelly, P. Kötter, D. LaBonte, D.C. Lamb, N. Lan, H. Liang, H. Liao, L. Liu, C. Luo, M. Lussier, R. Mao, P. Menard, S.L. Ooi, J.L. Revuelta, C.J. Roberts, M. Rose, P. Ross-Macdonald, B. Scherens, G. Schimmack, B. Shafer, D.D. Shoemaker, S. Sookhai-Mahadeo, R.K. Storms, J.N. Strathern, G. Valle, M. Voet, G. Volckaert, C. Wang, T.R. Ward, J. Wilhelmy, E.A. Winzeler, Y. Yang, G. Yen, E. Youngman, K. Yu, H. Bussey, J.D. Boeke, M. Snyder, P. Philippsen, R.W. Davis, and M. Johnston, "Functional profiling of the Saccharomyces cerevisiae genome.", Nature, 2002. http://www.ncbi.nlm.nih.gov/pubmed/12140549
- R. Serrano, D. Bernal, E. Simón, and J. Ariño, "Copper and iron are the limiting factors for growth of the yeast Saccharomyces cerevisiae in an alkaline environment.", The Journal of biological chemistry, 2004. http://www.ncbi.nlm.nih.gov/pubmed/14993228
- J. Ariño, "Integrative Responses to High pH Stress inS. cerevisiae", OMICS: A Journal of Integrative Biology, vol. 14, pp. 517-523, 2010. http://dx.doi.org/10.1089/omi.2010.0044
- P.M. Kane, "The where, when, and how of organelle acidification by the yeast vacuolar H+-ATPase.", Microbiology and molecular biology reviews : MMBR, 2006. http://www.ncbi.nlm.nih.gov/pubmed/16524922
- T.T. Diakov, and P.M. Kane, "Regulation of Vacuolar Proton-translocating ATPase Activity and Assembly by Extracellular pH", Journal of Biological Chemistry, vol. 285, pp. 23771-23778, 2010. http://dx.doi.org/10.1074/jbc.M110.110122
- S. Padilla-López, and D.A. Pearce, "Saccharomyces cerevisiae lacking Btn1p modulate vacuolar ATPase activity to regulate pH imbalance in the vacuole.", The Journal of biological chemistry, 2006. http://www.ncbi.nlm.nih.gov/pubmed/16423829
- S.C. Li, and P.M. Kane, "The yeast lysosome-like vacuole: Endpoint and crossroads", Biochimica et Biophysica Acta (BBA) - Molecular Cell Research, vol. 1793, pp. 650-663, 2009. http://dx.doi.org/10.1016/j.bbamcr.2008.08.003
- H.C. Causton, B. Ren, S.S. Koh, C.T. Harbison, E. Kanin, E.G. Jennings, T.I. Lee, H.L. True, E.S. Lander, and R.A. Young, "Remodeling of yeast genome expression in response to environmental changes.", Molecular biology of the cell, 2001. http://www.ncbi.nlm.nih.gov/pubmed/11179418
- D.A. Davis, "How human pathogenic fungi sense and adapt to pH: the link to virulence", Current Opinion in Microbiology, vol. 12, pp. 365-370, 2009. http://dx.doi.org/10.1016/j.mib.2009.05.006
- M.A. Peñalva, J. Tilburn, E. Bignell, and H.N. Arst, "Ambient pH gene regulation in fungi: making connections", Trends in Microbiology, vol. 16, pp. 291-300, 2008. http://dx.doi.org/10.1016/j.tim.2008.03.006
- K. Selvig, and J.A. Alspaugh, "pH Response Pathways in Fungi: Adapting to Host-derived and Environmental Signals", Mycobiology, vol. 39, pp. 249-256, 2011. http://dx.doi.org/10.5941/MYCO.2011.39.4.249
- L. Viladevall, R. Serrano, A. Ruiz, G. Domenech, J. Giraldo, A. Barceló, and J. Ariño, "Characterization of the calcium-mediated response to alkaline stress in Saccharomyces cerevisiae.", The Journal of biological chemistry, 2004. http://www.ncbi.nlm.nih.gov/pubmed/15299026
- M. Kanzaki, M. Nagasawa, I. Kojima, C. Sato, K. Naruse, M. Sokabe, and H. Iida, "Molecular identification of a eukaryotic, stretch-activated nonselective cation channel.", Science (New York, N.Y.), 1999. http://www.ncbi.nlm.nih.gov/pubmed/10436155
- D. Engelberg, R. Perlman, and A. Levitzki, "Transmembrane signaling in Saccharomyces cerevisiae as a model for signaling in metazoans: State of the art after 25years", Cellular Signalling, vol. 26, pp. 2865-2878, 2014. http://dx.doi.org/10.1016/j.cellsig.2014.09.003
- M. Paidhungat, and S. Garrett, "A homolog of mammalian, voltage-gated calcium channels mediates yeast pheromone-stimulated Ca2+ uptake and exacerbates the cdc1(Ts) growth defect.", Molecular and cellular biology, 1997. http://www.ncbi.nlm.nih.gov/pubmed/9343395
- R. Serrano, H. Martín, A. Casamayor, and J. Ariño, "Signaling alkaline pH stress in the yeast Saccharomyces cerevisiae through the Wsc1 cell surface sensor and the Slt2 MAPK pathway.", The Journal of biological chemistry, 2006. http://www.ncbi.nlm.nih.gov/pubmed/17088254
- V. Dupres, D. Alsteens, S. Wilk, B. Hansen, J.J. Heinisch, and Y.F. Dufrêne, "The yeast Wsc1 cell surface sensor behaves like a nanospring in vivo", Nature Chemical Biology, vol. 5, pp. 857-862, 2009. http://dx.doi.org/10.1038/nchembio.220
- M.S. Cyert, "Calcineurin signaling in Saccharomyces cerevisiae: how yeast go crazy in response to stress.", Biochemical and biophysical research communications, 2003. http://www.ncbi.nlm.nih.gov/pubmed/14623300
- T. Nakamura, Y. Liu, D. Hirata, H. Namba, S. Harada, T. Hirokawa, and T. Miyakawa, "Protein phosphatase type 2B (calcineurin)-mediated, FK506-sensitive regulation of intracellular ions in yeast is an important determinant for adaptation to high salt stress conditions.", The EMBO journal, 1993. http://www.ncbi.nlm.nih.gov/pubmed/7693452
- I. Mendoza, F. Rubio, A. Rodriguez-Navarro, and J.M. Pardo, "The protein phosphatase calcineurin is essential for NaCl tolerance of Saccharomyces cerevisiae.", The Journal of biological chemistry, 1994. http://www.ncbi.nlm.nih.gov/pubmed/8132612
- A.L. Kullas, S.J. Martin, and D. Davis, "Adaptation to environmental pH: integrating the Rim101 and calcineurin signal transduction pathways.", Molecular microbiology, 2007. http://www.ncbi.nlm.nih.gov/pubmed/17927701
- A. Odom, S. Muir, E. Lim, D.L. Toffaletti, J. Perfect, and J. Heitman, "Calcineurin is required for virulence of Cryptococcus neoformans.", The EMBO journal, 1997. http://www.ncbi.nlm.nih.gov/pubmed/9184205
- A.M. Stathopoulos, and M.S. Cyert, "Calcineurin acts through the CRZ1/TCN1-encoded transcription factor to regulate gene expression in yeast.", Genes & development, 1997. http://www.ncbi.nlm.nih.gov/pubmed/9407035
- I. Mendizabal, G. Rios, J.M. Mulet, R. Serrano, and I.F. de Larrinoa, "Yeast putative transcription factors involved in salt tolerance.", FEBS letters, 1998. http://www.ncbi.nlm.nih.gov/pubmed/9559673
- D.P. Matheos, T.J. Kingsbury, U.S. Ahsan, and K.W. Cunningham, "Tcn1p/Crz1p, a calcineurin-dependent transcription factor that differentially regulates gene expression in Saccharomyces cerevisiae.", Genes & development, 1997. http://www.ncbi.nlm.nih.gov/pubmed/9407036
- I. Mendizabal, A. Pascual-Ahuir, R. Serrano, and I.F. de Larrinoa, "Promoter sequences regulated by the calcineurin-activated transcription factor Crz1 in the yeast ENA1 gene.", Molecular genetics and genomics : MGG, 2001. http://www.ncbi.nlm.nih.gov/pubmed/11523797
- H. Yoshimoto, K. Saltsman, A.P. Gasch, H.X. Li, N. Ogawa, D. Botstein, P.O. Brown, and M.S. Cyert, "Genome-wide analysis of gene expression regulated by the calcineurin/Crz1p signaling pathway in Saccharomyces cerevisiae.", The Journal of biological chemistry, 2002. http://www.ncbi.nlm.nih.gov/pubmed/12058033
- S. Thewes, "Calcineurin-Crz1 Signaling in Lower Eukaryotes", Eukaryotic Cell, vol. 13, pp. 694-705, 2014. http://dx.doi.org/10.1128/EC.00038-14
- R. Serrano, A. Ruiz, D. Bernal, J.R. Chambers, and J. Ariño, "The transcriptional response to alkaline pH in Saccharomyces cerevisiae: evidence for calcium-mediated signalling.", Molecular microbiology, 2002. http://www.ncbi.nlm.nih.gov/pubmed/12453218
- A. Ruiz, R. Serrano, and J. Ariño, "Direct Regulation of Genes Involved in Glucose Utilization by the Calcium/Calcineurin Pathway", Journal of Biological Chemistry, vol. 283, pp. 13923-13933, 2008. http://dx.doi.org/10.1074/jbc.M708683200
- K.A. Kafadar, H. Zhu, M. Snyder, and M.S. Cyert, "Negative regulation of calcineurin signaling by Hrr25p, a yeast homolog of casein kinase I.", Genes & development, 2003. http://www.ncbi.nlm.nih.gov/pubmed/14597664
- R. Sopko, D. Huang, N. Preston, G. Chua, B. Papp, K. Kafadar, M. Snyder, S.G. Oliver, M. Cyert, T.R. Hughes, C. Boone, and B. Andrews, "Mapping pathways and phenotypes by systematic gene overexpression.", Molecular cell, 2006. http://www.ncbi.nlm.nih.gov/pubmed/16455487
- K.A. Kafadar, and M.S. Cyert, "Integration of stress responses: modulation of calcineurin signaling in Saccharomyces cerevisiae by protein kinase A.", Eukaryotic cell, 2004. http://www.ncbi.nlm.nih.gov/pubmed/15470242
- V.L. Heath, S.L. Shaw, S. Roy, and M.S. Cyert, "Hph1p and Hph2p, novel components of calcineurin-mediated stress responses in Saccharomyces cerevisiae.", Eukaryotic cell, 2004. http://www.ncbi.nlm.nih.gov/pubmed/15189990
- F.J. Piña, A.F. O'Donnell, S. Pagant, H.L. Piao, J.P. Miller, S. Fields, E.A. Miller, and M.S. Cyert, "Hph1 and Hph2 Are Novel Components of the Sec63/Sec62 Posttranslational Translocation Complex That Aid in Vacuolar Proton ATPase Biogenesis", Eukaryotic Cell, vol. 10, pp. 63-71, 2011. http://dx.doi.org/10.1128/EC.00241-10
- C. Forster, and P.M. Kane, "Cytosolic Ca2+ homeostasis is a constitutive function of the V-ATPase in Saccharomyces cerevisiae.", The Journal of biological chemistry, 2000. http://www.ncbi.nlm.nih.gov/pubmed/10991947
- S.S. Su, and A.P. Mitchell, "Identification of functionally related genes that stimulate early meiotic gene expression in yeast.", Genetics, 1993. http://www.ncbi.nlm.nih.gov/pubmed/8417990
- B. Tréton, S. Blanchin-Roland, M. Lambert, A. Lépingle, and C. Gaillardin, "Ambient pH signalling in ascomycetous yeasts involves homologues of the Aspergillus nidulans genes palF and paIH.", Molecular & general genetics : MGG, 2000. http://www.ncbi.nlm.nih.gov/pubmed/10821185
- S. Piccirillo, M.G. White, J.C. Murphy, D.J. Law, and S.M. Honigberg, "The Rim101p/PacC Pathway and Alkaline pH Regulate Pattern Formation in Yeast Colonies", Genetics, vol. 184, pp. 707-716, 2010. http://dx.doi.org/10.1534/genetics.109.113480
- K. Obara, H. Yamamoto, and A. Kihara, "Membrane Protein Rim21 Plays a Central Role in Sensing Ambient pH in Saccharomyces cerevisiae", Journal of Biological Chemistry, vol. 287, pp. 38473-38481, 2012. http://dx.doi.org/10.1074/jbc.M112.394205
- T. Maeda, "The signaling mechanism of ambient pH sensing and adaptation in yeast and fungi", The FEBS Journal, vol. 279, pp. 1407-1413, 2012. http://dx.doi.org/10.1111/j.1742-4658.2012.08548.x
- K. Obara, and A. Kihara, "Signaling Events of the Rim101 Pathway Occur at the Plasma Membrane in a Ubiquitination-Dependent Manner", Molecular and Cellular Biology, vol. 34, pp. 3525-3534, 2014. http://dx.doi.org/10.1128/MCB.00408-14
- A. Galindo, A.M. Calcagno-Pizarelli, H.N. Arst, and M.�. Peñalva, "An ordered pathway for the assembly of ESCRT-containing fungal ambient pH signalling complexes at the plasma membrane", Journal of Cell Science, 2012. http://dx.doi.org/10.1242/jcs.098897
- A. Herrador, S. Herranz, D. Lara, and O. Vincent, "Recruitment of the ESCRT Machinery to a Putative Seven-Transmembrane-Domain Receptor Is Mediated by an Arrestin-Related Protein", Molecular and Cellular Biology, vol. 30, pp. 897-907, 2010. http://dx.doi.org/10.1128/MCB.00132-09
- M.A. Peñalva, D. Lucena-Agell, and H.N. Arst, "Liaison alcaline: Pals entice non-endosomal ESCRTs to the plasma membrane for pH signaling", Current Opinion in Microbiology, vol. 22, pp. 49-59, 2014. http://dx.doi.org/10.1016/j.mib.2014.09.005
- T.M. Lamb, and A.P. Mitchell, "The transcription factor Rim101p governs ion tolerance and cell differentiation by direct repression of the regulatory genes NRG1 and SMP1 in Saccharomyces cerevisiae.", Molecular and cellular biology, 2003. http://www.ncbi.nlm.nih.gov/pubmed/12509465
- V.K. Vyas, S. Kuchin, and M. Carlson, "Interaction of the repressors Nrg1 and Nrg2 with the Snf1 protein kinase in Saccharomyces cerevisiae.", Genetics, 2001. http://www.ncbi.nlm.nih.gov/pubmed/11404322
- S. Kuchin, V.K. Vyas, and M. Carlson, "Snf1 protein kinase and the repressors Nrg1 and Nrg2 regulate FLO11, haploid invasive growth, and diploid pseudohyphal differentiation.", Molecular and cellular biology, 2002. http://www.ncbi.nlm.nih.gov/pubmed/12024013
- S. Hong, and M. Carlson, "Regulation of snf1 protein kinase in response to environmental stress.", The Journal of biological chemistry, 2007. http://www.ncbi.nlm.nih.gov/pubmed/17438333
- A. Casamayor, R. Serrano, M. Platara, C. Casado, A. Ruiz, and J. Ariño, "The role of the Snf1 kinase in the adaptive response of Saccharomyces cerevisiae to alkaline pH stress", Biochemical Journal, vol. 444, pp. 39-49, 2012. http://dx.doi.org/10.1042/BJ20112099
- N.P. Mira, A.B. Lourenço, A.R. Fernandes, J.D. Becker, and I. Sá-Correia, "The RIM101 pathway has a role inSaccharomyces cerevisiaeadaptive response and resistance to propionic acid and other weak acids", FEMS Yeast Research, vol. 9, pp. 202-216, 2009. http://dx.doi.org/10.1111/j.1567-1364.2008.00473.x
- N.P. Mira, M. Palma, J.F. Guerreiro, and I. Sá-Correia, "Genome-wide identification of Saccharomyces cerevisiae genes required for tolerance to acetic acid", Microbial Cell Factories, vol. 9, 2010. http://dx.doi.org/10.1186/1475-2859-9-79
- N.P. Mira, J.D. Becker, and I. Sá-Correia, "Genomic Expression Program Involving the Haa1p-Regulon inSaccharomyces cerevisiaeResponse to Acetic Acid", OMICS: A Journal of Integrative Biology, vol. 14, pp. 587-601, 2010. http://dx.doi.org/10.1089/omi.2010.0048
- M. Nishizawa, M. Tanigawa, M. Hayashi, T. Maeda, Y. Yazaki, Y. Saeki, and A. Toh-e, "Pho85 Kinase, a Cyclin-Dependent Kinase, Regulates Nuclear Accumulation of the Rim101 Transcription Factor in the Stress Response of Saccharomyces cerevisiae", Eukaryotic Cell, vol. 9, pp. 943-951, 2010. http://dx.doi.org/10.1128/EC.00247-09
- F. Castrejon, A. Gomez, M. Sanz, A. Duran, and C. Roncero, "The RIM101 pathway contributes to yeast cell wall assembly and its function becomes essential in the absence of mitogen-activated protein kinase Slt2p.", Eukaryotic cell, 2006. http://www.ncbi.nlm.nih.gov/pubmed/16524906
- C.J. Nobile, N. Solis, C.L. Myers, A.J. Fay, J. Deneault, A. Nantel, A.P. Mitchell, and S.G. Filler, "Candida albicanstranscription factor Rim101 mediates pathogenic interactions through cell wall functions", Cellular Microbiology, vol. 10, pp. 2180-2196, 2008. http://dx.doi.org/10.1111/j.1462-5822.2008.01198.x
- T.R. O'Meara, S.M. Holmer, K. Selvig, F. Dietrich, and J.A. Alspaugh, "Cryptococcus neoformans Rim101 Is Associated with Cell Wall Remodeling and Evasion of the Host Immune Responses", mBio, vol. 4, 2013. http://dx.doi.org/10.1128/mBio.00522-12
- T.R. O'Meara, W. Xu, K.M. Selvig, M.J. O'Meara, A.P. Mitchell, and J.A. Alspaugh, "The Cryptococcus neoformans Rim101 Transcription Factor Directly Regulates Genes Required for Adaptation to the Host", Molecular and Cellular Biology, vol. 34, pp. 673-684, 2014. http://dx.doi.org/10.1128/MCB.01359-13
- B. Smets, R. Ghillebert, P. De Snijder, M. Binda, E. Swinnen, C. De Virgilio, and J. Winderickx, "Life in the midst of scarcity: adaptations to nutrient availability in Saccharomyces cerevisiae", Current Genetics, vol. 56, pp. 1-32, 2010. http://dx.doi.org/10.1007/s00294-009-0287-1
- L. Kraakman, K. Lemaire, P. Ma, A.W. Teunissen, M.C. Donaton, P. Van Dijck, J. Winderickx, J.H. de Winde, and J.M. Thevelein, "A Saccharomyces cerevisiae G-protein coupled receptor, Gpr1, is specifically required for glucose activation of the cAMP pathway during the transition to growth on glucose.", Molecular microbiology, 1999. http://www.ncbi.nlm.nih.gov/pubmed/10361302
- S. Colombo, P. Ma, L. Cauwenberg, J. Winderickx, M. Crauwels, A. Teunissen, D. Nauwelaers, J.H. de Winde, M.F. Gorwa, D. Colavizza, and J.M. Thevelein, "Involvement of distinct G-proteins, Gpa2 and Ras, in glucose- and intracellular acidification-induced cAMP signalling in the yeast Saccharomyces cerevisiae.", The EMBO journal, 1998. http://www.ncbi.nlm.nih.gov/pubmed/9628870
- C. Casado, A. González, M. Platara, A. Ruiz, and J. Ariño, "The role of the protein kinase A pathway in the response to alkaline pH stress in yeast", Biochemical Journal, vol. 438, pp. 523-533, 2011. http://dx.doi.org/10.1042/BJ20110607
- W. Görner, E. Durchschlag, M.T. Martinez-Pastor, F. Estruch, G. Ammerer, B. Hamilton, H. Ruis, and C. Schüller, "Nuclear localization of the C2H2 zinc finger protein Msn2p is regulated by stress and protein kinase A activity.", Genes & development, 1998. http://www.ncbi.nlm.nih.gov/pubmed/9472026
- J.M. Thevelein, and J.H. de Winde, "Novel sensing mechanisms and targets for the cAMP-protein kinase A pathway in the yeast Saccharomyces cerevisiae.", Molecular microbiology, 1999. http://www.ncbi.nlm.nih.gov/pubmed/10476026
- S. Broggi, E. Martegani, and S. Colombo, "Nuclear Ras2-GTP Controls Invasive Growth in Saccharomyces cerevisiae", PLoS ONE, vol. 8, pp. e79274, 2013. http://dx.doi.org/10.1371/journal.pone.0079274
- S. Colombo, D. Ronchetti, J.M. Thevelein, J. Winderickx, and E. Martegani, "Activation state of the Ras2 protein and glucose-induced signaling in Saccharomyces cerevisiae.", The Journal of biological chemistry, 2004. http://www.ncbi.nlm.nih.gov/pubmed/15339905
- Y. Takatsume, T. Ohdate, K. Maeta, W. Nomura, S. Izawa, and Y. Inoue, "Calcineurin/Crz1 destabilizes Msn2 and Msn4 in the nucleus in response to Ca2+ in Saccharomyces cerevisiae", Biochemical Journal, vol. 427, pp. 275-287, 2010. http://dx.doi.org/10.1042/BJ20091334
- J.L. DeRisi, V.R. Iyer, and P.O. Brown, "Exploring the metabolic and genetic control of gene expression on a genomic scale.", Science (New York, N.Y.), 1997. http://www.ncbi.nlm.nih.gov/pubmed/9381177
- J. Heyland, J. Fu, and L.M. Blank, "Correlation between TCA cycle flux and glucose uptake rate during respiro-fermentative growth of Saccharomyces cerevisiae", Microbiology, vol. 155, pp. 3827-3837, 2009. http://dx.doi.org/10.1099/mic.0.030213-0
- K. Hedbacker, and M. Carlson, "SNF1/AMPK pathways in yeast.", Frontiers in bioscience : a journal and virtual library, 2008. http://www.ncbi.nlm.nih.gov/pubmed/17981722
- R. Ghillebert, E. Swinnen, J. Wen, L. Vandesteene, M. Ramon, K. Norga, F. Rolland, and J. Winderickx, "The AMPK/SNF1/SnRK1 fuel gauge and energy regulator: structure, function and regulation", The FEBS Journal, vol. 278, pp. 3978-3990, 2011. http://dx.doi.org/10.1111/j.1742-4658.2011.08315.x
- P.M. Alepuz, K.W. Cunningham, and F. Estruch, "Glucose repression affects ion homeostasis in yeast through the regulation of the stress-activated ENA1 gene.", Molecular microbiology, 1997. http://www.ncbi.nlm.nih.gov/pubmed/9383192
- F. Portillo, J.M. Mulet, and R. Serrano, "A role for the non-phosphorylated form of yeast Snf1: tolerance to toxic cations and activation of potassium transport.", FEBS letters, 2005. http://www.ncbi.nlm.nih.gov/pubmed/15642368
- T. Ye, R. García-Salcedo, J. Ramos, and S. Hohmann, "Gis4, a new component of the ion homeostasis system in the yeast Saccharomyces cerevisiae.", Eukaryotic cell, 2006. http://www.ncbi.nlm.nih.gov/pubmed/17030993
- M. Platara, A. Ruiz, R. Serrano, A. Palomino, F. Moreno, and J. Ariño, "The transcriptional response of the yeast Na(+)-ATPase ENA1 gene to alkaline stress involves three main signaling pathways.", The Journal of biological chemistry, 2006. http://www.ncbi.nlm.nih.gov/pubmed/17023428
- A. Serra-Cardona, S. Petrezsélyová, D. Canadell, J. Ramos, and J. Ariño, "Coregulated Expression of the Na+/Phosphate Pho89 Transporter and Ena1 Na+-ATPase Allows Their Functional Coupling under High-pH Stress", Molecular and Cellular Biology, vol. 34, pp. 4420-4435, 2014. http://dx.doi.org/10.1128/MCB.01089-14
- M. Zhang, L. Galdieri, and A. Vancura, "The Yeast AMPK Homolog SNF1 Regulates Acetyl Coenzyme A Homeostasis and Histone Acetylation", Molecular and Cellular Biology, vol. 33, pp. 4701-4717, 2013. http://dx.doi.org/10.1128/MCB.00198-13
- A. Goldman, J. Roy, B. Bodenmiller, S. Wanka, C. Landry, R. Aebersold, and M. Cyert, "The Calcineurin Signaling Network Evolves via Conserved Kinase-Phosphatase Modules that Transcend Substrate Identity", Molecular Cell, vol. 55, pp. 422-435, 2014. http://dx.doi.org/10.1016/j.molcel.2014.05.012
- D.E. Levin, "Regulation of Cell Wall Biogenesis in Saccharomyces cerevisiae: The Cell Wall Integrity Signaling Pathway", Genetics, vol. 189, pp. 1145-1175, 2011. http://dx.doi.org/10.1534/genetics.111.128264
- B. Aguilar-Uscanga, and J.M. François, "A study of the yeast cell wall composition and structure in response to growth conditions and mode of cultivation.", Letters in applied microbiology, 2003. http://www.ncbi.nlm.nih.gov/pubmed/12904232
- R. García, C. Bermejo, C. Grau, R. Pérez, J.M. Rodríguez-Peña, J. Francois, C. Nombela, and J. Arroyo, "The global transcriptional response to transient cell wall damage in Saccharomyces cerevisiae and its regulation by the cell integrity signaling pathway.", The Journal of biological chemistry, 2004. http://www.ncbi.nlm.nih.gov/pubmed/14739279
- A. Lagorce, N.C. Hauser, D. Labourdette, C. Rodriguez, H. Martin-Yken, J. Arroyo, J.D. Hoheisel, and J. François, "Genome-wide analysis of the response to cell wall mutations in the yeast Saccharomyces cerevisiae.", The Journal of biological chemistry, 2003. http://www.ncbi.nlm.nih.gov/pubmed/12644457
- C. Bermejo, E. Rodríguez, R. García, J.M. Rodríguez-Peña, M.L. Rodríguez de la Concepción, C. Rivas, P. Arias, C. Nombela, F. Posas, and J. Arroyo, "The Sequential Activation of the Yeast HOG and SLT2 Pathways Is Required for Cell Survival to Cell Wall Stress", Molecular Biology of the Cell, vol. 19, pp. 1113-1124, 2008. http://dx.doi.org/10.1091/mbc.E07-08-0742
- R. García, J.M. Rodríguez-Peña, C. Bermejo, C. Nombela, and J. Arroyo, "The High Osmotic Response and Cell Wall Integrity Pathways Cooperate to Regulate Transcriptional Responses to Zymolyase-induced Cell Wall Stress in Saccharomyces cerevisiae", Journal of Biological Chemistry, vol. 284, pp. 10901-10911, 2009. http://dx.doi.org/10.1074/jbc.M808693200
- A. Ruiz, and J. Ariño, "Function and regulation of the Saccharomyces cerevisiae ENA sodium ATPase system.", Eukaryotic cell, 2007. http://www.ncbi.nlm.nih.gov/pubmed/17951516
- K. Backhaus, D. Rippert, C.J. Heilmann, A.G. Sorgo, C.G. de Koster, F.M. Klis, R. Rodicio, and J.J. Heinisch, "Mutations in SNF1 complex genes affect yeast cell wall strength", European Journal of Cell Biology, vol. 92, pp. 383-395, 2013. http://dx.doi.org/10.1016/j.ejcb.2014.01.001
- C. Dubacq, A. Chevalier, and C. Mann, "The protein kinase Snf1 is required for tolerance to the ribonucleotide reductase inhibitor hydroxyurea.", Molecular and cellular biology, 2004. http://www.ncbi.nlm.nih.gov/pubmed/14993292
- M. Pérez-Sampietro, C. Casas, and E. Herrero, "The AMPK Family Member Snf1 Protects Saccharomyces cerevisiae Cells upon Glutathione Oxidation", PLoS ONE, vol. 8, pp. e58283, 2013. http://dx.doi.org/10.1371/journal.pone.0058283
- M. Pérez-Sampietro, and E. Herrero, "The PacC-family protein Rim101 prevents selenite toxicity in Saccharomyces cerevisiae by controlling vacuolar acidification", Fungal Genetics and Biology, vol. 71, pp. 76-85, 2014. http://dx.doi.org/10.1016/j.fgb.2014.09.001
- M. Conrad, J. Schothorst, H.N. Kankipati, G. Van Zeebroeck, M. Rubio-Texeira, and J.M. Thevelein, "Nutrient sensing and signaling in the yeastSaccharomyces cerevisiae", FEMS Microbiology Reviews, vol. 38, pp. 254-299, 2014. http://dx.doi.org/10.1111/1574-6976.12065
- T.M. Lamb, W. Xu, A. Diamond, and A.P. Mitchell, "Alkaline Response Genes of Saccharomyces cerevisiaeand Their Relationship to the RIM101 Pathway", Journal of Biological Chemistry, vol. 276, pp. 1850-1856, 2001. http://dx.doi.org/10.1074/jbc.M008381200
- N.G. De Luca, and P.M. Wood, "Iron uptake by fungi: contrasted mechanisms with internal or external reduction.", Advances in microbial physiology, 2000. http://www.ncbi.nlm.nih.gov/pubmed/10907554
- S.R. Davis-Kaplan, D.M. Ward, S.L. Shiflett, and J. Kaplan, "Genome-wide analysis of iron-dependent growth reveals a novel yeast gene required for vacuolar acidification.", The Journal of biological chemistry, 2003. http://www.ncbi.nlm.nih.gov/pubmed/14594803
- W. Hansberg, R. Salas-Lizana, and L. Domínguez, "Fungal catalases: Function, phylogenetic origin and structure", Archives of Biochemistry and Biophysics, vol. 525, pp. 170-180, 2012. http://dx.doi.org/10.1016/j.abb.2012.05.014
- M. Nishizawa, Y. Katou, K. Shirahige, and A. Toh-e, "Yeast Pho85 kinase is required for proper gene expression during the diauxic shift.", Yeast (Chichester, England), 2004. http://www.ncbi.nlm.nih.gov/pubmed/15334555
- M. Bun-ya, K. Shikata, S. Nakade, C. Yompakdee, S. Harashima, and Y. Oshima, "Two new genes, PHO86 and PHO87, involved in inorganic phosphate uptake in Saccharomyces cerevisiae.", Current genetics, 1996. http://www.ncbi.nlm.nih.gov/pubmed/8598055
- R. Ghillebert, E. Swinnen, P. De Snijder, B. Smets, and J. Winderickx, "Differential roles for the low-affinity phosphate transporters Pho87 and Pho90 inSaccharomyces cerevisiae", Biochemical Journal, vol. 434, pp. 243-251, 2011. http://dx.doi.org/10.1042/BJ20101118
- D.D. Wykoff, and E.K. O'Shea, "Phosphate transport and sensing in Saccharomyces cerevisiae.", Genetics, 2001. http://www.ncbi.nlm.nih.gov/pubmed/11779791
- J. GOODMAN, and A. ROTHSTEIN, "The active transport of phosphate into the yeast cell.", The Journal of general physiology, 1957. http://www.ncbi.nlm.nih.gov/pubmed/13439168
- R.A. Zvyagilskaya, F. Lundh, D. Samyn, J. Pattison-Granberg, J. Mouillon, Y. Popova, J.M. Thevelein, and B.L. Persson, "Characterization of the Pho89 phosphate transporter by functional hyperexpression inSaccharomyces cerevisiae", FEMS Yeast Research, vol. 8, pp. 685-696, 2008. http://dx.doi.org/10.1111/j.1567-1364.2008.00408.x
- J. Mouillon, and B.L. Persson, "New aspects on phosphate sensing and signalling in Saccharomyces cerevisiae.", FEMS yeast research, 2006. http://www.ncbi.nlm.nih.gov/pubmed/16487340
- C.D. Castro, A.J. Meehan, A.P. Koretsky, and M.M. Domach, "In situ 31P nuclear magnetic resonance for observation of polyphosphate and catabolite responses of chemostat-cultivated Saccharomyces cerevisiae after alkalinization.", Applied and environmental microbiology, 1995. http://www.ncbi.nlm.nih.gov/pubmed/8534109
- L. Lichko, T. Kulakovskaya, N. Pestov, and I. Kulaev, "Inorganic polyphosphates and exopolyphosphatases in cell compartments of the yeast Saccharomyces cerevisiae under inactivation of PPX1 and PPN1 genes.", Bioscience reports, 2006. http://www.ncbi.nlm.nih.gov/pubmed/16779667
- M.R. Thomas, and E.K. O'Shea, "An intracellular phosphate buffer filters transient fluctuations in extracellular phosphate levels.", Proceedings of the National Academy of Sciences of the United States of America, 2005. http://www.ncbi.nlm.nih.gov/pubmed/15972809
- F.M. Freimoser, H.C. Hürlimann, C.A. Jakob, T.P. Werner, and N. Amrhein, "Systematic screening of polyphosphate (poly P) levels in yeast mutant cells reveals strong interdependence with primary metabolism.", Genome biology, 2006. http://www.ncbi.nlm.nih.gov/pubmed/17107617
- B.L. Persson, J. Petersson, U. Fristedt, R. Weinander, A. Berhe, and J. Pattison, "Phosphate permeases of Saccharomyces cerevisiae: structure, function and regulation.", Biochimica et biophysica acta, 1999. http://www.ncbi.nlm.nih.gov/pubmed/10548719
- M. Nishizawa, "The regulators of yeast PHO system participate in the transcriptional regulation of G1 cyclin under alkaline stress conditions.", Yeast (Chichester, England), 2015. http://www.ncbi.nlm.nih.gov/pubmed/25582350
- J.L. Crespo, K. Daicho, T. Ushimaru, and M.N. Hall, "The GATA transcription factors GLN3 and GAT1 link TOR to salt stress in Saccharomyces cerevisiae.", The Journal of biological chemistry, 2001. http://www.ncbi.nlm.nih.gov/pubmed/11457832
- A. Rodríguez-Navarro, and B. Benito, "Sodium or potassium efflux ATPase", Biochimica et Biophysica Acta (BBA) - Biomembranes, vol. 1798, pp. 1841-1853, 2010. http://dx.doi.org/10.1016/j.bbamem.2010.07.009
- J.E. Pérez-Ortín, L. de Miguel-Jiménez, and S. Chávez, "Genome-wide studies of mRNA synthesis and degradation in eukaryotes", Biochimica et Biophysica Acta (BBA) - Gene Regulatory Mechanisms, vol. 1819, pp. 604-615, 2012. http://dx.doi.org/10.1016/j.bbagrm.2011.12.002
- J.E. Pérez-Ortín, P. Alepuz, S. Chávez, and M. Choder, "Eukaryotic mRNA Decay: Methodologies, Pathways, and Links to Other Stages of Gene Expression", Journal of Molecular Biology, vol. 425, pp. 3750-3775, 2013. http://dx.doi.org/10.1016/j.jmb.2013.02.029
- D. Canadell, J. García-Martínez, P. Alepuz, J.E. Pérez-Ortín, and J. Ariño, "Impact of high pH stress on yeast gene expression: A comprehensive analysis of mRNA turnover during stress responses", Biochimica et Biophysica Acta (BBA) - Gene Regulatory Mechanisms, vol. 1849, pp. 653-664, 2015. http://dx.doi.org/10.1016/j.bbagrm.2015.04.001
- H. Yazawa, H. Iwahashi, Y. Kamisaka, K. Kimura, and H. Uemura, "Production of polyunsaturated fatty acids in yeast Saccharomyces cerevisiae and its relation to alkaline pH tolerance", Yeast, vol. 26, pp. 167-184, 2009. http://dx.doi.org/10.1002/yea.1659
- W.J. Steinbach, R.A. Cramer, B.Z. Perfect, C. Henn, K. Nielsen, J. Heitman, and J.R. Perfect, "Calcineurin inhibition or mutation enhances cell wall inhibitors against Aspergillus fumigatus.", Antimicrobial agents and chemotherapy, 2007. http://www.ncbi.nlm.nih.gov/pubmed/17502415
- D. Sanglard, F. Ischer, O. Marchetti, J. Entenza, and J. Bille, "Calcineurin A of Candida albicans: involvement in antifungal tolerance, cell morphogenesis and virulence.", Molecular microbiology, 2003. http://www.ncbi.nlm.nih.gov/pubmed/12753189
- M. Del Poeta, M.C. Cruz, M.E. Cardenas, J.R. Perfect, and J. Heitman, "Synergistic antifungal activities of bafilomycin A(1), fluconazole, and the pneumocandin MK-0991/caspofungin acetate (L-743,873) with calcineurin inhibitors FK506 and L-685,818 against Cryptococcus neoformans.", Antimicrobial agents and chemotherapy, 2000. http://www.ncbi.nlm.nih.gov/pubmed/10681348
- M. Cornet, and C. Gaillardin, "pH Signaling in Human Fungal Pathogens: a New Target for Antifungal Strategies", Eukaryotic Cell, vol. 13, pp. 342-352, 2014. http://dx.doi.org/10.1128/EC.00313-13
- K. Jung, A.K. Strain, K. Nielsen, K. Jung, and Y. Bahn, "Two cation transporters Ena1 and Nha1 cooperatively modulate ion homeostasis, antifungal drug resistance, and virulence of Cryptococcus neoformans via the HOG pathway", Fungal Genetics and Biology, vol. 49, pp. 332-345, 2012. http://dx.doi.org/10.1016/j.fgb.2012.02.001
- P. Landraud, S. Chuzeville, G. Billon-Grande, N. Poussereau, and C. Bruel, "Adaptation to pH and Role of PacC in the Rice Blast Fungus Magnaporthe oryzae", PLoS ONE, vol. 8, pp. e69236, 2013. http://dx.doi.org/10.1371/journal.pone.0069236
ACKNOWLEDGMENTS
The authors wish to thank former members of the Yeast Molecular Biology research group, Raquel Serrano, Laia Viladevall, Amparo Ruiz, Maria Platara, Asier González and Carlos Casado, who made many relevant contributions to these studies. Thanks are also given to M.A. Peñalva for critical reading of part of the manuscript and to J. Clotet (Universitat Internacional de Catalunya) for the Pho4-GFP plasmid. Recent work in the authors’ laboratory was funded by grants EUI2009-04147 (ERA-NET SysMo2), BFU2011-30197-C3-01 and BFU2014-54591-C2-1-P (Ministerio de Economia y Competitividad, Spain, and Fondo Europeo de Desarrollo Regional (FEDER)), and Ajut 2014SGR-4 (Generalitat de Catalunya) to JA.
J.A. was the recipient of an Institució Catalana de Recerca i Estudis Avançats (ICREA) Academia 2009 Award (Generalitat de Catalunya). D.C. was recipient of a pre-doctoral fellowship (FPU) from the Spanish Ministry of Education. A.S-C. was recipient of a PIF Fellowship (Universitat Autònoma de Barcelona).
COPYRIGHT
© 2015

Coordinate responses to alkaline pH stress in budding yeast by Albert Serra-Cardona et al. is licensed under a Creative Commons Attribution 4.0 International License.









