
Regulation of extracellular vesicles for protein secretion in Aspergillus nidulans
This study reveals that Aspergillus nidulans boosts extracellular vesicle production when ER-trafficked enzymes are induced, uncovering how fungi remodel their secretome through vesicle-mediated secretion to adapt to changing environments and biofilm formation.

Transcriptomic response to different heme sources in Trypanosoma cruzi epimastigotes
This study uncovers how the Chagas disease parasite adapts to changes in heme, an essential molecule for its survival, providing transcriptional clues to heme metabolism and identifying a previously unreported heme-binding protein in T. cruzi.

Luminal acetylation of microtubules is not essential for Plasmodium berghei and Toxoplasma gondii survival
Acetylation of α-tubulin at lysine 40 is not essential for cytoskeletal stability in Plasmodium berghei or Toxoplasma gondii, suggesting redundancy and plasticity in microtubule regulation in these parasites.

The dual-site agonist for human M2 muscarinic receptors Iper-8-naphtalimide induces mitochondrial dysfunction in Saccharomyces cerevisiae
S. cerevisiae is a model to study human GPCRs. N-8-Iper, active against glioblastoma via M2 receptor, causes mitochondrial damage in yeast by binding Ste2, highlighting evolutionary conservation of GPCRs.

Integrative Omics reveals changes in the cellular landscape of peroxisome-deficient pex3 yeast cells
To uncover the consequences of peroxisome deficiency, we compared Saccharomyces cerevisiae wild-type with pex3 cells, which lack peroxisomes, employing quantitative proteomics and transcriptomics technologies.

The mechanism of Tat-dependent protein translocation
Brüser and SandersThis review integrates mechanistically relevant biochemical, molecular, and structural studies on Tat-dependent translocation of folded proteins into an in its molecular detail new comprehensive explanation of how the Tat system mediates protein transport.

TOR-dependent regulation of the yeast homolog of the juvenile Batten Disease-associated gene CLN3
Pillalamarri et al.This study identifies conditions and genes that induce BTN1 expression in yeast. We show that BTN1 expression is regulated by translational control and by the mTOR1 pathway. An understanding of when and why BTN1 expression will aid in understanding the expression of CLN3, which may be helpful in the treatment of this devastating disease.

Overcoming phagocytosis resistance of hypervirulent Klebsiella pneumoniae by directly targeting capsules
Tsubaki et al.This study highlights a promising strategy for disarming hypervirulent K. pneumoniae by directly targeting its key virulence factors and provides novel insights into antibacterial therapeutic approaches against this clinically significant pathogen.
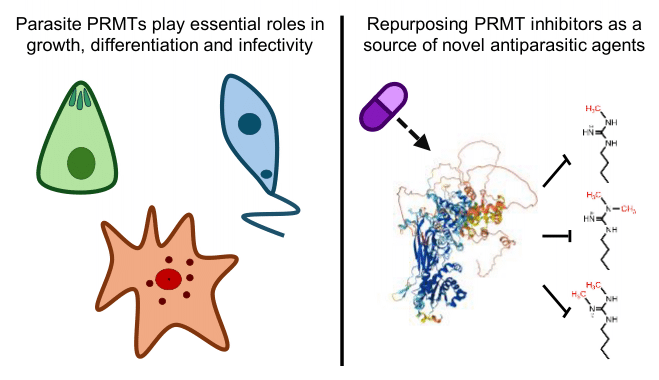
Protein arginine methyltransferases in protozoan parasites: a new path for antiparasitic chemotherapy?
Campagnaro et al.This review discusses the activity and the relevance of arginine methyltransferases for the survival of pathogenic kinetoplastids, apicomplexans and amoebas, and how these enzymes could be exploited as drug targets.

VapA/Scs2 sustains polarized growth in Aspergillus nidulans by maintaining AP-2-mediated apical endocytosis
Georgiou et al.To explore the functional significance of ER–PM contact sites in filamentous fungi, we identified and genetically characterized all Aspergillus nidulans proteins homologous to Snc2/VAP, Ist2, or tricalbins.

Genetic make-up and regulation of the L-lysine biosynthesis pathway in Vibrio natriegens
Straube et al.This study analysed the make-up and regulation of the biosynthetic pathway for L-lysine and related L-aspartate family amino acids (AFAAs) in Vibrio natriegens DSM759 to provide a comprehensive basis for future metabolic engineering endeavours aiming at developing this strain into an amino acid overproducer.
A central role for TOR signalling in a yeast model for juvenile CLN3 disease
Yeasts provide an excellent genetically tractable eukaryotic system for investigating the function of genes in their biological context, and are especially relevant for those conserved genes that cause disease. Bond et al. study the role of btn1, the orthologue of a human gene that underlies an early onset neurodegenerative disease (juvenile CLN3 disease, neuronal ceroid lipofuscinosis (NCLs) or Batten disease) in the fission yeast Schizosaccharomyces pombe.
Oxygen availability strongly affects chronological lifespan and thermotolerance in batch cultures of Saccharomyces cerevisiae
Stationary-phase (SP) batch cultures of Saccharomyces cerevisiae, in which growth has been arrested by carbon-source depletion, are widely applied to study chronological lifespan, quiescence and SP-associated robustness. Based on this type of experiments, typically performed under aerobic conditions, several roles of oxygen in aging have been proposed. However, SP in anaerobic yeast cultures has not been investigated in detail. Here, we use the unique capability of S. cerevisiae to grow in the complete absence of oxygen to directly compare SP in aerobic and anaerobic bioreactor cultures. This comparison revealed strong positive effects of oxygen availability on adenylate energy charge, longevity and thermotolerance during SP. A low thermotolerance of…
DNA damage checkpoint adaptation genes are required for division of cells harbouring eroded telomeres
In budding yeast, telomerase and the Cdc13p protein are two key players acting to ensure telomere stability. This article shows that while the capping process can be flexible, it takes a very specific genetic setup to allow a change from canonical capping to alternative capping.
The MAPKKKs Ste11 and Bck1 jointly transduce the high oxidative stress signal through the cell wall integrity MAP kinase pathway
Oxidative stress stimulates the Rho1 GTPase, which in turn induces the cell wall integrity (CWI) MAP kinase cascade. CWI activation promotes stress-responsive gene expression through activation of transcription factors (Rlm1, SBF) and nuclear release and subsequent destruction of the repressor cyclin C. This study reports that, in response to high hydrogen peroxide exposure, or in the presence of constitutively active Rho1, cyclin C still translocates to the cytoplasm and is degraded in cells lacking Bck1, the MAPKKK of the CWI pathway.
Formyl-methionine as a degradation signal at the N-termini of bacterial proteins
Varshavsky and colleagues solve a long-standing mystery in proteolysis! In bacteria, all nascent proteins bear the pretranslationally formed N-terminal formyl-methionine (fMet) residue. The fMet residue is cotranslationally deformylated by a ribosome-associated deformylase. The formylation of N-terminal Met in bacterial proteins is not strictly essential for either translation or cell viability. Moreover, protein synthesis by the cytosolic ribosomes of eukaryotes does not involve the formylation of N-terminal Met. What, then, is the main biological function of this metabolically costly, transient, and not strictly essential modification of N‑terminal Met, and why has Met formylation not been eliminated during bacterial evolution? One possibility is that the similarity of the formyl and acetyl groups, their identical locations in…
A single mutation in the 15S rRNA gene confers non sense suppressor activity and interacts with mRF1 the release factor in yeast mitochondria
This article presents the nucleotide sequence of the mim3-1 mitochondrial ribosomal suppressor, acting on ochre mitochondrial mutations and one frameshift mutation in Saccharomyces cerevisiae. A hypothetical mechanism of suppression by “ribosome shifting” is also discussed in view of the nature of mutations suppressed and not suppressed.
The lysosomotropic drug LeuLeu-OMe induces lysosome disruption and autophagy-independent cell death in Trypanosoma brucei
Trypanosoma brucei is a blood-borne, protozoan parasite that causes African sleeping sickness in humans and nagana in animals. The current chemotherapy relies on only a handful of drugs that display undesirable toxicity, poor efficacy and drug-resistance. In this study, we explored the use of lysosomotropic drugs to induce bloodstream form T. brucei cell death via lysosome destabilization. We measured drug concentrations that inhibit cell proliferation by 50% (IC50) for several compounds, chosen based on their lysosomotropic effects previously reported in Plasmodium falciparum. The lysosomal effects and cell death induced by L-leucyl-L-leucyl methyl ester (LeuLeu-OMe) were further analyzed by flow cytometry and immunofluorescence analyses of different lysosomal markers…
In Entamoeba histolytica, a BspA family protein is required for chemotaxis toward tumour necrosis factor
Background: Entamoeba histolytica cell migration is essential for the development of human amoebiasis (an infectious disease characterized by tissue invasion and destruction). The tissue inflammation associated with tumour necrosis factor (TNF) secretion by host cells is a well-documented feature of amoebiasis. Tumour necrosis factor is a chemoattractant for E. histolytica, and the parasite may have a TNF receptor at its cell surface. Methods: confocal microscopy, RNA Sequencing, bioinformatics, RNA antisense techniques and histological analysis of human colon explants were used to characterize the interplay between TNF and E. histolytica. Results: an antibody against human TNF receptor 1 (TNFR1) stained the E. histolytica trophozoite…
Microevolution of the pathogenic yeasts Candida albicans and Candida glabrata during antifungal therapy and host infection
Pais et al.This review explores how Candida albicans and Candida glabrata, common fungal pathogens resistant to antifungal therapy, adapt and evolve within different environments, aiming to identify stable adaptive mechanisms as potential drug targets.
The extracellular matrix of mycobacterial biofilms: could we shorten the treatment of mycobacterial infections?
Chakraborty and KumarThe article discusses the challenges presented by biofilms formed by non-tuberculous mycobacteria (NTM) species, which can lead to persistent infections that are difficult to treat due to phenotypic drug tolerance. The role of various cell wall components in mycobacterial biofilm formation is outlined, with a particular focus on Mycobacterium tuberculosis.
Guidelines for DNA recombination and repair studies: Cellular assays of DNA repair pathways
Klein et al.DNA recombination, repair and mutagenesis assays are powerful tools but each comes with its particular advantages and limitations. Here the most commonly used assays are reviewed, discussed, and presented as the guidelines for future studies.
Guidelines for DNA recombination and repair studies: Mechanistic assays of DNA repair processes
Klein et al.Mechanistic assays of DNA repair processes are a powerful tools but each comes with its particular advantages and limitations. Here the most commonly used assays are reviewed, discussed, and presented as the guidelines for future studies.
Imbalance in gut microbes from babies born to obese mothers increases gut permeability and myeloid cell adaptations that provoke obesity and NAFLD
Soderborg and FriedmanThis article comments on work published by Soderborg et al. (Nat Commun, 2018), which demonstrates a causative role of early life microbiome dysbiosis in infants born to mothers with obesity in novel pathways that promote developmental programming of NAFLD.
Retroviral integration site selection: a running Gag?
Lesbats and ParissiIn this article, the authors comment on the study “Structural basis for spumavirus GAG tethering to chromatin” by Lesbats et al. (Proc Natl Acad Sci, 2018) that revealed that the Gag protein of the spumaretrovirus prototype foamy virus (PFV) directly interacts with the nucleosome acidic patch, acting as a chromatin tether, and its disruption leads to delocalization of viral particles and integration sites, shedding light on the importance of retroviral structural proteins in the selection of integration sites.
Insights into the host-pathogen interaction: C. albicans manipulation of macrophage pyroptosis
O’Meara and CowenIn this article, the authors comment on the study “High-Throughput Screening Identifies Genes Required for Candida albicans Induction of Macrophage Pyroptosis” by O’Meara et al. (MBio, 2018) that provides a comprehensive analysis of the genetic circuitry in both Candida albicans and host macrophages that leads to pyroptosis, revealing the impact of altered pyroptosis on infection, the role of pyroptosis in facilitating neutrophil accumulation at the site of C. albicans infection, and the decoupling of inflammasome priming and activation in the response to C. albicans infection, thus shedding new light on the factors governing the outcomes of this interaction.
A comparative approach to decipher intestinal animal-microbe associations
NakashimaIn this article, the authors comment on the study “Chitin-based barrier immunity and its loss predated mucus-colonization by indigenous gut microbiota” by Nakashima et al. (Nat Commun, 2018) that used comparative analyses of chordates to investigate the development of animal-microbe associations, suggesting that microbial colonization of the mucus layer over mammalian gastrointestinal epithelium was established upon the loss of ancestral chitin-based barrier immunity, providing insights into the establishment of these associations in an evolutionary context.
Pathways of host cell exit by intracellular pathogens
Flieger et al.This review provides an overview of the diverse host cell exit strategies employed by intracellular-living bacterial, fungal, and protozoan pathogens, highlighting the commonalities and system-specific variations of these strategies, and discussing potential microbial molecules involved in host cell exit as targets for future intervention approaches.
The emerging role of complex modifications of tRNALysUUU in signaling pathways
Patrick C. Thiaville and Valérie de Crécy-LagardThis comment discusses the article “Loss of wobble uridine modification in tRNA anticodons interferes with TOR pathway signaling” by Scheidt et al (Microbial Cell, 2014).
Only functional localization is faithful localization
Roland LillThis article comments on work published by Peleh et al. (Microbial Cell 2014), which analyzes the localization of Dre2 in Saccharomyces cerevisiae.
One cell, one love: a journal for microbial research
Didac Carmona-Gutierrez et al.In this inaugural article of Microbial Cell, we highlight the importance of microbial research in general and the journal’s intention to serve as a publishing forum that supports and enfolds the scientific diversity in this area as it provides a unique, high-quality and universally accessible source of information and inspiration.
What’s the role of autophagy in trypanosomes?
Katherine Figarella and Néstor L. UzcáteguiThis article comments on Proto et al. (Microbial Cell, 2014), who report first insights into the molecular mechanism of autophagy in African trypanosomes by generating reporter bloodstream form cell lines.
Can’t find what you’re looking for?
You can browse all our issues and published articles here.
FAQs
Peer-reviewed, open-access research using unicellular organisms (and multicellular microorganisms) to understand cellular responses and human disease.
The journal (founded in 2014) is led by its Editors-in-Chief Frank Madeo, Didac Carmona-Gutierrez, and Guido Kroemer
Microbial Cell has been publishing original scientific literature since 2014, and from the very beginning has been managed by active scientists through an independent Publishing House (Shared science Publishers). The journal was conceived as a platform to acknowledge the importance of unicellular organisms, both as model systems as well as in the biological context of human health and disease.
Ever since, Microbial Cell has very positively developed and strongly grown into a respected journal in the unicellular research community and even beyond. This scientific impact is reflected in the yearly number of citations obtained by articles published in Microbial Cell, as recorded by the Web of Science (Clarivate, formerly Thomson/Reuters):
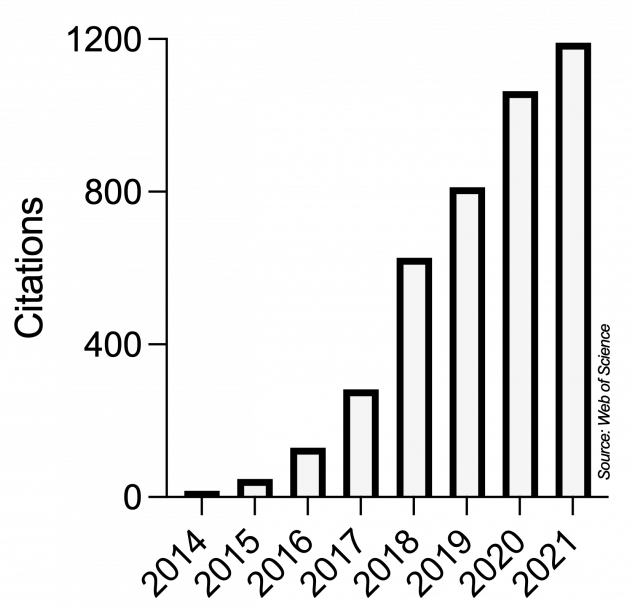
The scientific impact of Microbial Cell is also mirrored in a series of milestones:
2015: Microbial Cell is included in the Emerging Sources Citation Index (ESCI), a selection of developing journals drafted by Clarivate Analytics based on the candidate’s publishing standards, quality, editorial content, and citation data. Note: As an ESCI-selected journal, Microbial Cell is currently being evaluated in a rigorous and long process to determine an inclusion in the Science Citation Index Expanded (SCIE), which allows the official calculation of Clarivate Analytics’ impact factor.
2016: Microbial Cell is awarded the so-called DOAJ Seal by the selective Directory of Open Access Journals (DOAJ). The DOAJ Seal is an exclusive mark of certification for open access journals granted by DOAJ to journals that adhere to outstanding best practice and achieve an extra high and clear commitment to open access and high publishing standards.
2017: Microbial Cell is included in Pubmed Central (PMC), allowing the archiving of all the journal’s articles in PMC and PubMed.
2019: Microbial Cell is indexed in the prestigious abstract and citation database Scopus after a thorough selection process. This also means that Microbial Cell obtains, for the first time, an official Scopus CiteScore as well as an official journal ranking in the Scimago Journal and Country Ranking.
2022: Microbial Cell’s CiteScore reaches a value of 7.2 for the year 2021, positioning Microbial Cell among the top microbiology journals (previously available CiteScores: 2019: 5.4; 2020: 5.1).
2022: Microbial Cell is indexed in the highly selective Science Citation Index Expanded™, which covers approx. 9,500 of the world’s most impactful journals across 178 scientific disciplines. In their journal selection and curation process, Clarivate´s editors apply 24 ‘quality’ criteria and four ‘impact’ criteria to select the most influential journals in their respective fields. This selection is also a pre-requisite for inclusion in the JCR, which features the impact factor.
2022: Microbial Cell is listed in the Journal Citation Reports™ (JCR), and obtains its first official Journal Impact Factor™ (JIF) for the year 2021: 5.316.Check Article Types and Manuscript Preparation guidelines. Submit online via Scholastica.



Metabolic pathways further increase the complexity of cell size control in budding yeast
Jorrit M. EnserinkThis article comments on work published by Soma et al. (Microbial Cell, 2014), which teased apart the effect of metabolism and growth rate on setting of critical cell size in Saccharomyces cerevisiae.