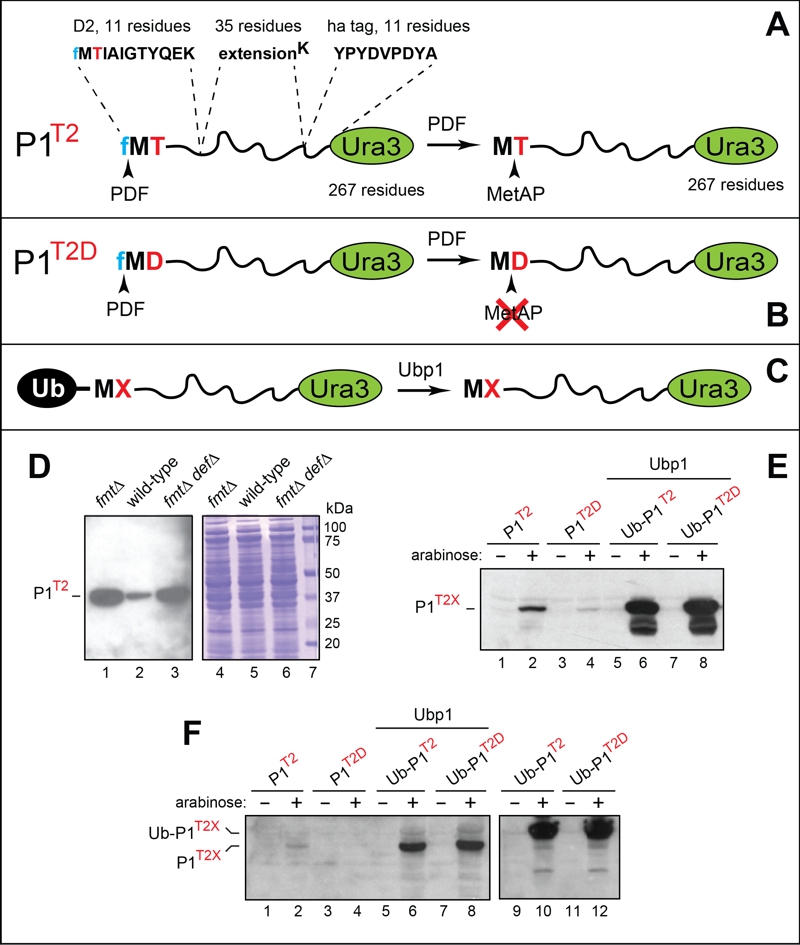
FIGURE 3: Analyses of reporter proteins in wild-type and formylation-lacking fmt E. coli.
(A) Design of the P1T2 (D21-11-eK-ha-Ura3) reporter protein. The term P1T2 (protein-1 containing Thr at position 2) denotes a chimeric reporter containing the indicated N‑terminal segments upstream of the 267-residue S. cerevisiae Ura3 moiety. Arrowheads indicate deformylation of N-terminal fMet by the PDF deformylase and the subsequent removal of Met by MetAP. See the main text for details.
(B) Same as in (A) but the otherwise identical P1T2D reporter contains Asp (D) at position 2. The rate of PDF-mediated deformylation of N-terminal fMet with Asp at position 2 is at least 10-fold lower than the rate of deformylation with Thr at position 2. Another difference between P1T2 and P1T2D is the retention of N-terminal Met in P1T2D. See the main text for details and citations.
(C) The use of ubiquitin (Ub) fusions to generate P1T2 and P1T2D through the removal of the N-terminal Ub moiety by the S. cerevisiae Ubp1 deubiquitylase expressed in E. coli.
(D) Immunoblotting analyses, after SDS-PAGE, of the P1T2 reporter protein expressed in fmt (lane 1), wild-type (lane 2) and fmt def (lane 3) E. coli. Lanes 4-6, the corresponding total protein patterns (Coomassie staining). Lane 7, molecular mass markers.
(E) Immunoblotting analyses, after SDS-PAGE, of the P1T2 and P1T2D reporters expressed from the Para promoter in wild-type E. coli (lanes 1-4), and of the Ub fusions Ub-P1T2 and Ub-P1T2D in wild-type E. coli expressing the S. cerevisiae Ubp1 deubiquitylase (lanes 5-8).
(F) Same as in (E), but independent experiments, in addition to expressing Ub-P1T2 and Ub-P1T2D in the absence of coexpressed yeast Ubp1 (lanes 9-12). See the main text for details and citations.