Back to article: Ydj1 governs fungal morphogenesis and stress response, and facilitates mitochondrial protein import via Mas1 and Mas2
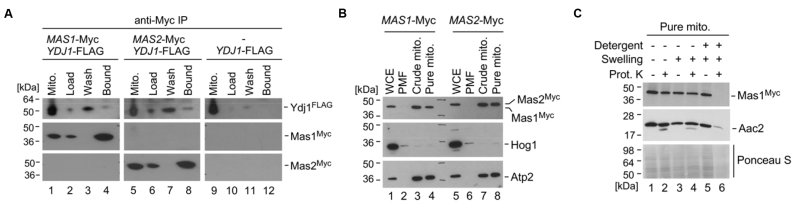
FIGURE 5: Ydj1 interacts with Mas1 and Mas2, which are localised to the mitochondria. (A) Gradient purified mitochondria from 2×FLAG-YDJ1/ydj1, 2×FLAG-YDJ1/ydj1 MAS1-Myc and 2×FLAG-YDJ1/ydj1 MAS2-Myc incubated for 6 hours at 37oC were lysed and used for the co-immunoprecipitation assay. A physical interaction between Ydj1 and Mas1 or Mas2 was determined by positive anti-FLAG and anti-Myc (Santa Cruz Biotechnology) antibody staining in the Bound frac-tions. (B) Whole-cell extracts (WCE), post-mitochondrial fractions (PMFs), crude and gradient-purified (pure) mitochondria were obtained from yeast cultures grown in YPD at 30oC for 16 hours. Fractions were ana-lysed by SDS-PAGE and immunoblotting with antibodies against the Myc epitope (Roche) used to tag Mas1 and Mas2. The purity of the mitochondrial fractions was determined by the absence of a Hog1-specific band normally present in the cyto-solic fractions. Antibodies against the inner mitochondrial protein Atp2 were used as a marker for the mitochondrial fraction. (C) Mas1 is localised to the mitochondrial matrix. Gradient-purified mitochondria harboring Myc-tagged Mas1 were subjected to osmotic disruption of the outer mitochondrial membrane (Swelling) and lysis with dodecyl malto-side (Detergent) to disrupt the inner membrane in combination with Proteinase K (Prot. K) treatment. The fractions were subjected to Western blot analysis using anti-Myc antibodies to visualise Mas1 localisation, and anti-serum against the inner mitochondrial membrane protein Aac2 used as a marker for submitochondrial localization. Ponceau S staining was used to ascertain the loading of the mitochondrial fractions.