
Regulation of extracellular vesicles for protein secretion in Aspergillus nidulans
This study reveals that Aspergillus nidulans boosts extracellular vesicle production when ER-trafficked enzymes are induced, uncovering how fungi remodel their secretome through vesicle-mediated secretion to adapt to changing environments and biofilm formation.

Transcriptomic response to different heme sources in Trypanosoma cruzi epimastigotes
This study uncovers how the Chagas disease parasite adapts to changes in heme, an essential molecule for its survival, providing transcriptional clues to heme metabolism and identifying a previously unreported heme-binding protein in T. cruzi.

Luminal acetylation of microtubules is not essential for Plasmodium berghei and Toxoplasma gondii survival
Acetylation of α-tubulin at lysine 40 is not essential for cytoskeletal stability in Plasmodium berghei or Toxoplasma gondii, suggesting redundancy and plasticity in microtubule regulation in these parasites.

The dual-site agonist for human M2 muscarinic receptors Iper-8-naphtalimide induces mitochondrial dysfunction in Saccharomyces cerevisiae
S. cerevisiae is a model to study human GPCRs. N-8-Iper, active against glioblastoma via M2 receptor, causes mitochondrial damage in yeast by binding Ste2, highlighting evolutionary conservation of GPCRs.

Integrative Omics reveals changes in the cellular landscape of peroxisome-deficient pex3 yeast cells
To uncover the consequences of peroxisome deficiency, we compared Saccharomyces cerevisiae wild-type with pex3 cells, which lack peroxisomes, employing quantitative proteomics and transcriptomics technologies.

The mechanism of Tat-dependent protein translocation
Brüser and SandersThis review integrates mechanistically relevant biochemical, molecular, and structural studies on Tat-dependent translocation of folded proteins into an in its molecular detail new comprehensive explanation of how the Tat system mediates protein transport.

TOR-dependent regulation of the yeast homolog of the juvenile Batten Disease-associated gene CLN3
Pillalamarri et al.This study identifies conditions and genes that induce BTN1 expression in yeast. We show that BTN1 expression is regulated by translational control and by the mTOR1 pathway. An understanding of when and why BTN1 expression will aid in understanding the expression of CLN3, which may be helpful in the treatment of this devastating disease.

Overcoming phagocytosis resistance of hypervirulent Klebsiella pneumoniae by directly targeting capsules
Tsubaki et al.This study highlights a promising strategy for disarming hypervirulent K. pneumoniae by directly targeting its key virulence factors and provides novel insights into antibacterial therapeutic approaches against this clinically significant pathogen.
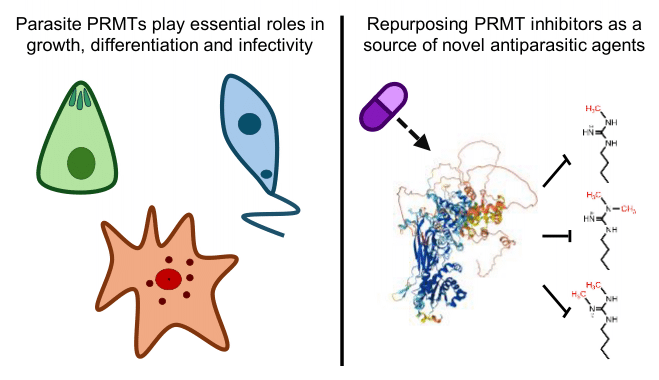
Protein arginine methyltransferases in protozoan parasites: a new path for antiparasitic chemotherapy?
Campagnaro et al.This review discusses the activity and the relevance of arginine methyltransferases for the survival of pathogenic kinetoplastids, apicomplexans and amoebas, and how these enzymes could be exploited as drug targets.

VapA/Scs2 sustains polarized growth in Aspergillus nidulans by maintaining AP-2-mediated apical endocytosis
Georgiou et al.To explore the functional significance of ER–PM contact sites in filamentous fungi, we identified and genetically characterized all Aspergillus nidulans proteins homologous to Snc2/VAP, Ist2, or tricalbins.

Genetic make-up and regulation of the L-lysine biosynthesis pathway in Vibrio natriegens
Straube et al.This study analysed the make-up and regulation of the biosynthetic pathway for L-lysine and related L-aspartate family amino acids (AFAAs) in Vibrio natriegens DSM759 to provide a comprehensive basis for future metabolic engineering endeavours aiming at developing this strain into an amino acid overproducer.
Yeast can express and assemble bacterial secretins in the mitochondrial outer membrane
Secretins, essential components of bacterial secretion systems, can be expressed in yeast and show differential dependencies on mitochondrial import and assembly factors for membrane integration, suggesting diverse pathways for their assembly into the bacterial outer membrane.
Metabolic reprogramming of Salmonella infected macrophages and its modulation by iron availability and the mTOR pathway
This article shows that iron plays a critical role in both the immune response and metabolic reprogramming of macrophages during infection, influencing the TCA cycle and mTOR pathway, with implications for the growth of intracellular bacteria like Salmonella.
Transcriptomic and chemogenomic analyses unveil the essential role of Com2-regulon in response and tolerance of Saccharomyces cerevisiae to stress induced by sulfur dioxide
This article shows that in the presence of sulfur dioxide (SO2), the transcription factor Com2 plays a critical role in the tolerance and response of Saccharomyces cerevisiae, affecting the expression of a majority of SO2-activated genes and contributing to the protection against stress induced by SO2 at an enologically relevant pH.
Network dynamics of the yeast methyltransferome
This article presents a systematic genetic analysis of methyltransferases (MTases) under normal and stress conditions, uncovering the complex and adaptive nature of the methyltransferome and discovering a potential connection between phospholipid methylation and histone methylation, suggesting interplay between lipid homeostasis and epigenetic regulation.
Functional link between mitochondria and Rnr3, the minor catalytic subunit of yeast ribonucleotide reductase
This article shows that the carbon source affects the abundance of ribonucleotide reductase (RNR) subunits in yeast, with a novel Mec1 signaling axis regulating Rnr3 independently of known DNA damage response pathways, and reveals Rnr3’s unexpected role in mitochondrial function.
Mitochondria-Associated Membranes (MAMs) are involved in Bax mitochondrial localization and cytochrome c release
This study investigated the role of Mitochondria-Associated Membranes (MAMs) in the regulation of apoptosis by analyzing the localization and function of the pro-apoptotic protein Bax in yeast, finding that disruption of MAMs by deletion of the MDM34 gene affects Bax’s mitochondrial localization and the release of cytochrome c.
Chlamydia pneumoniae is present in the dental plaque of periodontitis patients and stimulates an inflammatory response in gingival epithelial cells
This study found that Chlamydia pneumoniae is present more frequently in the dental plaque of individuals with periodontal disease, can invade human gingival epithelial cells causing inflammatory responses, and may therefore be a contributing factor to periodontal disease and a potential indicator of risk.
Breaking the bad: Bacillus blocks fungal virulence factors
Mayer and KronstadThis article comments on work published by Mayer & Kronstad (mBio, 2017), which identified the soil bacterium, Bacillus safensis as a potent inhibitor of virulence factor production by two major fungal pathogens of humans, Cryptococcus neoformans, and Candida albicans.
The integrated stress response in budding yeast lifespan extension
PostnikoffThis article summarizes how the budding yeast Saccharomyces cerevisiae has been instrumental in unraveling the molecular and cellular determinants of aging, and how the induction of cellular stress responses has been associated with experimental lifespan extension, thus underscoring the value of yeast as a model for developing potential aging therapies for humans.
Macrophages as drivers of an opportunistic infection
Vergunst et al.This article comments on work published by Mesureur et al. (PloS Pathog, 2017), which shows that macrophages are essential for proliferation of B. cenocepacia in the host. This suggests a new paradigm for Bcc infections and urges the development of novel anti-infectious therapies to efficiently disarm these intrinsically antibiotic resistant facultative intracellular pathogens.
Exacerbating and reversing lysosomal storage diseases: from yeast to humans
RajakumarThis article summarizes the use of yeast models in advancing our understanding of lysosomal storage diseases (LSDs), where they have been instrumental in researching LSD mechanisms, screening for therapeutic compounds, and exploring genetic and gene-environment interactions relevant to diseases like Batten disease, cystinosis, and Niemann-Pick type C disease, as well as their connection to broader health issues such as viral infections and obesity.
Live fast, die fast principle in a single cell of fission yeast
NakaokaThis article comments on a recent study (Nakaoka and Wakamoto, PLoS Biol, 2017), which developed a microfluidics-based platform to track multiple single cell lineages until death.
Out with the old: Hsp90 finds amino acid residue more useful than co-chaperone protein
Zuehlke and NeckersThis article comments on work published by Zuehlke et al (Nat Commun, 2017), which demonstrates that the function of one co-chaperone in yeast is replaced by posttranslational modification (PTM) of a single amino acid within Hsp90 in higher eukaryotes.
Having your cake and eating it – Staphylococcus aureus small colony variants can evolve faster growth rate without losing their antibiotic resistance
Gerrit Brandis et al.This article comments on work published by Cao et al. (mBio, 2027), which shows that Staphylococcus aureus can produce small colony variants (SCVs) that are challenging to detect and lead to persistent infections due to mutations affecting respiration and ATP production, with recent findings indicating various evolutionary paths for SCVs to increase growth rate while maintaining antibiotic resistance, suggesting greater adaptability and clinical challenge.
Transceptors as a functional link of transporters and receptors
George DiallinasA relative newcomer in environment sensing are the so called transceptors, membrane proteins that possess both solute transport and receptor-like signaling activities. Now, the transceptor concept is further enlarged to include micronutrient sensing via the iron and zinc high-affinity transporters of Saccharomyces cerevisiae.
S. pombe placed on the prion map
Jacqueline HaylesThis article comments on work published by Sideri et al. (Microbial Cell, 2017), which identified the Ctr4 prion in S. pombe.
Using microbes as a key tool to unravel the mechanism of autophagy and the functions of the ATG proteins
Mario Mauthe and Fulvio ReggioriMicrobes have served to discover and characterize unconventional functions of the ATG proteins, which are uncoupled from their role in autophagy. In our recent study, we have taken advantage of viruses as a screening tool to determine the extent of the unconventional functions of the ATG proteome and characterize one of them.
Autophagy: one more Nobel Prize for yeast
Zimmermann et al.The recent announcement of the 2016 Nobel Prize in Physiology or Medicine, awarded to Yoshinori Ohsumifor the discoveries of mechanisms governing autophagy, underscores the importance of intracellular degradation and recycling. Here we provide a quick historical overview that mirrors both the importance of autophagy as a conserved and essential process for cellular life and death as well as the crucial role of yeast in its mechanistic characterization.
Physiology, phylogeny, and LUCA
Martin et al.Genomes record their own history. But if we want to look all the way back to life’s beginnings some 4 billion years ago, the record of microbial evolution that is preserved in prokaryotic genomes is not easy to read. The classical approach has been to look for genes that are universally distributed. Another approach is to make all trees for all genes, and sift out the trees where signals have been overwritten by lateral gene transfer. What is left ought to be ancient. If we do that, what do we find?
Sexually transmitted infections: old foes on the rise
Carmona-Gutierrez et al.Sexually transmitted infections (STIs) are commonly spread via sexual contact. It is estimated that one million STIs are acquired every day worldwide. Besides their impact on sexual, reproductive and neonatal health, they can cause disastrous and life-threatening complications if left untreated. In addition to this personal burden, STIs also represent a socioeconomic problem, deriving in treatment costs of tremendous proportions. Despite a substantial progress in diagnosis, treatment and prevention, the incidence of many common STIs is increasing, and STIs continue to represent a global public health problem and a major cause for morbidity and mortality. With this Special Issue, Microbial Cell provides an in-depth overview of the eight major STIs, covering all relevant features of each infection.
Can’t find what you’re looking for?
You can browse all our issues and published articles here.
FAQs
Peer-reviewed, open-access research using unicellular organisms (and multicellular microorganisms) to understand cellular responses and human disease.
The journal (founded in 2014) is led by its Editors-in-Chief Frank Madeo, Didac Carmona-Gutierrez, and Guido Kroemer
Microbial Cell has been publishing original scientific literature since 2014, and from the very beginning has been managed by active scientists through an independent Publishing House (Shared science Publishers). The journal was conceived as a platform to acknowledge the importance of unicellular organisms, both as model systems as well as in the biological context of human health and disease.
Ever since, Microbial Cell has very positively developed and strongly grown into a respected journal in the unicellular research community and even beyond. This scientific impact is reflected in the yearly number of citations obtained by articles published in Microbial Cell, as recorded by the Web of Science (Clarivate, formerly Thomson/Reuters):
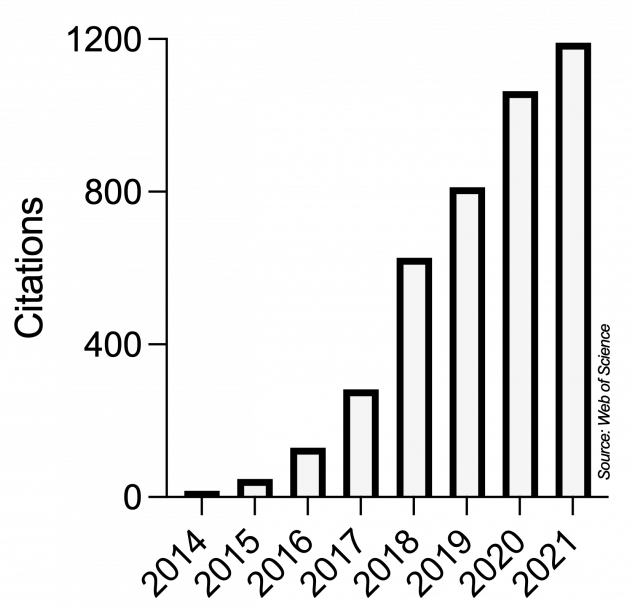
The scientific impact of Microbial Cell is also mirrored in a series of milestones:
2015: Microbial Cell is included in the Emerging Sources Citation Index (ESCI), a selection of developing journals drafted by Clarivate Analytics based on the candidate’s publishing standards, quality, editorial content, and citation data. Note: As an ESCI-selected journal, Microbial Cell is currently being evaluated in a rigorous and long process to determine an inclusion in the Science Citation Index Expanded (SCIE), which allows the official calculation of Clarivate Analytics’ impact factor.
2016: Microbial Cell is awarded the so-called DOAJ Seal by the selective Directory of Open Access Journals (DOAJ). The DOAJ Seal is an exclusive mark of certification for open access journals granted by DOAJ to journals that adhere to outstanding best practice and achieve an extra high and clear commitment to open access and high publishing standards.
2017: Microbial Cell is included in Pubmed Central (PMC), allowing the archiving of all the journal’s articles in PMC and PubMed.
2019: Microbial Cell is indexed in the prestigious abstract and citation database Scopus after a thorough selection process. This also means that Microbial Cell obtains, for the first time, an official Scopus CiteScore as well as an official journal ranking in the Scimago Journal and Country Ranking.
2022: Microbial Cell’s CiteScore reaches a value of 7.2 for the year 2021, positioning Microbial Cell among the top microbiology journals (previously available CiteScores: 2019: 5.4; 2020: 5.1).
2022: Microbial Cell is indexed in the highly selective Science Citation Index Expanded™, which covers approx. 9,500 of the world’s most impactful journals across 178 scientific disciplines. In their journal selection and curation process, Clarivate´s editors apply 24 ‘quality’ criteria and four ‘impact’ criteria to select the most influential journals in their respective fields. This selection is also a pre-requisite for inclusion in the JCR, which features the impact factor.
2022: Microbial Cell is listed in the Journal Citation Reports™ (JCR), and obtains its first official Journal Impact Factor™ (JIF) for the year 2021: 5.316.Check Article Types and Manuscript Preparation guidelines. Submit online via Scholastica.



Staphylococcus aureus type I signal peptidase: essential or not essential, that’s the question
Wouter L.W. Hazenbos et al.This article comments on work published by Morisaki et al. (mBio, 2016), which characterized a novel ABC transporter. This transporter apparently compensates for SpsB’s essential function by mediating alternative cleavage of a subset of proteins at a site distinct from the SpsB-cleavage site, leading to SpsB-independent secretion.