
Regulation of extracellular vesicles for protein secretion in Aspergillus nidulans
This study reveals that Aspergillus nidulans boosts extracellular vesicle production when ER-trafficked enzymes are induced, uncovering how fungi remodel their secretome through vesicle-mediated secretion to adapt to changing environments and biofilm formation.

Transcriptomic response to different heme sources in Trypanosoma cruzi epimastigotes
This study uncovers how the Chagas disease parasite adapts to changes in heme, an essential molecule for its survival, providing transcriptional clues to heme metabolism and identifying a previously unreported heme-binding protein in T. cruzi.

Luminal acetylation of microtubules is not essential for Plasmodium berghei and Toxoplasma gondii survival
Acetylation of α-tubulin at lysine 40 is not essential for cytoskeletal stability in Plasmodium berghei or Toxoplasma gondii, suggesting redundancy and plasticity in microtubule regulation in these parasites.

The dual-site agonist for human M2 muscarinic receptors Iper-8-naphtalimide induces mitochondrial dysfunction in Saccharomyces cerevisiae
S. cerevisiae is a model to study human GPCRs. N-8-Iper, active against glioblastoma via M2 receptor, causes mitochondrial damage in yeast by binding Ste2, highlighting evolutionary conservation of GPCRs.

Integrative Omics reveals changes in the cellular landscape of peroxisome-deficient pex3 yeast cells
To uncover the consequences of peroxisome deficiency, we compared Saccharomyces cerevisiae wild-type with pex3 cells, which lack peroxisomes, employing quantitative proteomics and transcriptomics technologies.

Regulation of extracellular vesicles for protein secretion in Aspergillus nidulans
Rebekkah E. Pope1, Patrick Ballmann2, Lisa Whitworth3 and Rolf A. Prade1,*
This study reveals that Aspergillus nidulans boosts extracellular vesicle production when ER-trafficked enzymes are induced, uncovering how fungi remodel their secretome through vesicle-mediated secretion to adapt to changing environments and biofilm formation.

Transcriptomic response to different heme sources in Trypanosoma cruzi epimastigotes
Evelyn Tevere1,a, María G. Mediavilla1,a, Cecilia B. Di Capua1, Marcelo L. Merli1, Carlos Robello2,3, Luisa Berná2,4 and Julia A. Cricco
This study uncovers how the Chagas disease parasite adapts to changes in heme, an essential molecule for its survival, providing transcriptional clues to heme metabolism and identifying a previously unreported heme-binding protein in T. cruzi.
Sir2 regulates selective autophagy in stationary-phase yeast cells
Ji-In Ryua, Juhye Junga, and Jeong-Yoon Kim
This study establishes Sir2 as a previously unrecognized regulator of selective autophagy during the stationary phase and highlight how cells dynamically control organelle degradation.
The transcription factors ADR1 or CAT8 are required for RTG pathway activation and evasion from yeast acetic acid-induced programmed cell death in raffinose
Luna Laera1,#, Nicoletta Guaragnella1,#, Maša Ždralević1,¶, Domenico Marzulli1, Zhengchang Liu2 and Sergio Giannattasio1
Yeast Saccharomyces cerevisiae grown on glucose undergoes programmed cell death (PCD) induced by acetic acid (AA-PCD), but evades PCD when grown in raffinose. This is due to concomitant relief of carbon catabolite repression (CCR) and activation of mitochondrial retrograde signaling. In this work, we investigated the relationships between the RTG and CCR pathways in the modulation of AA-PCD sensitivity under glucose repression or de-repression conditions. Our data show that simultaneous mitochondrial retrograde pathway activation and SNF1-dependent relief of CCR have a key role in central carbon metabolism reprogramming which modulates the yeast acetic acid-stress response.
The ubiquitin-conjugating enzyme, Ubc1, indirectly regulates SNF1 kinase activity via Forkhead-dependent transcription
Rubin Jiao1, Liubov Lobanova1, Amanda Waldner1, Anthony Fu1, Linda Xiao1, Troy A. Harkness1, and Terra G. Arnason1,2
The SNF1 kinase class of serine/threonine kinases, which includes the AMP-dependent protein kinase (AMPK) in other systems, are of widespread interest because of their important roles in glucose homeostasis, stress resistance, and aging. Our goal was to identify discrete ubiquitin-conjugating enzymes that are involved in SNF1 kinase activity in response to glucose levels and anticipated revealing those which are involved in Snf1-Ub attachment. Here, we report that the cell cycle and stress-related E2, Ubc1, indirectly affects SNF1 kinase activity not through stability, but through upstream events.
Phylogenetic profiles of all membrane transport proteins of the malaria parasite highlight new drug targets
January Weiner 3rd1 and Taco W.A. Kooij2
In order to combat the on-going malaria epidemic, discovery of new drug targets remains vital. Proteins that are essential to survival and specific to malaria parasites are key candidates. Here, we present a comprehensive orthology assignment of all Plasmodium falciparum putative membrane transport proteins and provide a detailed overview of the associated essential gene functions obtained through experimental genetics studies in human and murine model parasites.
VDAC regulates AAC-mediated apoptosis and cytochrome c release in yeast
Dário Trindade1,2, Clara Pereira3,4, Susana R. Chaves1, Stéphen Manon2, Manuela Côrte-Real1 and Maria João Sousa1
Mitochondrial outer membrane permeabilization is a key event in apoptosis processes leading to the release of lethal factors. In this study, we sought to determine whether Por1p functionally interacts with ADP/ATP carrier (AAC) proteins, as well as its contribution to cytochrome c release and yeast apoptosis induced by acetic acid treatment. Our data indicate that Por1p may regulate cell survival by acting as a negative regulator of AAC proteins in the apoptotic cascade.
Attenuation of polyglutamine-induced toxicity by enhancement of mitochondrial OXPHOS in yeast and fly models of aging
Andrea L. Ruetenik1,2,3, Alejandro Ocampo1,2,3,¶, Kai Ruan4,5,#, Yi Zhu4,5, Chong Li4,6, R. Grace Zhai1,4,5,6 and Antoni Barrientos1,2,3,5
Defects in mitochondrial biogenesis and function are common in many neurodegenerative disorders, including Huntington’s disease (HD). We could shown that enhancement of mitochondrial biogenesis protects against neurodegeneration in HD yeast and fly models. Our results suggest that therapeutic interventions aiming at the enhancement of mitochondrial respiration and OXPHOS could reduce polyQ toxicity and delay disease onset.
Cox1 mutation abrogates need for Cox23 in cytochrome c oxidase biogenesis
Richard Dela Cruz1,2, Mi-Young Jeong1 and Dennis R. Winge1
Cox23 is a known conserved assembly factor for cytochrome c oxidase, although its role in cytochrome c oxidase (CcO) biogenesis remains unresolved. To gain additional insights into its role, we isolated spontaneous suppressors of the respiratory growth defect in cox23∆ yeast cells. In this report, we describe the isolation of a robust suppressor of the respiratory defect in cox23∆ cells that mapped to the mitochondrial-encoded Cox1 subunit.
Increased spontaneous recombination in RNase H2-deficient cells arises from multiple contiguous rNMPs and not from single rNMP residues incorporated by DNA polymerase epsilon
Anastasiya Epshtein1, Catherine J. Potenski2, and Hannah L. Klein1
Ribonucleotides (rNMPs) can become embedded in DNA from insertion by DNA polymerases, failure to remove Okazaki fragment primers, R-loops that can prime replication, and RNA/cDNA-mediated recombination. We report here that recombination is not stimulated by rNMPs incorporated by the replicative polymerase epsilon. Instead, recombination seems to be stimulated by multiple contiguous rNMPs, which may arise from R-loops or replication priming events.
Construction and evaluation of yeast expression networks by database-guided predictions
Katharina Papsdorf1,#, Siyuan Sima1,#, Gerhard Richter2, Klaus Richter1
DNA-Microarrays are powerful tools to obtain expression data on the genome-wide scale. We set out to define a way to cluster microarray data according to their expressional relationship and to obtain information on the significance of this clustering approach.
Optogenetic monitoring identifies phosphatidylthreonine-regulated calcium homeostasis in Toxoplasma gondii
Arunakar Kuchipudi1, Ruben D. Arroyo-Olarte1, Friederike Hoffmann1, Volker Brinkmann2, Nishith Gupta1, 2
Toxoplasma gondii is an obligate intracellular parasite, which inflicts acute as well as chronic infections in a wide range of warm-blooded vertebrates. Using an optogenetic sensor to monitor subcellular calcium in this model intracellular pathogen we found a novel regulatory function of phosphatidylthreonine in calcium signaling.
Gut microbiota and ankylosing spondylitis: current insights and future challenges
Andrei Lobiuc1, Liliana Groppa2, Lia Chislari2, Eugeniu Russu2,3, Marinela Homitchi2,3, Camelia Ciorescu2,3, Sevag Hamamah4, I. Codruta Bran1 and Mihai Covasa1
This review explores the growing role of gut microbiota in AS and its potential to reshape targeted treatment strategies and facilitate development of adjunct therapies to address disease onset and progression.
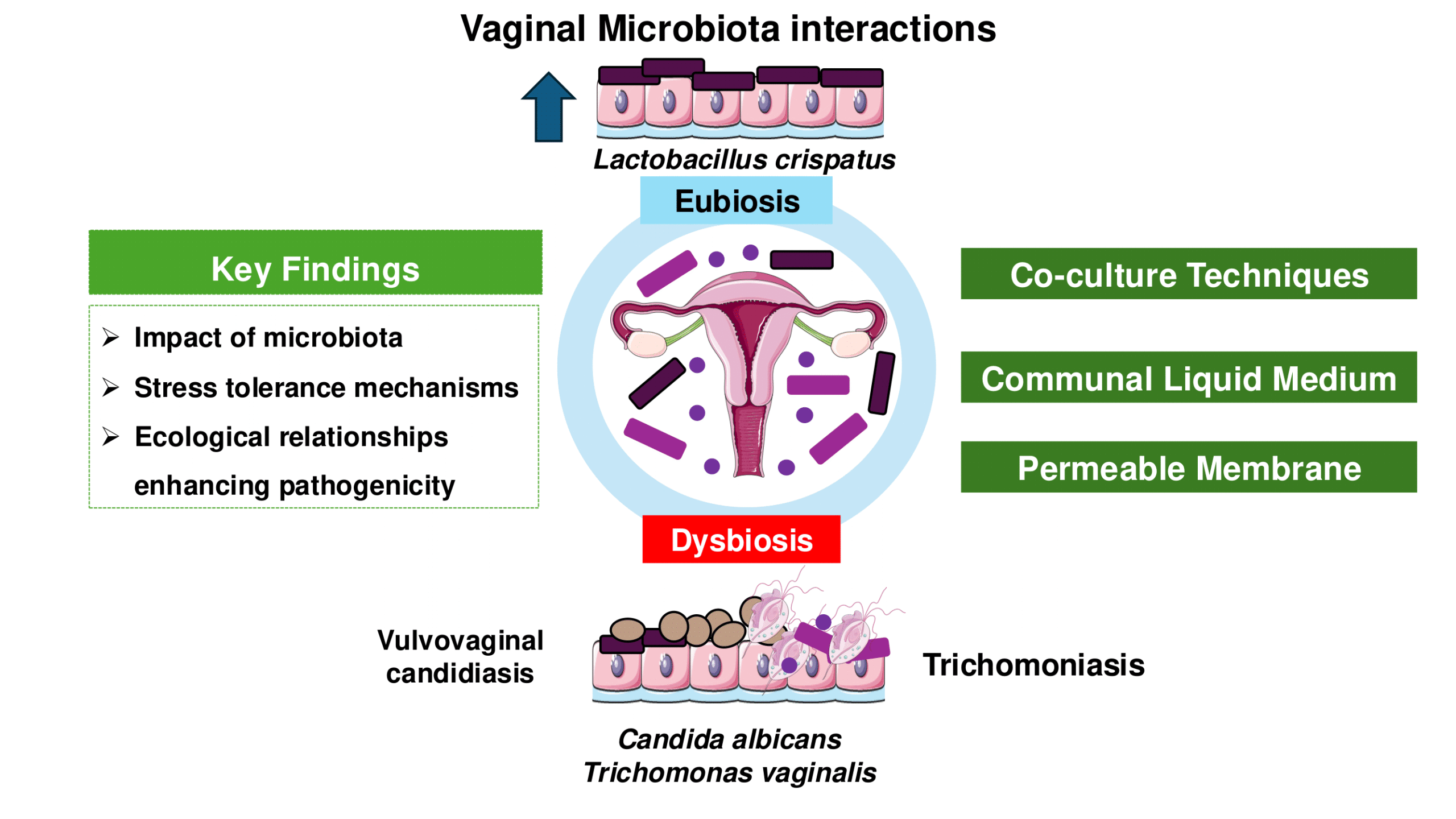
Advancements in vaginal microbiota, Trichomonas vaginalis, and vaginal cell interactions: Insights from co-culture assays
Fernanda Gomes Cardoso and Tiana Tasca
This review updates co-culture and co-incubation techniques for studying interactions of Lactobacillus spp., representing a pre-dominant member of the healthy vaginal microbiota; Candida spp., the most abundant yeast in the vagina, and T. vaginalis, responsible for the most widespread nonviral STI worldwide.
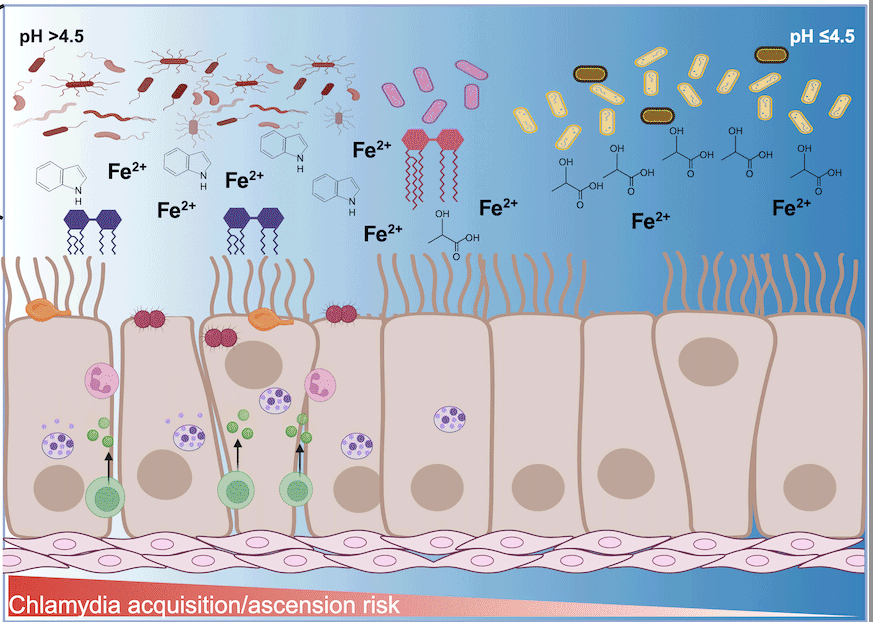
Influence of cervicovaginal microbiota on Chlamydia trachomatis infection dynamics
Emily Hand1, Indriati Hood-Pishchany1,2, Toni Darville1,2 and Catherine M. O’Connell2
This review examines the complex interplay between the cervicovaginal microbiome, C. trachomatis infection, and host immune responses, highlighting the role of metabolites such as short-chain and long-chain fatty acids, indole, and iron in modulating pathogen survival and host defenses.
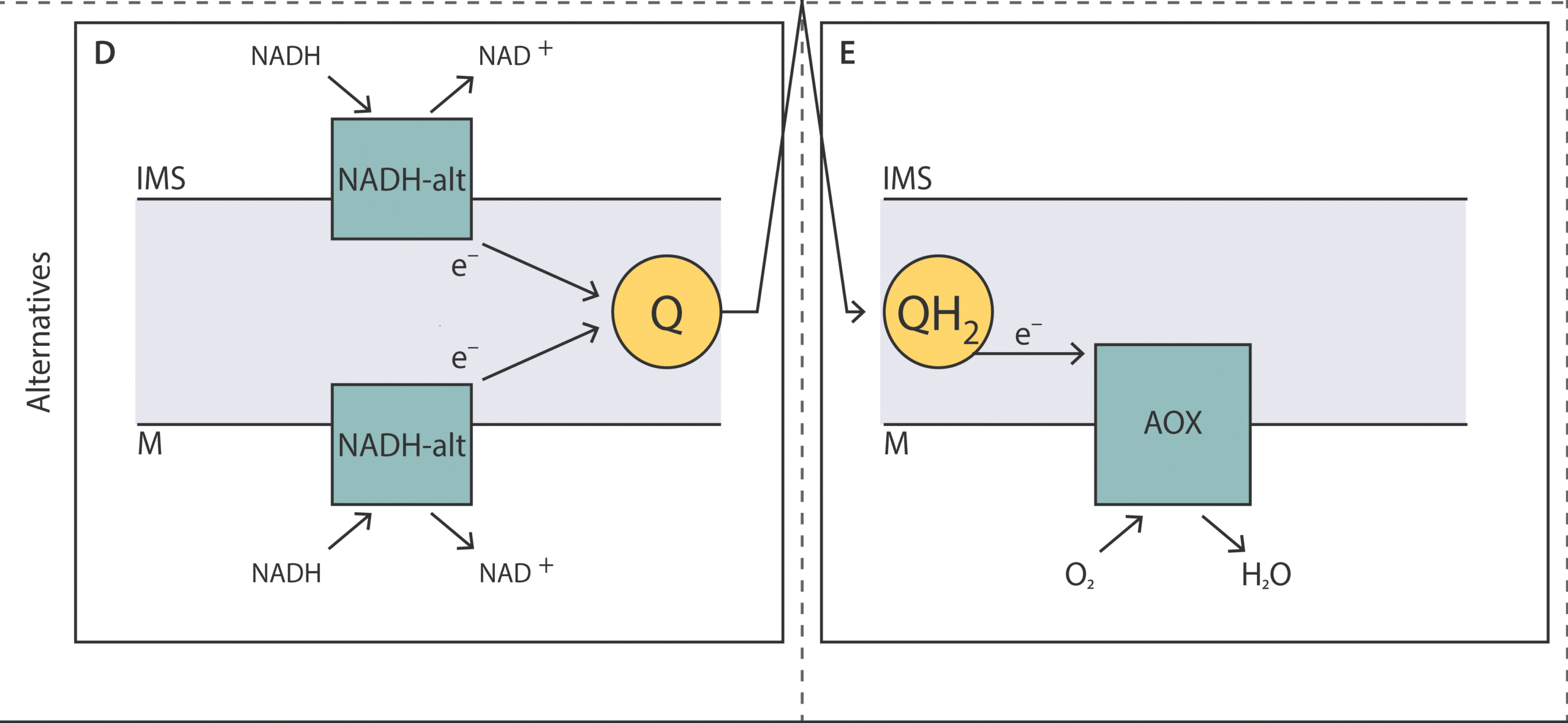
Unveiling the molecular architecture of the mitochondrial respiratory chain of Acanthamoeba castellanii
Christian Q. Scheckhuber1, Sutherland K. Maciver2 and Alvaro de Obeso Fernandez del Valle1
This review provides a comprehensive overview of the mitochondrial res-piratory chain in A. castellanii, focusing on the key alternative components involved in oxidative phosphorylation and their roles in energy metabolism, stress response, and adaptation to various conditions.

Paving the way for new antimicrobial peptides through molecular de-extinction
Karen O. Osiro1, Abel Gil-Ley2, Fabiano C. Fernandes1,3, Kamila B. S. de Oliveira2, Cesar de la Fuente-Nunez4-7, Octavio L. Franco1,2
The advancement of artificial intelligence and molecular de-extinction offers a valuable opportunity not only to discover new antimicrobials but also to provide accurate in silico predictions, thereby shortening the path to addressing the global antibiotic resistance crisis.

Efflux pumps: gatekeepers of antibiotic resistance in Staphylococcus aureus biofilms
Shweta Sinha1, Shifu Aggarwal2,3 and Durg Vijai Singh1
This review aims to elucidate the complex relationship between efflux pumps, antibiotic resistance and biofilm formation in S. aureus with the aim to aid in the development of potential therapeutic targets for combating S. aureus infections, especially those associated with biofilms.

Understanding the molecular mechanisms of human diseases: the benefits of fission yeasts
Lajos Acs-Szabo, Laszlo-Attila Papp and Ida Miklos
Here we collect the latest laboratory protocols and bioinformatics tools for the fission yeasts to highlight the many possibilities available to the research community. In addition, we present several limiting factors that everyone should be aware of when working with yeast models.
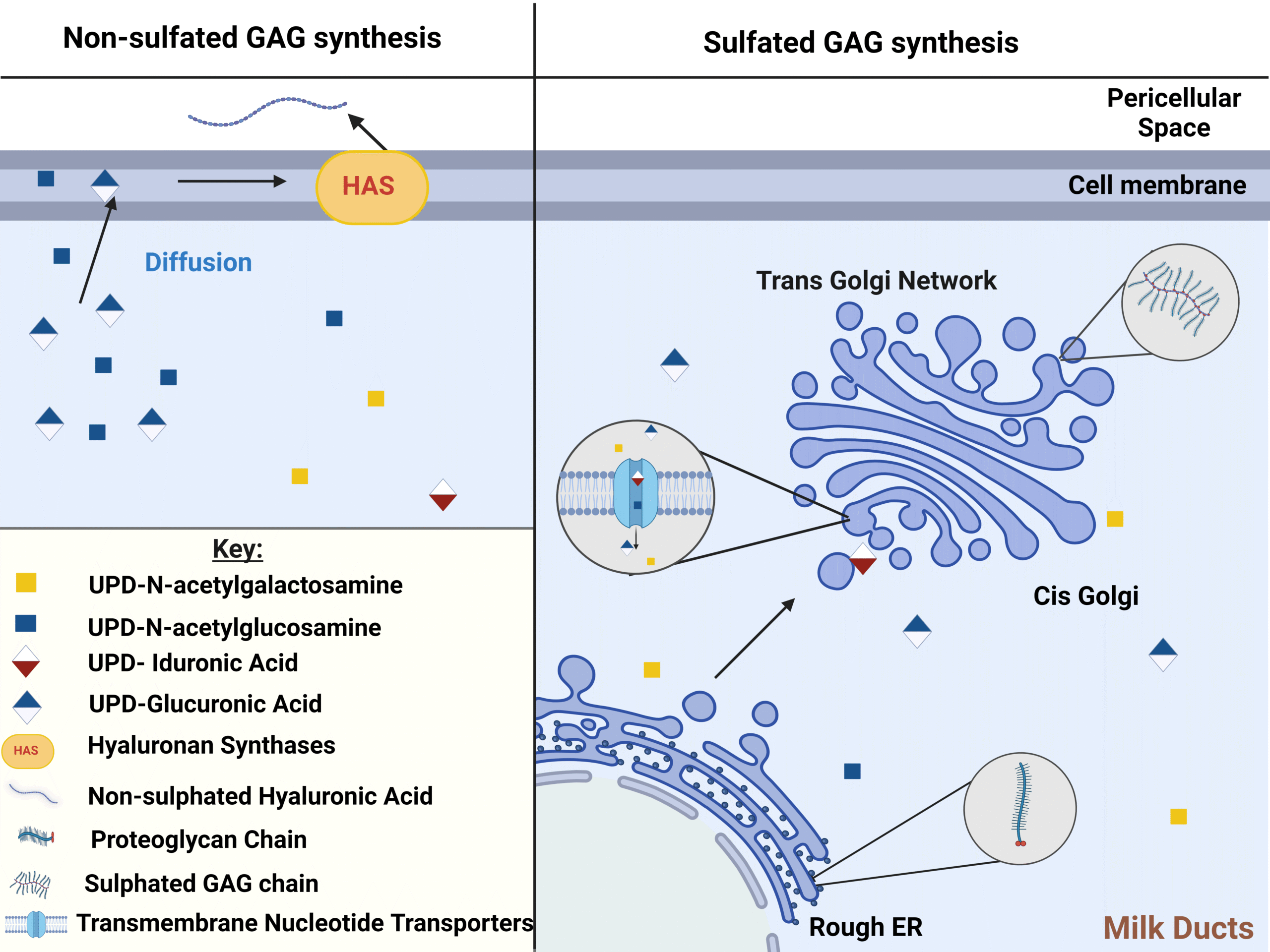
Characterising glycosaminoglycans in human breastmilk and their potential role in infant health
Melissa Greenwood1,2, Patricia Murciano-Martínez3, Janet Berrington4, Sabine L Flitsch5, Sean Austin2 and Christopher Stewart1
Glycosaminoglycans are bioactive components present in breast milk and play a potential key role in determining infant health yet are overlooked by many contemporary studies. This review explores their relevance, use and characterisation techniques.
Non-genetic impact factors on chronological lifespan and stress resistance of baker’s yeast
Michael Sauer and Diethard Mattanovich
This article comments on work published by Bisschops et al. (Microbial Cell, 2015), which illustrates how important the choice of the experimental setup is and how culture conditions influcence cellular aging and survival in biotechnological processes.
What’s old is new again: yeast mutant screens in the era of pooled segregant analysis by genome sequencing
Chris Curtin and Toni Cordente
This article comments on work published by Den Abt et al. (Microbial Cell, 2016), which identified genes involved in ethyl acetate formation in a yeast mutant screen based on a new approach combining repeated rounds of chemical mutagenesis and pooled segregant analysis by whole genome sequencing.
The complexities of bacterial-fungal interactions in the mammalian gastrointestinal tract
Eduardo Lopez-Medina1 and Andrew Y. Koh2
This article comments on work published by Lopez-Medina et al. (PLoS Pathog, 2015) and Fan et al. (Nat Med, 2015), which utilize an “artificial” niche, the antibiotic-treated gut with concomitant pathogenic microbe expansion, to gain insight in bacterial-fungal interactions in clinically common scenarios.
Gearing up for survival – HSP-containing granules accumulate in quiescent cells and promote survival
Ruofan Yu and Weiwei Dang
This article comments on work published by Lee et al. (Microbial Cell, 2016), which reports that distinct granules are formed in quiescent and non-quiescent cells, which determines their respective cell fates.
Yeast screening platform identifies FDA-approved drugs that reduce Aβ oligomerization
Triana Amen1,2 and Daniel Kaganovich1
This article comments on work published by Park et al. (Microbial Cell, 2016), which discovered a number of small molecules capable of modulating Aβ aggregation in a yeast model.
Groupthink: chromosomal clustering during transcriptional memory
Kevin A. Morano
In this article, the authors comment on the study “NO1 transcriptional memory leads to DNA zip code-dependent interchromosomal clustering.” by Brickner et al. (Microbial Cell, 2015), discussing the importance and molecular mechanisms of chromosomal clustering during transcriptional memory.
Yeast proteinopathy models: a robust tool for deciphering the basis of neurodegeneration
Amit Shrestha1, 2 and Lynn A. Megeney1, 2, 3
Protein quality control or proteostasis is an essential determinant of basic cell health and aging. Eukaryotic cells have evolved a number of proteostatic mechanisms to ensure that proteins retain functional conformation, or are rapidly degraded when proteins misfold or self-aggregate. This article discusses the use of budding yeast as a robust proxy to study the intersection between proteostasis and neurodegenerative disease.
Microbial Cell
is an open-access, peer-reviewed journal that publishes exceptionally relevant research works that implement the use of unicellular organisms (and multicellular microorganisms) to understand cellular responses to internal and external stimuli and/or human diseases.
you can trust
Can’t find what you’re looking for?
You can browse all our issues and published articles here.
FAQs
Peer-reviewed, open-access research using unicellular organisms (and multicellular microorganisms) to understand cellular responses and human disease.
The journal (founded in 2014) is led by its Editors-in-Chief Frank Madeo, Didac Carmona-Gutierrez, and Guido Kroemer
Microbial Cell has been publishing original scientific literature since 2014, and from the very beginning has been managed by active scientists through an independent Publishing House (Shared science Publishers). The journal was conceived as a platform to acknowledge the importance of unicellular organisms, both as model systems as well as in the biological context of human health and disease.
Ever since, Microbial Cell has very positively developed and strongly grown into a respected journal in the unicellular research community and even beyond. This scientific impact is reflected in the yearly number of citations obtained by articles published in Microbial Cell, as recorded by the Web of Science (Clarivate, formerly Thomson/Reuters):

The scientific impact of Microbial Cell is also mirrored in a series of milestones:
2015: Microbial Cell is included in the Emerging Sources Citation Index (ESCI), a selection of developing journals drafted by Clarivate Analytics based on the candidate’s publishing standards, quality, editorial content, and citation data. Note: As an ESCI-selected journal, Microbial Cell is currently being evaluated in a rigorous and long process to determine an inclusion in the Science Citation Index Expanded (SCIE), which allows the official calculation of Clarivate Analytics’ impact factor.
2016: Microbial Cell is awarded the so-called DOAJ Seal by the selective Directory of Open Access Journals (DOAJ). The DOAJ Seal is an exclusive mark of certification for open access journals granted by DOAJ to journals that adhere to outstanding best practice and achieve an extra high and clear commitment to open access and high publishing standards.
2017: Microbial Cell is included in Pubmed Central (PMC), allowing the archiving of all the journal’s articles in PMC and PubMed.
2019: Microbial Cell is indexed in the prestigious abstract and citation database Scopus after a thorough selection process. This also means that Microbial Cell obtains, for the first time, an official Scopus CiteScore as well as an official journal ranking in the Scimago Journal and Country Ranking.
2022: Microbial Cell’s CiteScore reaches a value of 7.2 for the year 2021, positioning Microbial Cell among the top microbiology journals (previously available CiteScores: 2019: 5.4; 2020: 5.1).
2022: Microbial Cell is indexed in the highly selective Science Citation Index Expanded™, which covers approx. 9,500 of the world’s most impactful journals across 178 scientific disciplines. In their journal selection and curation process, Clarivate´s editors apply 24 ‘quality’ criteria and four ‘impact’ criteria to select the most influential journals in their respective fields. This selection is also a pre-requisite for inclusion in the JCR, which features the impact factor.
2022: Microbial Cell is listed in the Journal Citation Reports™ (JCR), and obtains its first official Journal Impact Factor™ (JIF) for the year 2021: 5.316.Check Article Types and Manuscript Preparation guidelines. Submit online via Scholastica.


Similar environments but diverse fates: Responses of budding yeast to nutrient deprivation.
Saul M. Honigberg
Diploid budding yeast (Saccharomyces cerevisiae) can adopt one of several alternative differentiation fates in response to nutrient limitation, and each of these fates provides distinct biological functions. When different strain backgrounds are taken into account, these various fates occur in response to similar environmental cues, are regulated by the same signal transduction pathways, and share many of the same master regulators. I propose that the relationships between fate choice, environmental cues and signaling pathways are not Boolean, but involve graded levels of signals, pathway activation and master-regulator activity.