
Regulation of extracellular vesicles for protein secretion in Aspergillus nidulans
This study reveals that Aspergillus nidulans boosts extracellular vesicle production when ER-trafficked enzymes are induced, uncovering how fungi remodel their secretome through vesicle-mediated secretion to adapt to changing environments and biofilm formation.

Transcriptomic response to different heme sources in Trypanosoma cruzi epimastigotes
This study uncovers how the Chagas disease parasite adapts to changes in heme, an essential molecule for its survival, providing transcriptional clues to heme metabolism and identifying a previously unreported heme-binding protein in T. cruzi.

Luminal acetylation of microtubules is not essential for Plasmodium berghei and Toxoplasma gondii survival
Acetylation of α-tubulin at lysine 40 is not essential for cytoskeletal stability in Plasmodium berghei or Toxoplasma gondii, suggesting redundancy and plasticity in microtubule regulation in these parasites.

The dual-site agonist for human M2 muscarinic receptors Iper-8-naphtalimide induces mitochondrial dysfunction in Saccharomyces cerevisiae
S. cerevisiae is a model to study human GPCRs. N-8-Iper, active against glioblastoma via M2 receptor, causes mitochondrial damage in yeast by binding Ste2, highlighting evolutionary conservation of GPCRs.

Integrative Omics reveals changes in the cellular landscape of peroxisome-deficient pex3 yeast cells
To uncover the consequences of peroxisome deficiency, we compared Saccharomyces cerevisiae wild-type with pex3 cells, which lack peroxisomes, employing quantitative proteomics and transcriptomics technologies.

TOR-dependent regulation of the yeast homolog of the juvenile Batten Disease-associated gene CLN3
Pillalamarri et al.This study identifies conditions and genes that induce BTN1 expression in yeast. We show that BTN1 expression is regulated by translational control and by the mTOR1 pathway. An understanding of when and why BTN1 expression will aid in understanding the expression of CLN3, which may be helpful in the treatment of this devastating disease.

Overcoming phagocytosis resistance of hypervirulent Klebsiella pneumoniae by directly targeting capsules
Tsubaki et al.This study highlights a promising strategy for disarming hypervirulent K. pneumoniae by directly targeting its key virulence factors and provides novel insights into antibacterial therapeutic approaches against this clinically significant pathogen.
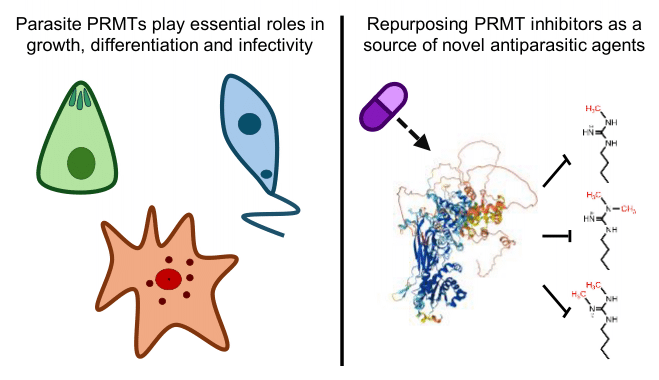
Protein arginine methyltransferases in protozoan parasites: a new path for antiparasitic chemotherapy?
Campagnaro et al.This review discusses the activity and the relevance of arginine methyltransferases for the survival of pathogenic kinetoplastids, apicomplexans and amoebas, and how these enzymes could be exploited as drug targets.

VapA/Scs2 sustains polarized growth in Aspergillus nidulans by maintaining AP-2-mediated apical endocytosis
Georgiou et al.To explore the functional significance of ER–PM contact sites in filamentous fungi, we identified and genetically characterized all Aspergillus nidulans proteins homologous to Snc2/VAP, Ist2, or tricalbins.

Genetic make-up and regulation of the L-lysine biosynthesis pathway in Vibrio natriegens
Straube et al.This study analysed the make-up and regulation of the biosynthetic pathway for L-lysine and related L-aspartate family amino acids (AFAAs) in Vibrio natriegens DSM759 to provide a comprehensive basis for future metabolic engineering endeavours aiming at developing this strain into an amino acid overproducer.

Regulation of extracellular vesicles for protein secretion in Aspergillus nidulans
Pope et al.This study reveals that Aspergillus nidulans boosts extracellular vesicle production when ER-trafficked enzymes are induced, uncovering how fungi remodel their secretome through vesicle-mediated secretion to adapt to changing environments and biofilm formation.

Transcriptomic response to different heme sources in Trypanosoma cruzi epimastigotes
Tevere et al.This study uncovers how the Chagas disease parasite adapts to changes in heme, an essential molecule for its survival, providing transcriptional clues to heme metabolism and identifying a previously unreported heme-binding protein in T. cruzi.
Sir2 regulates selective autophagy in stationary-phase yeast cells
Ryu et al.This study establishes Sir2 as a previously unrecognized regulator of selective autophagy during the stationary phase and highlight how cells dynamically control organelle degradation.
 Fasnacht et al.
Fasnacht et al.
Ampicillin treatment in persister cell studies may cause non-physiological artifacts
This study shows at the example of L2 how insufficient purification of ampicillin persister cells can lead to the generation of non-physiological artifacts and provides a novel tool to improve the removal of residual cell debris.

Ampicillin treatment in persister cell studies may cause non-physiological artifacts
Fasnacht et al.This study shows at the example of L2 how insufficient purification of ampicillin persister cells can lead to the generation of non-physiological artifacts and provides a novel tool to improve the removal of residual cell debris.
 Yao et al.
Yao et al.
Clostridium scindens promotes gallstone formation by inducing intrahepatic neutrophil extracellular traps through CXCL1 produced by colonic epithelial cells
Through in vivo and in vitro experiments, we validated the reliability of C. scindens stimulating colonic epithelial cells to produce TLR2, activating the NF-κB signaling pathway, promoting CXCL1 expres-sion, and inducing intrahepatic neutrophil NETosis, which may be associated with gallstone formation.

Clostridium scindens promotes gallstone formation by inducing intrahepatic neutrophil extracellular traps through CXCL1 produced by colonic epithelial cells
Yao et al.Through in vivo and in vitro experiments, we validated the reliability of C. scindens stimulating colonic epithelial cells to produce TLR2, activating the NF-κB signaling pathway, promoting CXCL1 expres-sion, and inducing intrahepatic neutrophil NETosis, which may be associated with gallstone formation.

Integrative Omics reveals changes in the cellular landscape of peroxisome-deficient pex3 yeast cells
To uncover the consequences of peroxisome deficiency, we compared Saccharomyces cerevisiae wild-type with pex3 cells, which lack peroxisomes, employing quantitative proteomics and transcriptomics technologies.

Integrative Omics reveals changes in the cellular landscape of peroxisome-deficient pex3 yeast cells
To uncover the consequences of peroxisome deficiency, we compared Saccharomyces cerevisiae wild-type with pex3 cells, which lack peroxisomes, employing quantitative proteomics and transcriptomics technologies.
 Mayer et al.
Mayer et al.
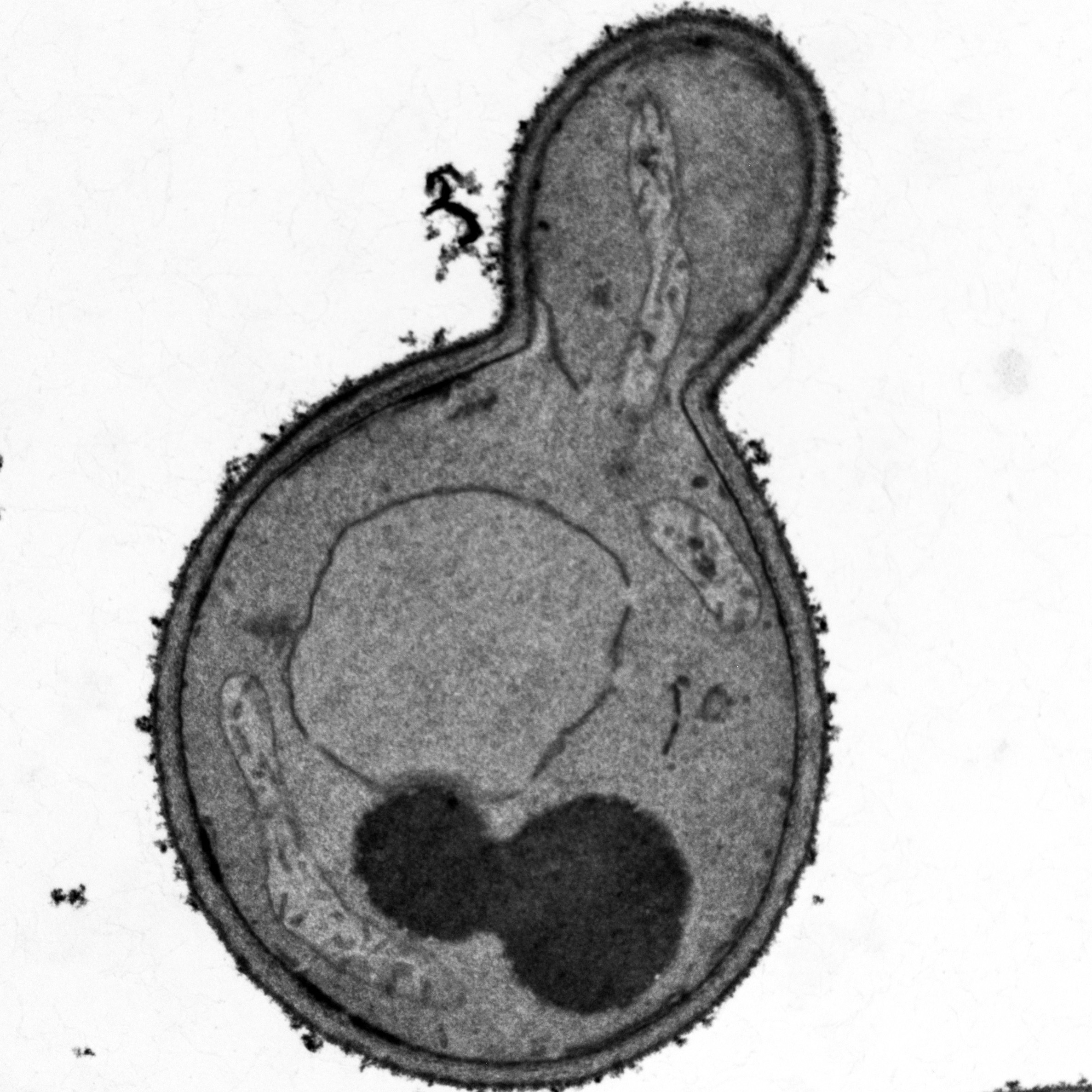
Microwave-assisted preparation of yeast cells for ultrastructural analysis by electron microscopy
Budding yeast Saccharomyces cerevisiae is widely used as a model organism to study the biogenesis and architecture of organellar membranes, which can be visualized by transmission electron microscopy (TEM).

Microwave-assisted preparation of yeast cells for ultrastructural analysis by electron microscopy
Mayer et al.Budding yeast Saccharomyces cerevisiae is widely used as a model organism to study the biogenesis and architecture of organellar membranes, which can be visualized by transmission electron microscopy (TEM).
 Vanetti et al.
Vanetti et al.
A complex remodeling of cellular homeostasis distinguishes RSV/SARS-CoV-2 co-infected A549-hACE2 expressing cell lines
Given the common tropism of SARS-CoV-2 and RSV, and the unclear consequences of their mutual influence, we developed an in vitro lung epithelial cell model to study the molecular mechanisms and cellular pathways modulated in viral co-infection.

A complex remodeling of cellular homeostasis distinguishes RSV/SARS-CoV-2 co-infected A549-hACE2 expressing cell lines
Vanetti et al.Given the common tropism of SARS-CoV-2 and RSV, and the unclear consequences of their mutual influence, we developed an in vitro lung epithelial cell model to study the molecular mechanisms and cellular pathways modulated in viral co-infection.
 Yang et al.
Yang et al.
Fecal gelatinase does not predict mortality in patients with alcohol-associated hepatitis
This study aimed to investigate the significance of fecal gelatinase on clinical outcomes in patients with alcohol-associated hepatitis. In conclusion, in our cohort, fecal gelatinase does not predict mortality and does not indicate higher disease severity in patients with alcohol-associated hepatitis.

Fecal gelatinase does not predict mortality in patients with alcohol-associated hepatitis
Yang et al.This study aimed to investigate the significance of fecal gelatinase on clinical outcomes in patients with alcohol-associated hepatitis. In conclusion, in our cohort, fecal gelatinase does not predict mortality and does not indicate higher disease severity in patients with alcohol-associated hepatitis.

Protein arginine methyltransferases in protozoan parasites: a new path for antiparasitic chemotherapy?
Campagnaro et al.This review discusses the activity and the relevance of arginine methyltransferases for the survival of pathogenic kinetoplastids, apicomplexans and amoebas, and how these enzymes could be exploited as drug targets.
Gut microbiota and ankylosing spondylitis: current insights and future challenges
Lobiuc et al.This review explores the growing role of gut microbiota in AS and its potential to reshape targeted treatment strategies and facilitate development of adjunct therapies to address disease onset and progression.
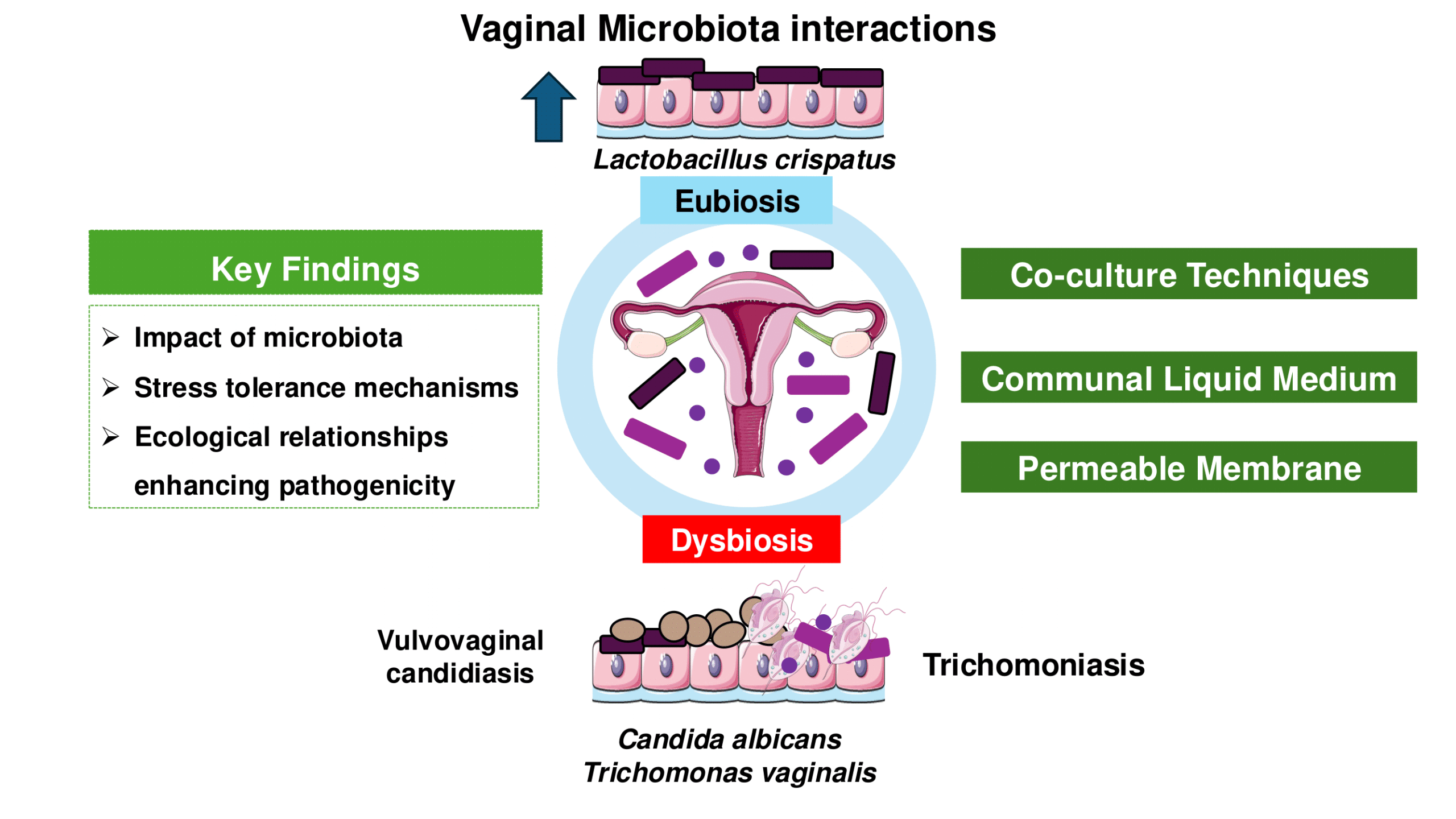
Advancements in vaginal microbiota, Trichomonas vaginalis, and vaginal cell interactions: Insights from co-culture assays
Cardoso and TascaThis review updates co-culture and co-incubation techniques for studying interactions of Lactobacillus spp., representing a pre-dominant member of the healthy vaginal microbiota; Candida spp., the most abundant yeast in the vagina, and T. vaginalis, responsible for the most widespread nonviral STI worldwide.
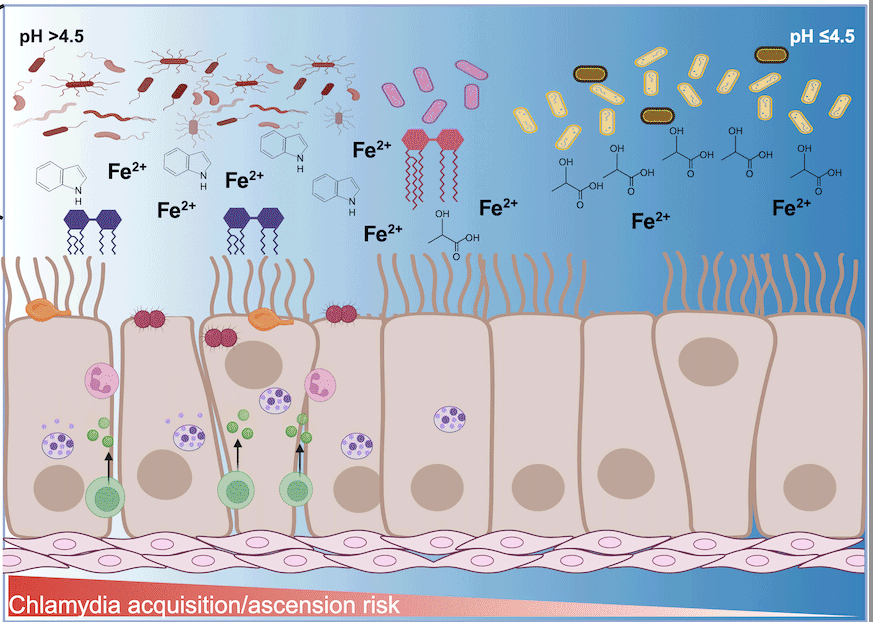
Influence of cervicovaginal microbiota on Chlamydia trachomatis infection dynamics
Hand et al.This review examines the complex interplay between the cervicovaginal microbiome, C. trachomatis infection, and host immune responses, highlighting the role of metabolites such as short-chain and long-chain fatty acids, indole, and iron in modulating pathogen survival and host defenses.
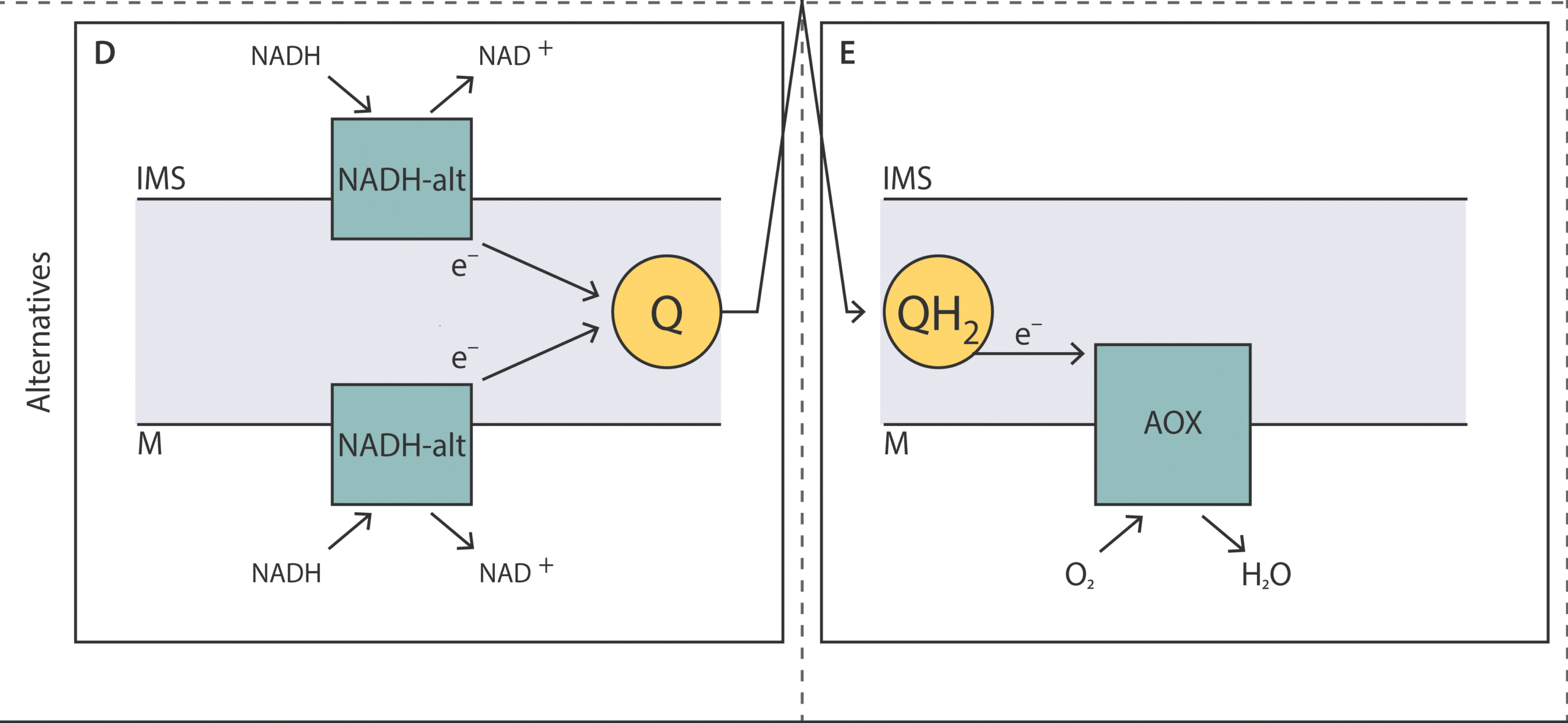
Unveiling the molecular architecture of the mitochondrial respiratory chain of Acanthamoeba castellanii
Scheckhuber et al.This review provides a comprehensive overview of the mitochondrial res-piratory chain in A. castellanii, focusing on the key alternative components involved in oxidative phosphorylation and their roles in energy metabolism, stress response, and adaptation to various conditions.

Paving the way for new antimicrobial peptides through molecular de-extinction
Osiro et al.The advancement of artificial intelligence and molecular de-extinction offers a valuable opportunity not only to discover new antimicrobials but also to provide accurate in silico predictions, thereby shortening the path to addressing the global antibiotic resistance crisis.

Efflux pumps: gatekeepers of antibiotic resistance in Staphylococcus aureus biofilms
Sinha et al.This review aims to elucidate the complex relationship between efflux pumps, antibiotic resistance and biofilm formation in S. aureus with the aim to aid in the development of potential therapeutic targets for combating S. aureus infections, especially those associated with biofilms.

Understanding the molecular mechanisms of human diseases: the benefits of fission yeasts
Acs-Szabo et al.Here we collect the latest laboratory protocols and bioinformatics tools for the fission yeasts to highlight the many possibilities available to the research community. In addition, we present several limiting factors that everyone should be aware of when working with yeast models.
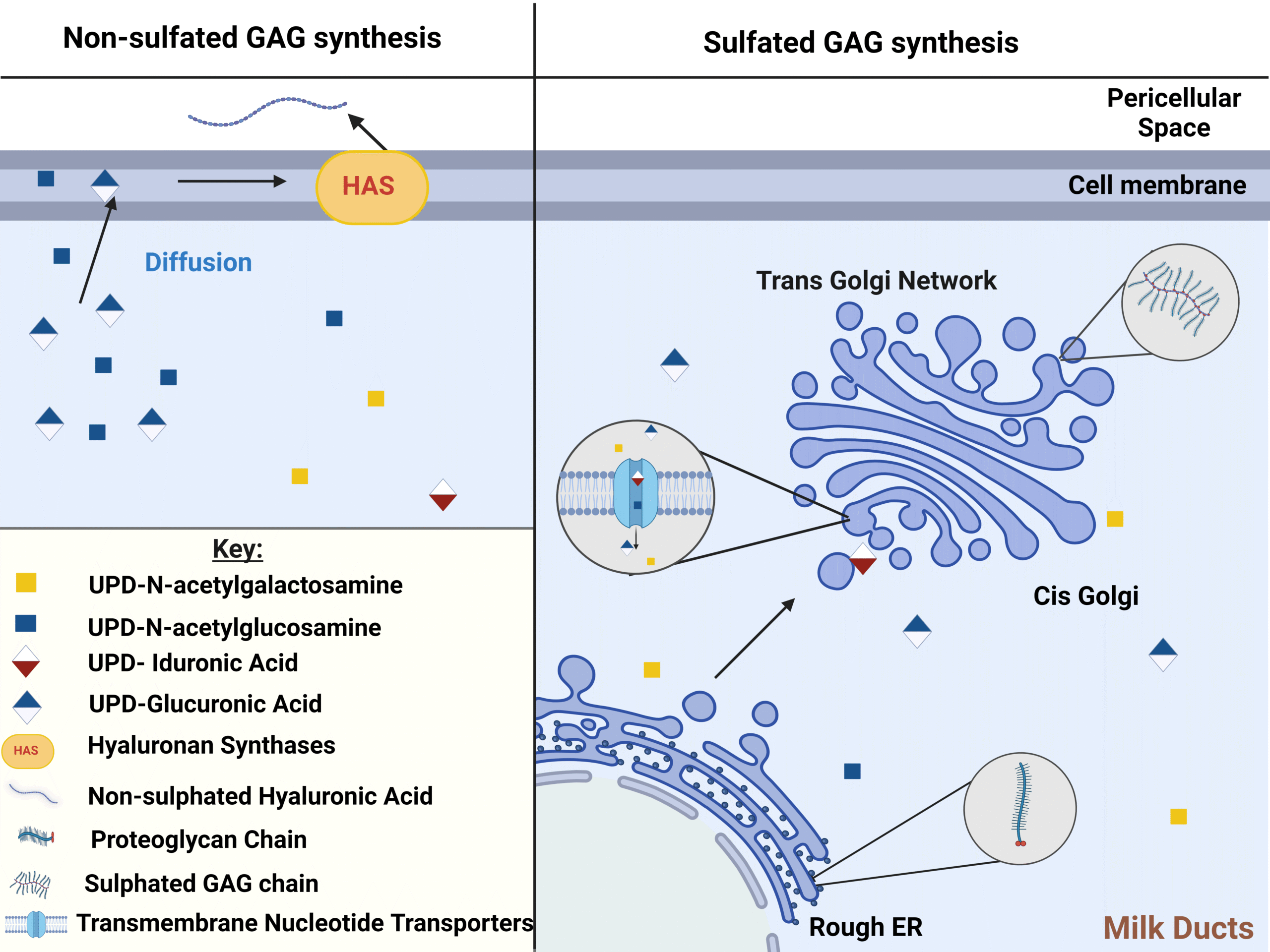
Characterising glycosaminoglycans in human breastmilk and their potential role in infant health
Greenwood et al.Glycosaminoglycans are bioactive components present in breast milk and play a potential key role in determining infant health yet are overlooked by many contemporary studies. This review explores their relevance, use and characterisation techniques.
Targeting GATA transcription factors – a novel strategy for anti-aging interventions?
Zimmermann et al.This article comments on work published by Carmona-Gutierrez et al. (Nat Commun., 2019), which identified a natural compound, 4,4′-dimethoxychalcone, inducing autophagy and prolonging lifespan in different organisms through a mechanism that involves GATA transcription factors.
In the beginning was the word: How terminology drives our understanding of endosymbiotic organelles
OborníkThis In the Pit article argues that the naming conventions for biological entities influence research perspectives and methodologies, advocating for mitochondria and plastids to be classified and named as bacteria due to their endosymbiotic origins, with potential implications for our understanding of bacterial prevalence, definitions of the microbiome and multicellularity, and the concept of endosymbiotic domestication.
What’s in a name? How organelles of endosymbiotic origin can be distinguished from endosymbionts
GruberThis In the Pit article suggests redefining the relationship between hosts and endosymbionts, like mitochondria and plastids, as a single species based on “sexual symbiont integration,” the loss of independent speciation, and congruence in genetic recombination and population sizes, rather than solely on historic classifications or structural properties.
Microbial wars: competition in ecological niches and within the microbiome
Bauer et al.In this Editorial Bauer et al. provide a brief overview on microbial competition and discuss some of its roles and consequences that directly affect humans.
Exploring the mechanism of amebic trogocytosis: the role of amebic lysosomes
Gilmartin and PetriIn this article, the authors comment on the study “Inhibition of Amebic Lysosomal Acidification Blocks Amebic Trogocytosis and Cell Killing” by Gilmartin et al. (MBio, 2017), discussing the the role of amebic lysosomes in Trogocytosis, the intracellular transfer of fragments of cell material.
Uncovering the hidden: complexity and strategies for diagnosing latent tuberculosis
Flores-ValdezThis editorial postulates that advanced proteomic and transcriptomic techniques are evolving and may enhance the detection of latent tuberculosis, thereby distinguishing true M. tuberculosis infections from other conditions, which is vital for controlling potential reactivation and transmission.
The Yin & Yang of Mitochondrial Architecture – Interplay of MICOS and F1Fo-ATP synthase in cristae formation
Rampelt and van der LaanThis Editorial posits that mitochondrial cristae architecture is shaped by the interplay of MICOS and ATP synthase, with a recent study illuminating their roles in cristae formation and maintenance.
When a ribosomal protein grows up – the ribosome assembly path of Rps3
Brigitte PertschyThis article comments on two papers by Mitterer et al., which followed yeast protein Rps3, highlighting the sophisticated mechanisms for protein protection, nuclear transport, and integration into pre-ribosomal particles for final assembly with 40S subunits.
Can’t find what you’re looking for?
You can browse all our issues and published articles here.
FAQs
Peer-reviewed, open-access research using unicellular organisms (and multicellular microorganisms) to understand cellular responses and human disease.
The journal (founded in 2014) is led by its Editors-in-Chief Frank Madeo, Didac Carmona-Gutierrez, and Guido Kroemer
Microbial Cell has been publishing original scientific literature since 2014, and from the very beginning has been managed by active scientists through an independent Publishing House (Shared science Publishers). The journal was conceived as a platform to acknowledge the importance of unicellular organisms, both as model systems as well as in the biological context of human health and disease.
Ever since, Microbial Cell has very positively developed and strongly grown into a respected journal in the unicellular research community and even beyond. This scientific impact is reflected in the yearly number of citations obtained by articles published in Microbial Cell, as recorded by the Web of Science (Clarivate, formerly Thomson/Reuters):
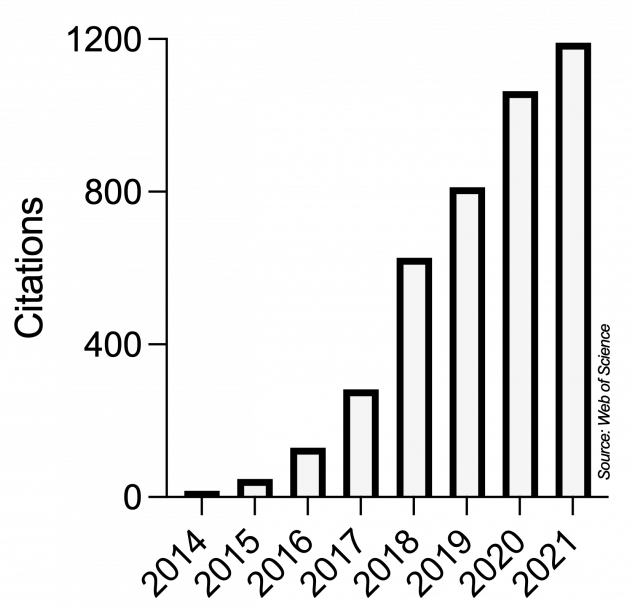
The scientific impact of Microbial Cell is also mirrored in a series of milestones:
2015: Microbial Cell is included in the Emerging Sources Citation Index (ESCI), a selection of developing journals drafted by Clarivate Analytics based on the candidate’s publishing standards, quality, editorial content, and citation data. Note: As an ESCI-selected journal, Microbial Cell is currently being evaluated in a rigorous and long process to determine an inclusion in the Science Citation Index Expanded (SCIE), which allows the official calculation of Clarivate Analytics’ impact factor.
2016: Microbial Cell is awarded the so-called DOAJ Seal by the selective Directory of Open Access Journals (DOAJ). The DOAJ Seal is an exclusive mark of certification for open access journals granted by DOAJ to journals that adhere to outstanding best practice and achieve an extra high and clear commitment to open access and high publishing standards.
2017: Microbial Cell is included in Pubmed Central (PMC), allowing the archiving of all the journal’s articles in PMC and PubMed.
2019: Microbial Cell is indexed in the prestigious abstract and citation database Scopus after a thorough selection process. This also means that Microbial Cell obtains, for the first time, an official Scopus CiteScore as well as an official journal ranking in the Scimago Journal and Country Ranking.
2022: Microbial Cell’s CiteScore reaches a value of 7.2 for the year 2021, positioning Microbial Cell among the top microbiology journals (previously available CiteScores: 2019: 5.4; 2020: 5.1).
2022: Microbial Cell is indexed in the highly selective Science Citation Index Expanded™, which covers approx. 9,500 of the world’s most impactful journals across 178 scientific disciplines. In their journal selection and curation process, Clarivate´s editors apply 24 ‘quality’ criteria and four ‘impact’ criteria to select the most influential journals in their respective fields. This selection is also a pre-requisite for inclusion in the JCR, which features the impact factor.
2022: Microbial Cell is listed in the Journal Citation Reports™ (JCR), and obtains its first official Journal Impact Factor™ (JIF) for the year 2021: 5.316.Check Article Types and Manuscript Preparation guidelines. Submit online via Scholastica.




Sulfur dioxide resistance in Saccharomyces cerevisiae: beyond SSU1
García-Ríos and GuillamónThis article discusses the importance of understanding sulfite resistance in Saccharomyces cerevisiae due to its use in winemaking and the potential role of the transcription factor Com2. While the SSU1 gene and its activity have been correlated with sulfite tolerance, the work by Lage et al. (2019) indicates that Com2 might control a large percentage of the genes activated by SO2 and contribute to the yeast’s protective response, offering new insights into the molecular factors influencing this oenological trait.