Back to article: The interplay between transcription and mRNA degradation in Saccharomyces cerevisiae
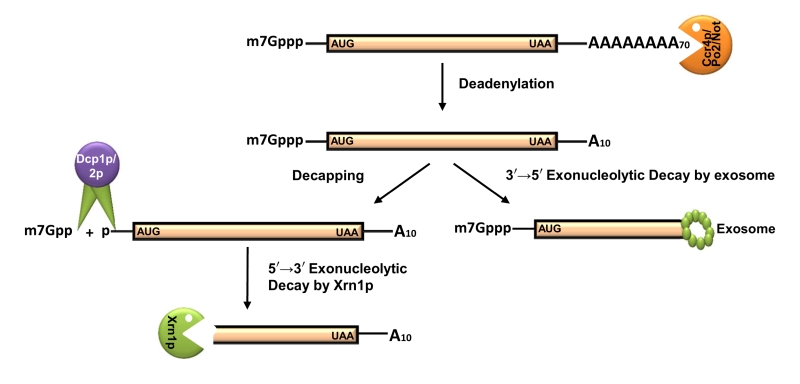
FIGURE 2: Default pathway of mRNA degradation in S. cerevisiae. Almost all mRNAs undergo decay by the deadenylation-dependent pathway. Thereby, the poly(A) tail is gradually and progressively shortened by the deadenylase activity of the Ccr4/Pop2/Not complex. Following deadenylation, the mRNA can be degraded by one of two mechanisms. The major mechanism involves decapping by Dcp1p/2p, following a 5’→3’ decay by Xrn1p. The minor mechanism includes a 3’→5’ decay by the cytoplasmic exosome and Ski7p. AUG and UAA are indicating the beginning and end of the ORF carried by the message. Only relevant decay components are shown by annotated symbols. Proteins which remain associated to translating/degrading mRNAs during different stages of decay are not shown.