Reviews:
Microbial Cell, Vol. 5, No. 2, pp. 63 - 73; doi: 10.15698/mic2018.02.612
Ras signalling in pathogenic yeasts
1 Kent Fungal Group, School of Biosciences, University of Kent, Canterbury, Kent, United Kingdom, CT2 7NJ.
2 Laboratoire national de santé, 1, Rue Louis Rech, L-3555 Dudelange, Luxembourg.
Keywords: Ras signalling, C. albicans, C. neoformans, pathogenicity, morphogenesis, biofilm.
Abbreviations:
GEF – guanine nucleotide exchange, factor,
MAP – mitogen-activated protein,
MTL – mating type locus,
PKA – protein kinase A.
Received originally: 22/09/2017 Received in revised form: 09/12/2017
Accepted: 11/12/2017
Published: 18/12/2017
Correspondence:
Campbell W. Gourlay, Kent Fungal Group, School of Biosciences, University of Kent, Canterbury, Kent, United Kingdom, CT2 7NJ; C.W.Gourlay@kent.ac.uk
Conflict of interest statement: The authors declare no conflict of interest.
Please cite this article as: Daniel R. Pentland, Elliot Piper-Brown, Fritz A. Mühlschlegel and Campbell W. Gourlay (2017). Ras signalling in pathogenic yeasts. Microbial Cell 5(2): 63-73. doi: 10.15698/mic2018.02.612
Abstract
The small GTPase Ras acts as a master regulator of growth, stress response and cell death in eukaryotic cells. The control of Ras activity is fundamental, as highlighted by the oncogenic properties of constitutive forms of Ras proteins. Ras also plays a crucial role in the pathogenicity of fungal pathogens where it has been found to regulate a number of adaptions required for virulence. The importance of Ras in fungal disease raises the possibility that it may provide a useful target for the development of new treatments at a time when resistance to available antifungals is increasing. New findings suggest that important regulatory sequences found within fungal Ras proteins that are not conserved may prove useful in the development of new antifungals. Here we review the roles of Ras protein function and signalling in the major human yeast pathogens Candida albicans and Cryptococcus neoformans and discuss the potential for targeting Ras as a novel approach to anti-fungal therapy.
RAS PROTEINS – FORM AND FUNCTION
The Ras superfamily consists of small G-binding proteins that have been divided into five main groups on the basis of sequence and functional similarity: Ras, Rho, Arf/Sar, Rab and Ran [1]. As with all G proteins, Ras activation is dependent on the exchange of bound GDP for GTP and deactivation via hydrolysis of GTP to GDP. Cycling between active and non-active states is regulated by Guanine nucleotide exchange factors (GEF), dissociation inhibitor’s (GDI) and GTPase Activating Proteins (GAP) accessory proteins [2]. The activation and deactivation cycle of Ras proteins couple a range of stimuli to effector proteins and as such these G-proteins serve as regulatory “switches” within a variety of cellular processes. The importance of Ras signalling is highlighted by the dramatic effects that can be observed upon inappropriate activation. For example mutations that lead to the constitutive activation of Ras signalling have been estimated to occur in ~50% of all tumours [3]. In addition, Ras proteins play important roles in the regulation of growth and adaption in fungal cells. In this review we will focus on Ras protein function in the major human yeast pathogens Candida albicans (C. albicans) and Cryptococcus neoformans (C. neoformans).
RAS SIGNALLING IN THE FUNGAL PATHOGEN OF HUMANS C. ALBICANS
C. albicans virulence and pathogenicity
C. albicans is a commensal organism that is commonly found on the mucosal surfaces of the oral cavity, gastrointestinal tract and genitourinary tract of healthy individuals [4][5]. However C. albicans is also a well characterised opportunistic pathogen [4][6] and a serious health-risk amongst immunocompromised individuals, such as those suffering from HIV infection, or persons living with indwelling medical devices such as catheters or voice prostheses [7][8]. The infections caused by C. albicans range from superficial infection of mucosal and non-mucosal surfaces (candidiasis) to a full systemic infection (candidaemia) also affecting internal organs [4]. Superficial mucosal surface infections can be readily treated with a range of antifungals. For example, vulvovaginal candidiasis (VVC) is usually successfully treated with azole antifungals like fluconazole [9]. However, candidaemia is associated with a high mortality even when treated with a variety of classes of antifungal agents [10]. As with many C. albicans infections, weakened immune defences are a significant risk factor for developing candidaemia. In healthy individuals, neutrophils provide suitable defence against C. albicans. As such, neutropenia, either as a result of particular blood cancers or treatment with immunosuppressants, significantly increases the risk of developing candidaemia. Furthermore, damage to the mucosa of the gastrointestinal tract, for example due to surgery, is also a risk factor as it enables the spread of C. albicans [11]. The symptoms of candidaemia range from fever and chills which do not abate following antibiotic treatment to severe sepsis or septic shock similar to that of bacterial septicaemia [9]. However, a lack of precise symptoms can lead to delayed diagnosis and required antifungal treatment leading to increased mortality [12]. It has been reported that even a delay of as little as 12-24h can double mortality rate [13]. Due to this, it has been suggested to prophylactically administer antifungals after any event which is likely to increase the risk of candidaemia, such as after abdominal surgery or bone marrow transplant [9]. Although the majority of cases of candidiasis and candidaemia are caused by C. albicans, there are other species within the Candida genus which are also pathogenic in humans. These include Candida glabrata, Candida tropicalis, Candida dubliniensis and Candida parapsilosis. Candida species have increasingly become associated with nosocomial infections [6]; in fact, C. albicans is recognised as the fourth most common cause of all hospital-acquired infections in the USA [4].
–
Morphogenesis, pathogenicity and Ras signalling in C. albicans
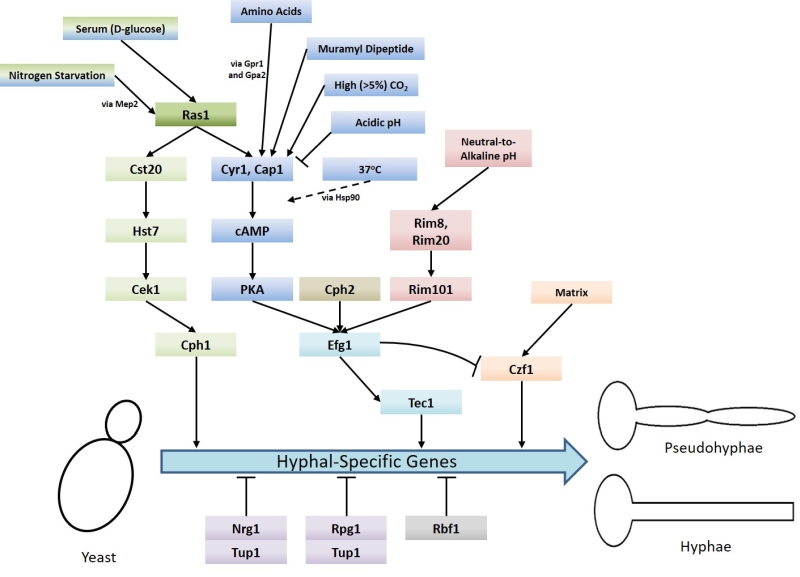
An important aspect of C. albicans biology, in terms of pathogenesis, is its ability to undergo morphogenesis from a yeast, to pseudohyphal or hyphal forms in response to environmental cues. The virulence of C. albicans is closely linked with the capacity to switch between these forms. Hyphal C. albicans cells are frequently located at sites of tissue invasion, moreover, cells which are unable to readily form hyphae exhibit reduced virulence [4]. However as strains that are incapable of growing in the yeast form also have less virulence it has been proposed that both the yeast and hyphal forms play important roles during infection [14][15]. Ras signalling is crucial to the integration of environmental cues with morphogenesis and C. albicans possesses two Ras genes – RAS1 and RAS2, which encode the Ras1 protein and a highly divergent Ras-like protein termed Ras2 [16]. The importance of Ras1 signalling to the virulence of C. albicans is demonstrated by the fact that mutants which lack Ras1 are defective in their ability to undergo hyphal transition and exhibit reduced virulence in mouse infection models [17]. Ras signalling is now known to mediate the induction of hyphal growth in response to a variety of environmental cues including growth at 37°C (via alleviation of Hsp90-mediated repression of the Ras1-cAMP-PKA pathway) [18][19], exposure to high levels of CO2 [20], N-acetylglucosamine[21] and serum exposure [22]. These environmental signals are transduced through the cyclic AMP-protein kinase A and a mitogen-activated protein (MAP) kinase pathways (Figure 1) [4]. These pathways culminate in the regulation of transcription factors which control the expression of hyphal-specific genes (HSGs) such as Als3 (adhesin) [23], Hwp1 (invasin) [24], Hyr1 (host immune response modulator) [25], and Hgc1 (hyphal-specific cyclin) [26].
–
Upon activation, Ras1 directly interacts with and activates Cyr1 (the C. albicans adenylate cyclase), causing an increase in the production of the second messenger cAMP [17]. cAMP causes the derepression of two isoforms of protein kinase A (PKA) by triggering the dissociation of the PKA regulatory subunit (Bcy1) from the catalytic subunits (Tpk1 or Tpk2). The activation of PKA stimulates several processes within the cell including the yeast-to-hyphae switch [27]. PKA is believed to phosphorylate the transcription factor Efg1 on threonine-206, thereby activating it and resulting in the expression of HSGs [28]. The Tpk1 and Tpk2 isoforms have some redundant functions in C. albicans, however, they also have specific roles in filamentation. For example, Tpk1 is necessary for the expression of genes encoding proteins involved in branched chain amino acid biosynthesis, and Tpk2 negatively regulates iron uptake genes and positively regulates those associated with trehalose degradation and water homeostasis [29].
–
A number of environmental signals promote the yeast-to-hyphae switch via the Ras1-Cyr1-PKA pathway. Some of these signals, including CO2 interface directly with the Cyr1 adenylate cyclase to activate it [20]. CO2 is able to do this because, unlike most signalling molecules, it is able to enter the cell by simple diffusion and is maintained in the cell as HCO3– via conversion by a carbonic anhydrase encoded by NCE103 [20]. It has recently been discovered that the expression of NCE103 is controlled in response to CO2 availability by the bZIP transcription factor Rca1; Rca1 is regulated in a CO2-dependent manner by the Sch9 kinase via a cascade mediated by lipid/Pkh1/2 signalling [30]. A lysine residue at position 1373 is critical for CO2 activation of Cyr1. This lysine residue is located in the C-terminal catalytic domain and makes up a receptor site which detects increased HCO3– levels [31], leading to increased cAMP production and activation of PKA filamentation [20]. The response of C. albicans to CO2 is of interest because within a mammalian host, the levels of CO2 are approximately 150x that of normal air (~5% compared to 0.03%). It may be the case that high levels of CO2, such as are found within the upper respiratory tract, may promote C. albicans colonisation. However, it is interesting to note that other pathogenic Candida species, including C. dubliniensis, C. glabrata, C. paropsilosis and C. krusei, do not undergo the yeast-to-hyphae transition in response to elevated CO2 [20]. Although this does not rule out the fact that the adenylyl cyclase of the latter species is activated by carbon dioxide/bicarbonate; the physiological significance of CO2 sensing with regards to Candida infection remains to be determined.
–
Muramyl dipeptide (MDP), the minimal biologically active subunit of bacterial peptidoglycan, also induces C. albicans filamentation by acting directly upon Cyr1 [32]. A further signal which can cause morphogenesis through direct interaction with Cyr1 are amino acids. Amino acids, when in the presence of glucose, activate Cyr1 via upstream signalling through the G-protein coupled receptor Gpr1 and its Gα protein Gpa2 [33]. Upon its activation by Gpr1, Gpa2 is believed to bind to a Gα domain on the Cyr1 adenylate cyclase thereby activating it [33]. This binding of Gpa2 to a fungal adenylate cyclase Gα domain has been demonstrated in fission yeast [34] but it is yet to be proved experimentally in C. albicans.
–
In contrast, acidic pH causes a reduction in signalling through the Ras1-Cyr1-PKA pathway via a Ras1-independent downregulation of Cyr1 activity [35]. C. albicans cells grown at pH 4 in hyphae-inducing conditions do not form hyphae, instead remaining as yeast or pseudohyphal cells and this is not reliant upon Ras1. It has also been observed that low extracellular pH results in fast and sustained decreases in intracellular pH which potentially contributes to reduced cAMP signalling through the reduction of intracellular bicarbonate levels [35].
–
The yeast-to-hyphae switch in response to exposure to serum relies on Ras1 signalling upstream of Cyr1 [22]. The component of serum principally responsible for the induction of hyphal growth is D-glucose which is able to activate both the Ras1-Cyr1-PKA pathway and the MAP-kinase pathway [22][36]. The precise mechanism of Ras1 activation by D-glucose in C. albicans remains to be elucidated but in Saccharomyces cerevisiae (S. cerevisiae) it depends on both an intracellular phosphorylated form of D-glucose and a G-protein coupled receptor Gpr1 with its Gα protein Gpa2 [37]. Gpr1-type receptors have been characterised in C. albicans [33] and so a similar mechanism for Ras1 activation may exist in this pathogen. However, deletion of either CaGpr1 or CaGpa2 had no effect on D-glucose-mediated cAMP signalling, but deletion of CaCdc25 (the C. albicans Ras1 GEF) or CaRas1 eliminated this signalling [33]. These findings indicate Ras1 activation via Cdc25 is the primary mechanism by which D-glucose induces morphogenesis. The response of C. albicans to D-glucose is of physiological relevance because links between candida infection and hyperglycaemia [38] as well as insulin-dependent diabetes mellitus [39] have been reported. Moreover, C. albicans cells have increased resistance to oxidative and cationic stresses upon exposure to levels of glucose that may be found in the bloodstream [40].
–
Ras1 has also been shown to regulate hyphal transition in response to other environmental cues, such as nitrogen starvation [41], via MAP kinase signalling. As with the Cyr1/PKA pathway, the regulatory MAP kinase cascade is also activated by Ras1 [17] and consist of the kinases Cst20, Hst7 and Cek1 [42] [43][44][45][46]. Ras1/MAPK signalling culminates in the phosphorylation and activation of the transcription factor Cph1, which in turn promotes the expression of HSGs (Figure 1) [33]. It has been shown that inactivation of the Ras1-Cyr1-PKA pathway inhibits filamentous growth in the majority of usual hyphae-inducing conditions, however, inactivation of the MAP-kinase pathway only prevents filamentous growth in only a specific subset of conditions [43].
–
The CaRas2 protein contains several variations in conserved motifs typically thought to be critical for Ras-related activities and is thus considered an unusual Ras protein. Sequence alignment using BLAST has shown that the C. albicans Ras2 protein only has 25-30% identity with all other fungal Ras proteins in the database except a Ras-like protein only found in Candida dubliniensis (80% identity) [47]. When RAS2 is deleted in a ras1∆/∆ background, intracellular cAMP levels are restored to approximately 30% of wild type levels (ras1∆/∆ mutant has a 20x reduction in cAMP). Ras1 and Ras2 may therefore exhibit antagonistic roles in C. albicans [47]. This is intriguing since the deletion of RAS2 in a ras1∆/∆ background results in a significantly increased defect in hyphal morphogenesis. Nevertheless as ras2∆ mutants themselves exhibit normal hyphal development [47] the role of Ras2 in morphogenesis and pathogenesis in C. albicans has yet to be elucidated.
–
In addition to the signalling pathways which drive the yeast-to-hyphae switch, there are also negative regulators that are controlled by Ras signalling (Figure 1). Hyphal-specific genes are repressed by the global-repressor Tup1 [48] via the specific DNA-binding proteins Nrg1 [49][50] and Rfg1 [51]; the deletions of each of these three proteins results in C. albicans cells which are constitutively hyphal even under non-hyphal inducing conditions [48][49][50][51]. Approximately half of the genes found to be upregulated during hyphal development in response to 37°C and serum are repressed by Tup1 and Nrg1 or Rfg1, suggesting that repression removal is a crucial step in the yeast-to-hyphae switch [52]. Consistently, it has been found that hyphal-inducing conditions such as serum exposure and growth at 37°C cause a reduction in the expression levels of NRG1, leading to the conclusion that one way in which repression of hyphal-specific genes is overcome during the yeast-to-hyphae switch is via down-regulation of the repressors [50]. Ras1-Cyr1-PKA pathway activation results in the prompt but short-term removal of Nrg1 from the promoters of hyphal-specific genes. The maintenance of this repression elimination, and hence hyphal development, is achieved through the subsequent recruitment of the Hda1 histone deacetylase which deacetylates a subunit of NuA4 histone acetyltransferase, causing it to also be removed from the promoter. This results in the coiling of the portion of the promoter containing the Nrg1 binding site, preventing the re-binding of Nrg1. It is important to note the removal of Nrg1 is an absolute prerequisite for the Hda1 recruitment [53].
–
Ras signalling and white-opaque switching in C. albicans
C. albicans was traditionally considered to be asexual, only existing as an obligate diploid [54]. However, it has now been discovered that mating occurs between homozygous diploid mating type-like (MTL) a and α strains in this organism, producing an a/α tetraploid product [54][55] which then undergoes ‘concerted chromosome loss’ to form diploid progeny [56]. C. albicans has also been reported to have a viable haploid state which can mate to restore the diploid form [57]. While this is similar to the mating program in S. cerevisiae (two haploid mating types; a and α which combine to generate an a/α diploid product) [58] it differs in several key respects.
–
Mating in C. albicans is reliant on a reversible phenotypic switch between two states termed ‘white’ and ‘opaque’. Only the ‘opaque’ state is capable of mating efficiently; ‘opaque’ cells have been demonstrated to mate approximately 106 fold more readily than ‘white’ cells [59]. ‘White’ cells are fairly round and form white, dome-shaped colonies on solid agar, they also express a specific set of genes. In contrast, ‘opaque’ cells tend to be larger and more oblong, forming darker colonies which grow flatter against solid agar. ‘Opaque’ cells also express a specific set of genes which differ from those expressed in ‘white’ cells [60].
–
This unusual mating program involving a reversible phenotypic switch hitherto seems to be unique to C. albicans as well as the very closely related fungal species C. dubliniensis [61]. It appears that ‘white’ cells are better suited for growth and survival within a mammalian host. Therefore, it is likely this unusual mating program has evolved to allow C. albicans to survive the variety of environments within a mammalian host while still being able to produce mating-competent cells [59].
–
Only C. albicans cells which are homozygous at the MTL locus (a/a or α/α) are capable of reversibly switching between ‘white’ and ‘opaque’ states, and are thus capable of efficient mating [55][62]. This is because two homeodomain proteins called Mtla1 and Mtlα2, encoded by the MTLa and MTLα alleles respectively, work together to inhibit white-opaque switching [59]. These proteins are both present in MTL heterozygous cells (a/α) and thus white-opaque switching cannot occur, only one of these two proteins is present in MTL homozygous cells, meaning white-opaque switching is not suppressed and mating (between a/a cells and α/α cells) can take place.
–
The transcription factor Wor1 is the master regulator of the white-opaque switch and acts in an all-or-nothing manner; it is virtually undetectable in white cells but highly expressed in opaque cells (expression is approximately 47-fold higher in opaque cells) [63]. Wor1 has been shown to control the expression of its own gene WOR1 in either a positive feedback or double-negative feedback loop and drives the C. albicans cell into the opaque state [64]. Due to WOR1 being repressed by the Mtla1 and Mtlα2 proteins [63], Wor1 is not present in MTL heterozygous cells (a/α) cells and ectopic WOR1 expression in these cells causes them to undergo the white-opaque switch [64].
–
The Ras1-Cyr1-PKA pathway is known to have a role in the white-opaque switch. High levels of CO2 can induce this switch; in 20% CO2 the switch has been reported to occur with up to 105x more frequency compared to normal air [65]. In a ras1Δ/Δ mutant and a cdc35Δ/Δ mutant (which lacks the CO2-responsive Cyr1 adenylate cyclase) white-opaque switching is reduced in both normal air and 1% CO2 compared to wild-type but normal in 20% CO2 [65]. This suggests signalling through the Ras1-Cyr1-PKA pathway is important for the switch in normal air and moderate CO2 but not in very high levels of CO2.
–
High N-acetylglucosamine levels also induce the white-opaque switch, and this switch in response to N-acetylglucosamine is significantly diminished from 90.5±3.8% cells in the wild-type to 11.2±1.5% in a ras1Δ/Δ mutant [66]. Likewise, in a cdc35Δ/Δ mutant, 8.0±3.5% of cells undergo the white-opaque response in the presence of N-acetylglucosamine compared to 86.9±4.3% in the wild-type [66]. These results suggest the N-acetylglucosamine switch occurs via signalling through the Ras1-Cyr1-PKA pathway. Furthermore, when the master switch regulator Wor1 is overexpressed in a ras1Δ/Δ, cdc35Δ/Δ, tpk1Δ/Δ or tpk2Δ/Δ background, the cells are driven into the opaque state. Conversely, a wor1Δ/Δ mutant does not switch in the presence of N-acetylglucosamine [66]. Wor1 contains a consensus PKA phosphorylation motif with a phosphorylatable threonine at residue 67 [64], and it has been demonstrated that this threonine is absolutely required for white-opaque switching in response to N-acetylglucosamine [66]. These results imply that the transcription factor Wor1 functions downstream of the Ras1-Cyr1-PKA pathway to induce white-opaque switching in the presence of N-acetylglucosamine.
–
In addition to Wor1 there are also other transcription factors which act as switch regulators, specifically; Efg1 (itself Ras1-regulated), Czf1, Wor2 [67] and Wor3 [68]. Binding sites for Wor1 have been identified upstream of EFG1, CZF1 and WOR2, indicating that these transcription factors function in a regulatory circuit composing positive-feedback loops with the Ras1-regulated Wor1 in a central position [67].
RAS SIGNALLING AND C. ALBICANS BIOFILM FORMATION
Biofilms are structured communities of microorganisms which are attached to either a living or non-living surface. The cells are often encased within a matrix of self-made extracellular polymeric substance (EPS); this EPS is composed of DNA [69][70], lipids [69], proteins [69][71] and polysaccharides [69]. Medically, biofilms are of particular importance because it is thought that a significant percentage of human microbial infections include biofilm formation [72][73][74]. Moreover, cells which reside within biofilms have distinctive phenotypes compared to planktonic cells, for example, they exhibit increased resistance to antibiotic and antifungal drugs. The reasons for this increased resistance are complex but include the presence of an extracellular matrix reducing the ability of antimicrobial agents to reach the cells, metabolic differences (such as modulation of glycolysis, ergosterol biosynthesis and mitochondrial respiration) [75] inherent to biofilms and upregulation of efflux pumps [76]. C. albicans biofilms are usually composed of a mixture of morphological forms; typically yeast, pseudohyphal and true hyphal cells are all present within a mature biofilm [6][77][78]. The formation of a biofilm is the result of a very precise and complex series of events that are divided into distinct stages; attachment, initiation, maturation and dispersal. Biofilm formation is therefore complex and highly regulated with more than 1000 genes found to be upregulated during biofilm development [79].
–
Hyphal cells are important for the formation of C.albicans biofilms, one reason for this is that the expression of several cell surface adhesins, such as Hwp1 and Eap1, is increased during hyphal growth. These adhesins are required for the initial attachment phase of biofilm formation, and as a result it means Ras signalling is strongly linked to their development [80]. This is highlighted by the finding that the hyphal-defective mutant efg1∆/∆ is unable to form biofilms [81]. Rather than the true basal layer which wild-type C. albicans cells form, efg1∆/∆ mutants produce very few surface-attached cells. Despite this the surface-attached mutant cells do display resistance to both fluconazole and amphotericin B [81]. These are important observations that suggest surface-adhesion is sufficient to induce an antifungal resistance response in biofilms [78].
–
The transcription factor Bcr1, which is upregulated by Tec1 (Figure 1) is an important regulator of C. albicans biofilm formation. The bcr1∆/∆ mutant is unable to form biofilms and also cannot switch to hyphal growth under certain conditions. However, when present within mixed biofilms formed using wild-type cells, bcr1∆/∆ mutant cells can form hyphae [82]. Interestingly, bcr1∆/∆ hyphal cells themselves are unable to adhere to surfaces and initiate biofilm formation. Bcr1 upregulates a number of genes which encode cell wall proteins, including the adhesins Als1, Als3, and Hwp1 [79]. It is likely therefore that hyphal associated cell wall composition is crucial for biofilm formation. Ras1-Cyr1-PKA signalling is important in this respect as it regulates the expression of adhesins, such as Als1, via its control of the activity of the key transcription factors Efg1, Tec1 and Bcr1 [79]. cAMP/PKA signalling is also likely to impact upon biofilm formation with respect to CO2 levels. For example, a local accumulation of CO2 within C. albicans colonies was sufficient to induce filamentous growth [31]. Although the precise roles have yet to be determined, it will be interesting to examine how CO2 signalling contributes to biofilm establishment in vivo. The final stage of biofilm development is the dispersal stage in which a mature biofilm begins to ‘throw’ fragments off in order to establish additional biofilms elsewhere [78]. The transcription factor Nrg1, whose degradation is inhibited by the C. albicans quorum sensing molecule farnesol, has been shown to promote biofilm cell dispersion [83]. As Ras activation can influence Nrg1 levels, and as farnesol has been shown to promote Ras1 degradation it will be of interest to investigate how Ras signalling influences the biofilm dispersion process.
RAS SIGNALLING AND VIRULENCE IN CRYPTOCOCCUS NEOFORMANS
The prototypical species in the Cryptococcus genus is C. neoformans which is an encapsulated, pleomorphic yeast [84]. Similar to C. albicans, C. neoformans is an opportunistic human pathogen with infection primarily being associated with a compromised immune system; cryptococcal infections (cryptococcosis) are a particular a problem amongst HIV/AIDS patients [85]. The Centers for Disease Control and Prevention estimates the number of deaths attributed to cryptococcal meningitis in HIV/AIDS patients is as high as 181000 per year [86]. C. neoformans possesses two RAS genes, denoted RAS1 and RAS2, which encode the Ras1 and Ras2 proteins respectively [87]. The Ras1 protein is highly conserved: it is a homolog of the traditional Ras proteins in mammalian cells such as H-RAS [84] and has been demonstrated to be essential for multiple processes in C. neoformans including growth at 37°C (Ras1 is not needed for growth at 30°C), mating, agar adherence and filamentation [88][89]. Due to its crucial role in these processes, particularly thermotolerance, Ras1 is considered as an important virulence factor. Indeed, the ∆ras1 mutant strain is avirulent in a rabbit model of cryptococcal meningitis [88]. Moreover, introduction of a dominant active RAS1 allele (RAS1Q67L), which was constructed by introducing a point mutation in the active site of the GTPase domain of Ras1, resulted in significant increases in filamentation and agar invasion of haploid C. neoformans cells [88]. These are two properties which are very important for virulence, supporting the conclusion that Ras1 signalling is vital to the pathogenicity of C. neoformans. It is important to note that in other model systems, including Caenorhabditis elegans [90] and Drosophila melanogaster [91], the ∆ras1 mutant strain exhibited decreased virulence at lower temperatures. This implies that, at least for non-mammalian model systems, Ras1 signalling may have significant functions in the pathogenicity of C. neoformans besides thermotolerance.
–
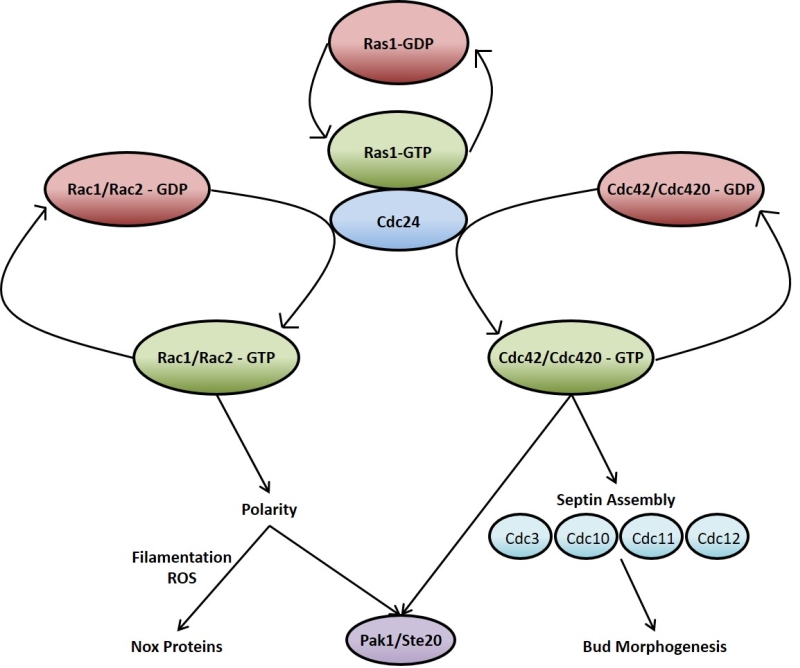
Ras signalling in C. neoformans, much like in mammalian cells, acts through and/or in concert with other Rho-type GTPases such as Cdc42 and Rac [89][92][93][94][95]. As is the case with RAS, C. neoformans possesses duplicate copies of CDC42 (CDC42 and CDC420) [92] and RAC (RAC1 and RAC2) [93]. The overexpression of any of these genes overcomes the thermotolerance deficiencies of the ∆ras1 mutant, supporting a model whereby these GTPases act downstream of Ras1 (Figure 2) [89][95]. At 37°C, ∆ras1 C. neoformans mutants amass a number of defects in cell cycle progression, specifically concerning cell polarisation and cytokinesis. While neither is required for viability when the organism is not under stress, the two Cdc42 paralogues are essential for septin organisation and effective cytokinesis at 37°C [92]. Ras1 is also important in the activation of Cdc42 and Rac [94]. Indeed, Cdc24 which is the reported GEF for Cdc42, undergoes a GTP-dependent physical interaction with Ras1 [89][96]. Once activated, Cdc42 organises the septin proteins Cdc3, Cdc10, Cdc11 and Cdc12, causing them to localise to the bud neck ready for cytokinesis [94]. In addition to septin protein organisation, Ras1 signalling through Cdc42 is also necessary for normal bud morphology [94]. Cdc42 appears to be the more important to C. neoformans virulence since it, and not Cdc420, is upregulated during temperature stress and is necessary for virulence in a mouse model of Cryptococcus infection [92]. As previously mentioned, Ras1 is not required for growth of C. neoformans at 30°C. The reason for this appears to be due to basal levels of Cdc42 activity even in the absence of Ras1 signalling which is sufficient to allow proliferation at this lower temperature [92][94].
–
Although functional redundancy may exist between Cdc42 and Rac, the two Rac paralogues in C. neoformans are predominantly involved in cell polarisation and polarised growth; particularly in hyphal development when mating and in the transport of vesicles during the yeast phase of growth [93][95][97]. The ∆rac1 mutant strain, while still able to grow at 37°C, displays a significant defect in haploid filamentation along with reduced mating [95]. ∆rac1 and ∆rac2 mutant strains have increased yeast cell size [93], in keeping with the phenotype exhibited by ∆ras1 cells when grown at 37°C [88] as well as that shown by polarity mutants of the prototypical budding yeast S. cerevisiae [98]. This highlights the important role Ras1 signalling through Rac1 and Rac2 plays in cell polarity and this precisely impacts hyphal development and yeast cell size.
–
As with C. albicans, the ability to sense CO2 is of critical importance to C. neoformans. Capsule biosynthesis, which is a major virulence characteristic, is increased by high CO2 levels and this is mediated by the C. neoformans adenylate cyclase Cac1 [20]. Importantly, it has been demonstrated that a fragment of CnCac1 can restore hyphal development in response to elevated CO2 within a cyr1∆/∆ C. albicans mutant. This implies the link between CO2 sensing and cAMP could be a general feature of pathogenic yeasts since C. neoformans and C. albicans are evolutionarily distantly related but their adenylate cyclases are functionally highly conserved [20].
–
The C. neoformans Ras2 protein plays separable roles to those of Ras1. For instance, ∆ras2 cells do not exhibit reduced virulence in a mouse model of cryptococcosis [87]. Furthermore, ∆ras2 cells do not exhibit the same mating defects associated with the loss of RAS1 [87]. RAS2 is also found to be expressed at low levels compared to RAS1 and ∆ras2 mutants do not display any differences in growth or differentiation when compared to wild-type [87]. However, it is worth noting that overexpression of RAS2 in a ∆ras1 mutant is able to rescue the mating defect and partially restores its growth at high temperature [87]. Moreover, the double ∆ras1∆ras2 mutant has growth defects at all temperatures which are worse than with either single mutation alone [87]. These findings suggest that Ras1 and Ras2 protein may have some overlapping functions in growth, mating and virulence that have yet to be properly defined.
TARGETING RAS TO COMBAT YEAST PATHOGENESIS
Given its role in the virulence and pathogenic properties of multiple yeast (and other fungal) species, it would seem to have potential as anti-fungal drug target. Additionally, it has been shown that the manipulation of Ras signalling is an important control point in the activation of apoptosis in the budding yeast S. cerevisiae [99][100][101] and in C. albicans [102]. These findings present the possibility that the pharmacological manipulation of Ras signalling may be useful in the induction of yeast cell death. Some current antifungals do exert an effect on fungal Ras signalling but this is indirect. For example, amphotericin B inserts into ergosterol-containing fungal membranes to form aqueous pores [103] which cause a local thinning of the bilayer. This thinning of the bilayer has been proposed to force lipid-anchored Ras proteins into sterol-rich lipid rafts, promoting its interactions with downstream proteins and thus activating signalling [104]. Enhanced Ras signalling through pharmacological manipulation induces yeast apoptosis and oxidative damage via the cAMP-PKA pathway, indicated by the fact S. cerevisiae ∆ras1, ∆ras2, ∆tpk1, ∆tpk2 and ∆tpk3 mutants have reduced amphotericin B-induced reactive oxygen species production and hence are unaffected by the lethal effect of amphotericin B [105]. Other fungicides such as miconazole and ciclopirox have been reported to cause fungal cell death via a similar mechanism [105].
–
Ras proteins have proven difficult to directly target pharmacologically and because the GTPase domain is very highly conserved any attempt to target a fungal Ras may result in unfavourable side effects upon the host. Encouragingly a recent discovery does suggest that fungal Ras proteins may be targetable after all. A recent study demonstrated that the RasA protein of the human fungal pathogen Aspergillus fumigatus possesses a short N-terminal tail domain which is missing from Ras homologs in higher eukaryotes [106]. This domain takes the form of a short length of amino acid residues which terminates in an arginine and has been dubbed the invariant arginine domain (IRD). Through sequence alignment, it has been reported that the IRD is present in many different fungal pathogens including C. albicans and C. neoformans, as well as non-pathogenic fungi such as S. cerevisiae and S. pombe [106]. Mutation of the A. fumigatus RasA IRD decreased the activation of PKA as well as reducing the interaction of RasA with Cdc42 [106] which controls polarity and thus is essential for normal growth and cell division [107]. In line with this, mutational analyses confirmed that the IRD is necessary for polarised morphogenesis, a characteristic strongly linked with pathogenesis in A. fumigatus, and asexual development [106]. These findings point towards the possibility of designing drugs to specifically target the IRD as a new Ras-based pan-antifungal therapy. As an extension of this it may also be possible to target regions within the C-terminal variable domain of fungal Ras proteins to modulate its activity.
CONCLUSIONS
Ras signalling has proven to be an important component of the growth and adaptability of fungal cells, as is indeed the case in higher eukaryotes. It is also crucial to the virulence and pathogenic properties of fungal species. Ras signalling therefore represents an interesting therapeutic target if fungal specific targets can be found. This may be particularly effective when used in combination with other antifungal agents. A full understanding of the Ras signalling network and its effectors will be required to achieve this aim. As an example, it may be possible to identify a small molecule that can activate fungal Ras in C. albicans. This may, at first glance, seem an unlikely intervention given the role of Ras signalling in promoting growth. However, as we now also know that the activation of Ras sensitizes cells to cell death, such an approach may prove particularly effective when used in combination with existing antifungals. Given the rise of antifungal resistance and our limited number of existing targets such “knowledge based” approaches will doubtless prove crucial in future therapy development.
References
- H.R. Bourne, D.A. Sanders, and F. McCormick, "The GTPase superfamily: conserved structure and molecular mechanism", Nature, vol. 349, pp. 117-127, 1991. http://dx.doi.org/10.1038/349117a0
- M.S. Boguski, and F. McCormick, "Proteins regulating Ras and its relatives", Nature, vol. 366, pp. 643-654, 1993. http://dx.doi.org/10.1038/366643a0
- P.M. Campbell, and C.J. Der, "Oncogenic Ras and its role in tumor cell invasion and metastasis", Seminars in Cancer Biology, vol. 14, pp. 105-114, 2004. http://dx.doi.org/10.1016/j.semcancer.2003.09.015
- J. Berman, and P.E. Sudbery, "Candida albicans: A molecular revolution built on lessons from budding yeast", Nature Reviews Genetics, vol. 3, pp. 918-931, 2002. http://dx.doi.org/10.1038/nrg948
- S. Ganguly, and A.P. Mitchell, "Mucosal biofilms of Candida albicans", Current Opinion in Microbiology, vol. 14, pp. 380-385, 2011. http://dx.doi.org/10.1016/j.mib.2011.06.001
- L. Douglas, "Candida biofilms and their role in infection", Trends in Microbiology, vol. 11, pp. 30-36, 2003. http://dx.doi.org/10.1016/S0966-842X(02)00002-1
- J.S. Finkel, and A.P. Mitchell, "Genetic control of Candida albicans biofilm development", Nature Reviews Microbiology, vol. 9, pp. 109-118, 2010. http://dx.doi.org/10.1038/nrmicro2475
- M.J. Talpaert, A. Balfour, S. Stevens, M. Baker, F.A. Muhlschlegel, and C.W. Gourlay, "Candida biofilm formation on voice prostheses", Journal of Medical Microbiology, vol. 64, pp. 199-208, 2015. http://dx.doi.org/10.1099/jmm.0.078717-0
- J. Kim, and P. Sudbery, "Candida albicans, a major human fungal pathogen", The Journal of Microbiology, vol. 49, pp. 171-177, 2011. http://dx.doi.org/10.1007/s12275-011-1064-7
- C. Kibbler, S. Seaton, R. Barnes, W. Gransden, R. Holliman, E. Johnson, J. Perry, D. Sullivan, and J. Wilson, "Management and outcome of bloodstream infections due to Candida species in England and Wales", Journal of Hospital Infection, vol. 54, pp. 18-24, 2003. http://dx.doi.org/10.1016/S0195-6701(03)00085-9
- A.Y. Koh, J.R. Köhler, K.T. Coggshall, N. Van Rooijen, and G.B. Pier, "Mucosal Damage and Neutropenia Are Required for Candida albicans Dissemination", PLoS Pathogens, vol. 4, pp. e35, 2008. http://dx.doi.org/10.1371/journal.ppat.0040035
- M. Mikulska, V. Del Bono, S. Ratto, and C. Viscoli, "Occurrence, presentation and treatment of candidemia", Expert Review of Clinical Immunology, vol. 8, pp. 755-765, 2012. http://dx.doi.org/10.1586/eci.12.52
- S.I. Blot, K.H. Vandewoude, E.A. Hoste, and F.A. Colardyn, "Effects of nosocomial candidemia on outcomes of critically ill patients", The American Journal of Medicine, vol. 113, pp. 480-485, 2002. http://dx.doi.org/10.1016/S0002-9343(02)01248-2
- L. Laprade, V.L. Boyartchuk, W.F. Dietrich, and F. Winston, "Spt3 plays opposite roles in filamentous growth in Saccharomyces cerevisiae and Candida albicans and is required for C. albicans virulence.", Genetics, 2002. http://www.ncbi.nlm.nih.gov/pubmed/12072450
- B.R. Braun, W.S. Head, M.X. Wang, and A.D. Johnson, "Identification and characterization of TUP1-regulated genes in Candida albicans.", Genetics, 2000. http://www.ncbi.nlm.nih.gov/pubmed/10978273
- M. Uhl, "Haploinsufficiency-based large-scale forward genetic analysis of filamentous growth in the diploid human fungal pathogen C.albicans", The EMBO Journal, vol. 22, pp. 2668-2678, 2003. http://dx.doi.org/10.1093/emboj/cdg256
- E. Leberer, D. Harcus, D. Dignard, L. Johnson, S. Ushinsky, D.Y. Thomas, and K. Schröppel, "Ras links cellular morphogenesis to virulence by regulation of the MAP kinase and cAMP signalling pathways in the pathogenic fungus Candida albicans", Molecular Microbiology, vol. 42, pp. 673-687, 2001. http://dx.doi.org/10.1046/j.1365-2958.2001.02672.x
- W.M. Bell, and W.L. Chaffin, "Effect of yeast growth conditions on yeast-mycelial transition in Candida albicans.", Mycopathologia, 1983. http://www.ncbi.nlm.nih.gov/pubmed/6369144
- R.S. Shapiro, P. Uppuluri, A.K. Zaas, C. Collins, H. Senn, J.R. Perfect, J. Heitman, and L.E. Cowen, "Hsp90 Orchestrates Temperature-Dependent Candida albicans Morphogenesis via Ras1-PKA Signaling", Current Biology, vol. 19, pp. 621-629, 2009. http://dx.doi.org/10.1016/j.cub.2009.03.017
- T. Klengel, W. Liang, J. Chaloupka, C. Ruoff, K. Schröppel, J.R. Naglik, S.E. Eckert, E.G. Mogensen, K. Haynes, M.F. Tuite, L.R. Levin, J. Buck, and F.A. Mühlschlegel, "Fungal Adenylyl Cyclase Integrates CO2 Sensing with cAMP Signaling and Virulence", Current Biology, vol. 15, pp. 2021-2026, 2005. http://dx.doi.org/10.1016/j.cub.2005.10.040
- E. Mattia, G. Carruba, L. Angiolella, and A. Cassone, "Induction of germ tube formation by N-acetyl-D-glucosamine in Candida albicans: uptake of inducer and germinative response.", Journal of bacteriology, 1982. http://www.ncbi.nlm.nih.gov/pubmed/6752114
- Q. Feng, E. Summers, B. Guo, and G. Fink, "Ras signaling is required for serum-induced hyphal differentiation in Candida albicans.", Journal of bacteriology, 1999. http://www.ncbi.nlm.nih.gov/pubmed/10515923
- L.L. Hoyer, T.L. Payne, M. Bell, A.M. Myers, and S. Scherer, "Candida albicans ALS3 and insights into the nature of the ALS gene family", Current Genetics, vol. 33, pp. 451-459, 1998. http://dx.doi.org/10.1007/s002940050359
- J.F. Staab, S.D. Bradway, P.L. Fidel, and P. Sundstrom, "Adhesive and Mammalian Transglutaminase Substrate Properties of Candida albicans Hwp1", Science, vol. 283, pp. 1535-1538, 1999. http://dx.doi.org/10.1126/science.283.5407.1535
- D.A. Bailey, P.J. Feldmann, M. Bovey, N.A. Gow, and A.J. Brown, "The Candida albicans HYR1 gene, which is activated in response to hyphal development, belongs to a gene family encoding yeast cell wall proteins.", Journal of bacteriology, 1996. http://www.ncbi.nlm.nih.gov/pubmed/8808922
- X. Zheng, Y. Wang, and Y. Wang, "Hgc1, a novel hypha-specific G1 cyclin-related protein regulates Candida albicans hyphal morphogenesis", The EMBO Journal, vol. 23, pp. 1845-1856, 2004. http://dx.doi.org/10.1038/sj.emboj.7600195
- N.A.R. Gow, F.L. van de Veerdonk, A.J.P. Brown, and M.G. Netea, "Candida albicans morphogenesis and host defence: discriminating invasion from colonization", Nature Reviews Microbiology, vol. 10, pp. 112-122, 2011. http://dx.doi.org/10.1038/nrmicro2711
- D.P. Bockmühl, and J.F. Ernst, "A potential phosphorylation site for an A-type kinase in the Efg1 regulator protein contributes to hyphal morphogenesis of Candida albicans.", Genetics, 2001. http://www.ncbi.nlm.nih.gov/pubmed/11290709
- L.S. Robertson, H.C. Causton, R.A. Young, and G.R. Fink, "The yeast A kinases differentially regulate iron uptake and respiratory function", Proceedings of the National Academy of Sciences, vol. 97, pp. 5984-5988, 2000. http://dx.doi.org/10.1073/pnas.100113397
- S. Pohlers, R. Martin, T. Krüger, D. Hellwig, F. Hänel, O. Kniemeyer, H.P. Saluz, P. Van Dijck, J.F. Ernst, A. Brakhage, F.A. Mühlschlegel, and O. Kurzai, "Lipid Signaling via Pkh1/2 Regulates Fungal CO 2 Sensing through the Kinase Sch9", mBio, vol. 8, 2017. http://dx.doi.org/10.1128/mBio.02211-16
- R.A. Hall, L. De Sordi, D.M. MacCallum, H. Topal, R. Eaton, J.W. Bloor, G.K. Robinson, L.R. Levin, J. Buck, Y. Wang, N.A.R. Gow, C. Steegborn, and F.A. Mühlschlegel, "CO2 Acts as a Signalling Molecule in Populations of the Fungal Pathogen Candida albicans", PLoS Pathogens, vol. 6, pp. e1001193, 2010. http://dx.doi.org/10.1371/journal.ppat.1001193
- X. Xu, R.T.H. Lee, H. Fang, Y. Wang, R. Li, H. Zou, Y. Zhu, and Y. Wang, "Bacterial Peptidoglycan Triggers Candida albicans Hyphal Growth by Directly Activating the Adenylyl Cyclase Cyr1p", Cell Host & Microbe, vol. 4, pp. 28-39, 2008. http://dx.doi.org/10.1016/j.chom.2008.05.014
- M.M. Maidan, L. De Rop, J. Serneels, S. Exler, S. Rupp, H. Tournu, J.M. Thevelein, and P. Van Dijck, "The G Protein-coupled Receptor Gpr1 and the Gα Protein Gpa2 Act through the cAMP-Protein Kinase A Pathway to Induce Morphogenesis inCandida albicans", Molecular Biology of the Cell, vol. 16, pp. 1971-1986, 2005. http://dx.doi.org/10.1091/mbc.e04-09-0780
- F.D. Ivey, and C.S. Hoffman, "Direct activation of fission yeast adenylate cyclase by the Gpa2 Gα of the glucose signaling pathway", Proceedings of the National Academy of Sciences, vol. 102, pp. 6108-6113, 2005. http://dx.doi.org/10.1073/pnas.0502270102
- J.M. Hollomon, N. Grahl, S.D. Willger, K. Koeppen, and D.A. Hogan, "Global Role of Cyclic AMP Signaling in pH-Dependent Responses in Candida albicans", mSphere, vol. 1, 2016. http://dx.doi.org/10.1128/mSphere.00283-16
- D.A. Hudson, Q.L. Sciascia, R.J. Sanders, G.E. Norris, P.J.B. Edwards, P.A. Sullivan, and P.C. Farley, "Identification of the dialysable serum inducer of germ-tube formation in Candida albicans", Microbiology, vol. 150, pp. 3041-3049, 2004. http://dx.doi.org/10.1099/mic.0.27121-0
- F. Rolland, J.H. De Winde, K. Lemaire, E. Boles, J.M. Thevelein, and J. Winderickx, "Glucose‐induced cAMP signalling in yeast requires both a G‐protein coupled receptor system for extracellular glucose detection and a separable hexose kinase‐dependent sensing process", Molecular Microbiology, vol. 38, pp. 348-358, 2000. http://dx.doi.org/10.1046/j.1365-2958.2000.02125.x
- R. Goswami, V. Dadhwal, S. Tejaswi, K. Datta, A. Paul, R. Haricharan, U. Banerjee, and N. Kochupillai, "Species-specific Prevalence of Vaginal Candidiasis Among Patients with Diabetes Mellitus and its Relation to their Glycaemic Status", Journal of Infection, vol. 41, pp. 162-166, 2000. http://dx.doi.org/10.1053/jinf.2000.0723
- J. Guggenheimer, P.A. Moore, K. Rossie, D. Myers, M.B. Mongelluzzo, H.M. Block, R. Weyant, and T. Orchard, "Insulin-dependent diabetes mellitus and oral soft tissue pathologies. II. Prevalence and characteristics of Candida and candidal lesions", Oral Surgery, Oral Medicine, Oral Pathology, Oral Radiology, and Endodontology, vol. 89, pp. 570-576, 2000. http://dx.doi.org/10.1067/moe.2000.104477
- A. Rodaki, I.M. Bohovych, B. Enjalbert, T. Young, F.C. Odds, N.A. Gow, and A.J. Brown, "Glucose Promotes Stress Resistance in the Fungal PathogenCandida albicans", Molecular Biology of the Cell, vol. 20, pp. 4845-4855, 2009. http://dx.doi.org/10.1091/mbc.e09-01-0002
- K. Biswas, and J. Morschhäuser, "The Mep2p ammonium permease controls nitrogen starvation‐induced filamentous growth in Candida albicans", Molecular Microbiology, vol. 56, pp. 649-669, 2005. http://dx.doi.org/10.1111/j.1365-2958.2005.04576.x
- H. Liu, J. Köhler, and G.R. Fink, "Suppression of Hyphal Formation in Candida albicans by Mutation of a STE12 Homolog", Science, vol. 266, pp. 1723-1726, 1994. http://dx.doi.org/10.1126/science.7992058
- S. Kimura, and Y. Fukuyama, "Tubular cytoplasmic inclusions in a case of childhood dermatomyositis with migratory subcutaneous nodules.", European journal of pediatrics, 1977. http://www.ncbi.nlm.nih.gov/pubmed/891572
- E. Leberer, D. Harcus, I.D. Broadbent, K.L. Clark, D. Dignard, K. Ziegelbauer, A. Schmidt, N.A. Gow, A.J. Brown, and D.Y. Thomas, "Signal transduction through homologs of the Ste20p and Ste7p protein kinases can trigger hyphal formation in the pathogenic fungus Candida albicans.", Proceedings of the National Academy of Sciences of the United States of America, 1996. http://www.ncbi.nlm.nih.gov/pubmed/8917571
- C. Csank, K. Schröppel, E. Leberer, D. Harcus, O. Mohamed, S. Meloche, D.Y. Thomas, and M. Whiteway, "Roles of the Candida albicans mitogen-activated protein kinase homolog, Cek1p, in hyphal development and systemic candidiasis.", Infection and immunity, 1998. http://www.ncbi.nlm.nih.gov/pubmed/9596738
- C. Sánchez-Martínez, and J. Pérez-Martín, "Gpa2, a G-Protein α Subunit Required for Hyphal Development in Candida albicans", Eukaryotic Cell, vol. 1, pp. 865-874, 2002. http://dx.doi.org/10.1128/EC.1.6.865-874.2002
- Y. Zhu, H. Fang, Y. Wang, G. Zeng, X. Zheng, and Y. Wang, "Ras1 and Ras2 play antagonistic roles in regulating cellular cAMP level, stationary‐phase entry and stress response in Candida albicans", Molecular Microbiology, vol. 74, pp. 862-875, 2009. http://dx.doi.org/10.1111/j.1365-2958.2009.06898.x
- B.R. Braun, and A.D. Johnson, "Control of filament formation in Candida albicans by the transcriptional repressor TUP1.", Science (New York, N.Y.), 1997. http://www.ncbi.nlm.nih.gov/pubmed/9204892
- A.A. Murad, "NRG1 represses yeast-hypha morphogenesis and hypha-specific gene expression in Candida albicans", The EMBO Journal, vol. 20, pp. 4742-4752, 2001. http://dx.doi.org/10.1093/emboj/20.17.4742
- B.R. Braun, "NRG1, a repressor of filamentous growth in C.albicans, is down-regulated during filament induction", The EMBO Journal, vol. 20, pp. 4753-4761, 2001. http://dx.doi.org/10.1093/emboj/20.17.4753
- R.A. Khalaf, and R.S. Zitomer, "The DNA binding protein Rfg1 is a repressor of filamentation in Candida albicans.", Genetics, 2001. http://www.ncbi.nlm.nih.gov/pubmed/11290707
- D. Kadosh, and A.D. Johnson, "Induction of theCandida albicansFilamentous Growth Program by Relief of Transcriptional Repression: A Genome-wide Analysis", Molecular Biology of the Cell, vol. 16, pp. 2903-2912, 2005. http://dx.doi.org/10.1091/mbc.e05-01-0073
- Y. Lu, C. Su, A. Wang, and H. Liu, "Hyphal Development in Candida albicans Requires Two Temporally Linked Changes in Promoter Chromatin for Initiation and Maintenance", PLoS Biology, vol. 9, pp. e1001105, 2011. http://dx.doi.org/10.1371/journal.pbio.1001105
- C.M. Hull, R.M. Raisner, and A.D. Johnson, "Evidence for Mating of the "Asexual" Yeast Candida albicans in a Mammalian Host", Science, vol. 289, pp. 307-310, 2000. http://dx.doi.org/10.1126/science.289.5477.307
- A. Forche, K. Alby, D. Schaefer, A.D. Johnson, J. Berman, and R.J. Bennett, "The Parasexual Cycle in Candida albicans Provides an Alternative Pathway to Meiosis for the Formation of Recombinant Strains", PLoS Biology, vol. 6, pp. e110, 2008. http://dx.doi.org/10.1371/journal.pbio.0060110
- R.J. Bennett, "Completion of a parasexual cycle in Candida albicans by induced chromosome loss in tetraploid strains", The EMBO Journal, vol. 22, pp. 2505-2515, 2003. http://dx.doi.org/10.1093/emboj/cdg235
- M.A. Hickman, G. Zeng, A. Forche, M.P. Hirakawa, D. Abbey, B.D. Harrison, Y. Wang, C. Su, R.J. Bennett, Y. Wang, and J. Berman, "The ‘obligate diploid’ Candida albicans forms mating-competent haploids", Nature, vol. 494, pp. 55-59, 2013. http://dx.doi.org/10.1038/nature11865
- I. Herskowitz, "Life cycle of the budding yeast Saccharomyces cerevisiae.", Microbiological reviews, 1988. http://www.ncbi.nlm.nih.gov/pubmed/3070323
- M.G. Miller, and A.D. Johnson, "White-Opaque Switching in Candida albicans Is Controlled by Mating-Type Locus Homeodomain Proteins and Allows Efficient Mating", Cell, vol. 110, pp. 293-302, 2002. http://dx.doi.org/10.1016/S0092-8674(02)00837-1
- D.R. Soil, "Gene regulation during high-frequency switching in Candida albicans.", Microbiology (Reading, England), 1997. http://www.ncbi.nlm.nih.gov/pubmed/9043104
- C. Pujol, K.J. Daniels, S.R. Lockhart, T. Srikantha, J.B. Radke, J. Geiger, and D.R. Soll, "The Closely Related Species Candida albicans and Candida dubliniensis Can Mate", Eukaryotic Cell, vol. 3, pp. 1015-1027, 2004. http://dx.doi.org/10.1128/ec.3.4.1015-1027.2004
- S.R. Lockhart, C. Pujol, K.J. Daniels, M.G. Miller, A.D. Johnson, M.A. Pfaller, and D.R. Soll, "In Candida albicans, white-opaque switchers are homozygous for mating type.", Genetics, 2002. http://www.ncbi.nlm.nih.gov/pubmed/12399384
- A.E. Tsong, M.G. Miller, R.M. Raisner, and A.D. Johnson, "Evolution of a Combinatorial Transcriptional Circuit", Cell, vol. 115, pp. 389-399, 2003. http://dx.doi.org/10.1016/S0092-8674(03)00885-7
- G. Huang, H. Wang, S. Chou, X. Nie, J. Chen, and H. Liu, "Bistable expression of WOR1 , a master regulator of white–opaque switching in Candida albicans", Proceedings of the National Academy of Sciences, vol. 103, pp. 12813-12818, 2006. http://dx.doi.org/10.1073/pnas.0605270103
- G. Huang, T. Srikantha, N. Sahni, S. Yi, and D.R. Soll, "CO2 Regulates White-to-Opaque Switching in Candida albicans", Current Biology, vol. 19, pp. 330-334, 2009. http://dx.doi.org/10.1016/j.cub.2009.01.018
- G. Huang, S. Yi, N. Sahni, K.J. Daniels, T. Srikantha, and D.R. Soll, "N-Acetylglucosamine Induces White to Opaque Switching, a Mating Prerequisite in Candida albicans", PLoS Pathogens, vol. 6, pp. e1000806, 2010. http://dx.doi.org/10.1371/journal.ppat.1000806
- R.E. Zordan, M.G. Miller, D.J. Galgoczy, B.B. Tuch, and A.D. Johnson, "Interlocking Transcriptional Feedback Loops Control White-Opaque Switching in Candida albicans", PLoS Biology, vol. 5, pp. e256, 2007. http://dx.doi.org/10.1371/journal.pbio.0050256
- M.B. Lohse, A.D. Hernday, P.M. Fordyce, L. Noiman, T.R. Sorrells, V. Hanson-Smith, C.J. Nobile, J.L. DeRisi, and A.D. Johnson, "Identification and characterization of a previously undescribed family of sequence-specific DNA-binding domains", Proceedings of the National Academy of Sciences, vol. 110, pp. 7660-7665, 2013. http://dx.doi.org/10.1073/pnas.1221734110
- R. Zarnowski, W.M. Westler, G.A. Lacmbouh, J.M. Marita, J.R. Bothe, J. Bernhardt, A. Lounes-Hadj Sahraoui, J. Fontaine, H. Sanchez, R.D. Hatfield, J.M. Ntambi, J.E. Nett, A.P. Mitchell, and D.R. Andes, "Novel Entries in a Fungal Biofilm Matrix Encyclopedia", mBio, vol. 5, 2014. http://dx.doi.org/10.1128/mBio.01333-14
- M. Martins, P. Uppuluri, D.P. Thomas, I.A. Cleary, M. Henriques, J.L. Lopez-Ribot, and R. Oliveira, "Presence of Extracellular DNA in the Candida albicans Biofilm Matrix and its Contribution to Biofilms", Mycopathologia, vol. 169, pp. 323-331, 2009. http://dx.doi.org/10.1007/s11046-009-9264-y
- D.P. Thomas, S.P. Bachmann, and J.L. Lopez-Ribot, "Proteomics for the analysis of theCandida albicans biofilm lifestyle", PROTEOMICS, vol. 6, pp. 5795-5804, 2006. http://dx.doi.org/10.1002/pmic.200600332
- J.W. Costerton, P.S. Stewart, and E.P. Greenberg, "Bacterial Biofilms: A Common Cause of Persistent Infections", Science, vol. 284, pp. 1318-1322, 1999. http://dx.doi.org/10.1126/science.284.5418.1318
- R. Donlan, "Biofilm Formation: A Clinically Relevant Microbiological Process", Clinical Infectious Diseases, vol. 33, pp. 1387-1392, 2001. http://dx.doi.org/10.1086/322972
- R. Donlan, "Biofilms and Device-Associated Infections", Emerging Infectious Diseases, vol. 7, pp. 277-281, 2001. http://dx.doi.org/10.3201/eid0702.010226
- J. Verma-Gaur, Y. Qu, P.F. Harrison, T.L. Lo, T. Quenault, M.J. Dagley, M. Bellousoff, D.R. Powell, T.H. Beilharz, and A. Traven, "Integration of Posttranscriptional Gene Networks into Metabolic Adaptation and Biofilm Maturation in Candida albicans", PLOS Genetics, vol. 11, pp. e1005590, 2015. http://dx.doi.org/10.1371/journal.pgen.1005590
- S. Fanning, and A.P. Mitchell, "Fungal Biofilms", PLoS Pathogens, vol. 8, pp. e1002585, 2012. http://dx.doi.org/10.1371/journal.ppat.1002585
- G. Ramage, S.P. Saville, D.P. Thomas, and J.L. López-Ribot, "CandidaBiofilms: an Update", Eukaryotic Cell, vol. 4, pp. 633-638, 2005. http://dx.doi.org/10.1128/ec.4.4.633-638.2005
- J. Chandra, D.M. Kuhn, P.K. Mukherjee, L.L. Hoyer, T. McCormick, and M.A. Ghannoum, "Biofilm Formation by the Fungal Pathogen Candida albicans : Development, Architecture, and Drug Resistance", Journal of Bacteriology, vol. 183, pp. 5385-5394, 2001. http://dx.doi.org/10.1128/JB.183.18.5385-5394.2001
- C. Nobile, E. Fox, J. Nett, T. Sorrells, Q. Mitrovich, A. Hernday, B. Tuch, D. Andes, and A. Johnson, "A Recently Evolved Transcriptional Network Controls Biofilm Development in Candida albicans", Cell, vol. 148, pp. 126-138, 2012. http://dx.doi.org/10.1016/j.cell.2011.10.048
- J.V. Desai, and A.P. Mitchell, "Candida albicans Biofilm Development and Its Genetic Control", Microbiology Spectrum, vol. 3, 2015. http://dx.doi.org/10.1128/microbiolspec.MB-0005-2014
- G. Ramage, K. VandeWalle, J.L. López-Ribot, and B.L. Wickes, "The filamentation pathway controlled by the Efg1 regulator protein is required for normal biofilm formation and development inCandida albicans", FEMS Microbiology Letters, vol. 214, pp. 95-100, 2002. http://dx.doi.org/10.1111/j.1574-6968.2002.tb11330.x
- C.J. Nobile, and A.P. Mitchell, "Regulation of Cell-Surface Genes and Biofilm Formation by the C. albicans Transcription Factor Bcr1p", Current Biology, vol. 15, pp. 1150-1155, 2005. http://dx.doi.org/10.1016/j.cub.2005.05.047
- Y. Lu, C. Su, O. Unoje, and H. Liu, "Quorum sensing controls hyphal initiation in Candida albicans through Ubr1-mediated protein degradation", Proceedings of the National Academy of Sciences, vol. 111, pp. 1975-1980, 2014. http://dx.doi.org/10.1073/pnas.1318690111
- J. R. Fortwendel, "Ras-Mediated Signal Transduction and Virulence in Human Pathogenic Fungi", Fungal Genomics & Biology, vol. 02, 2012. http://dx.doi.org/10.4172/2165-8056.1000105
- B.J. Park, K.A. Wannemuehler, B.J. Marston, N. Govender, P.G. Pappas, and T.M. Chiller, "Estimation of the current global burden of cryptococcal meningitis among persons living with HIV/AIDS", AIDS, vol. 23, pp. 525-530, 2009. http://dx.doi.org/10.1097/QAD.0b013e328322ffac
- . Centers for Disease Control and Prevention, "Global Fungal Diseases", Available at: https://www.cdc.gov/fungal/global/index.htm. Accessed 09.03.2017, 2017.
- M.S. Waugh, C.B. Nichols, C.M. DeCesare, G.M. Cox, J. Heitman, and J.A. Alspaugh, "Ras1 and Ras2 contribute shared and unique roles in physiology and virulence of Cryptococcus neoformans The GenBank accession number for the RAS2 sequence of C. neoformans H99 is AF294349.", Microbiology, vol. 148, pp. 191-201, 2002. http://dx.doi.org/10.1099/00221287-148-1-191
- J.A. Alspaugh, L.M. Cavallo, J.R. Perfect, and J. Heitman, "RAS1 regulates filamentation, mating and growth at high temperature of Cryptococcus neoformans", Molecular Microbiology, vol. 36, pp. 352-365, 2000. http://dx.doi.org/10.1046/j.1365-2958.2000.01852.x
- C.B. Nichols, Z.H. Perfect, and J.A. Alspaugh, "A Ras1‐Cdc24 signal transduction pathway mediates thermotolerance in the fungal pathogen Cryptococcus neoformans", Molecular Microbiology, vol. 63, pp. 1118-1130, 2006. http://dx.doi.org/10.1111/j.1365-2958.2006.05566.x
- E. Mylonakis, F.M. Ausubel, J.R. Perfect, J. Heitman, and S.B. Calderwood, "Killing of Caenorhabditis elegans by Cryptococcus neoformans as a model of yeast pathogenesis", Proceedings of the National Academy of Sciences, vol. 99, pp. 15675-15680, 2002. http://dx.doi.org/10.1073/pnas.232568599
- Y. Apidianakis, L.G. Rahme, J. Heitman, F.M. Ausubel, S.B. Calderwood, and E. Mylonakis, "Challenge of Drosophila melanogaster with Cryptococcus neoformans and Role of the Innate Immune Response", Eukaryotic Cell, vol. 3, pp. 413-419, 2004. http://dx.doi.org/10.1128/ec.3.2.413-419.2004
- E.R. Ballou, C.B. Nichols, K.J. Miglia, L. Kozubowski, and J.A. Alspaugh, "Two CDC42 paralogues modulate Cryptococcus neoformans thermotolerance and morphogenesis under host physiological conditions", Molecular Microbiology, vol. 75, pp. 763-780, 2010. http://dx.doi.org/10.1111/j.1365-2958.2009.07019.x
- E.R. Ballou, K. Selvig, J.L. Narloch, C.B. Nichols, and J.A. Alspaugh, "Two Rac paralogs regulate polarized growth in the human fungal pathogen Cryptococcus neoformans", Fungal Genetics and Biology, vol. 57, pp. 58-75, 2013. http://dx.doi.org/10.1016/j.fgb.2013.05.006
- E.R. Ballou, L. Kozubowski, C.B. Nichols, and J.A. Alspaugh, "Ras1 Acts through Duplicated Cdc42 and Rac Proteins to Regulate Morphogenesis and Pathogenesis in the Human Fungal Pathogen Cryptococcus neoformans", PLoS Genetics, vol. 9, pp. e1003687, 2013. http://dx.doi.org/10.1371/journal.pgen.1003687
- M.A. Vallim, C.B. Nichols, L. Fernandes, K.L. Cramer, and J.A. Alspaugh, "A Rac Homolog Functions Downstream of Ras1 To Control Hyphal Differentiation and High-Temperature Growth in the Pathogenic Fungus Cryptococcus neoformans", Eukaryotic Cell, vol. 4, pp. 1066-1078, 2005. http://dx.doi.org/10.1128/EC.4.6.1066-1078.2005
- L. Kozubowski, and J. Heitman, "Septins enforce morphogenetic events during sexual reproduction and contribute to virulence of Cryptococcus neoformans", Molecular Microbiology, vol. 75, pp. 658-675, 2010. http://dx.doi.org/10.1111/j.1365-2958.2009.06983.x
- G. Shen, E. Zhou, J.A. Alspaugh, and P. Wang, "Wsp1 Is Downstream of Cin1 and Regulates Vesicle Transport and Actin Cytoskeleton as an Effector of Cdc42 and Rac1 in Cryptococcus neoformans", Eukaryotic Cell, vol. 11, pp. 471-481, 2012. http://dx.doi.org/10.1128/EC.00011-12
- J.M. Johnson, M. Jin, and D.J. Lew, "Symmetry breaking and the establishment of cell polarity in budding yeast", Current Opinion in Genetics & Development, vol. 21, pp. 740-746, 2011. http://dx.doi.org/10.1016/j.gde.2011.09.007
- C.W. Gourlay, and K.R. Ayscough, "Identification of an upstream regulatory pathway controlling actin-mediated apoptosis in yeast", Journal of Cell Science, vol. 118, pp. 2119-2132, 2005. http://dx.doi.org/10.1242/jcs.02337
- C.W. Gourlay, and K.R. Ayscough, "Actin-Induced Hyperactivation of the Ras Signaling Pathway Leads to Apoptosis in Saccharomyces cerevisiae", Molecular and Cellular Biology, vol. 26, pp. 6487-6501, 2006. http://dx.doi.org/10.1128/MCB.00117-06
- J.E. Leadsham, K. Miller, K.R. Ayscough, S. Colombo, E. Martegani, P. Sudbery, and C.W. Gourlay, "Whi2p links nutritional sensing to actin-dependent Ras-cAMP-PKA regulation and apoptosis in yeast", Journal of Cell Science, vol. 122, pp. 706-715, 2009. http://dx.doi.org/10.1242/jcs.042424
- A.J. Phillips, J.D. Crowe, and M. Ramsdale, "Ras pathway signaling accelerates programmed cell death in the pathogenic fungus Candida albicans", Proceedings of the National Academy of Sciences, vol. 103, pp. 726-731, 2006. http://dx.doi.org/10.1073/pnas.0506405103
- J. Brajtburg, and J. Bolard, "Carrier effects on biological activity of amphotericin B.", Clinical microbiology reviews, 1996. http://www.ncbi.nlm.nih.gov/pubmed/8894350
- B.E. Cohen, "The Role of Signaling via Aqueous Pore Formation in Resistance Responses to Amphotericin B", Antimicrobial Agents and Chemotherapy, vol. 60, pp. 5122-5129, 2016. http://dx.doi.org/10.1128/AAC.00878-16
- P. Belenky, D. Camacho, and J. Collins, "Fungicidal Drugs Induce a Common Oxidative-Damage Cellular Death Pathway", Cell Reports, vol. 3, pp. 350-358, 2013. http://dx.doi.org/10.1016/j.celrep.2012.12.021
- Q. Al Abdallah, T.S. Norton, A.M. Hill, L.L. LeClaire, and J.R. Fortwendel, "A Fungus-Specific Protein Domain Is Essential for RasA-Mediated Morphogenetic Signaling in Aspergillus fumigatus", mSphere, vol. 1, 2016. http://dx.doi.org/10.1128/mSphere.00234-16
- A.E. Adams, D.I. Johnson, R.M. Longnecker, B.F. Sloat, and J.R. Pringle, "CDC42 and CDC43, two additional genes involved in budding and the establishment of cell polarity in the yeast Saccharomyces cerevisiae.", The Journal of cell biology, 1990. http://www.ncbi.nlm.nih.gov/pubmed/2195038
ACKNOWLEDGMENTS
This work is supported by a Kent Cancer Trust and University of Kent Graduate Teaching Assistant studentship to D.R.Pentland and an industrial CASE studentship to E.Piper-Brown.
COPYRIGHT
© 2017

Ras signalling in pathogenic yeasts by Pentland et al. is licensed under a Creative Commons Attribution 4.0 International License.