
Regulation of extracellular vesicles for protein secretion in Aspergillus nidulans
This study reveals that Aspergillus nidulans boosts extracellular vesicle production when ER-trafficked enzymes are induced, uncovering how fungi remodel their secretome through vesicle-mediated secretion to adapt to changing environments and biofilm formation.

Transcriptomic response to different heme sources in Trypanosoma cruzi epimastigotes
This study uncovers how the Chagas disease parasite adapts to changes in heme, an essential molecule for its survival, providing transcriptional clues to heme metabolism and identifying a previously unreported heme-binding protein in T. cruzi.

Luminal acetylation of microtubules is not essential for Plasmodium berghei and Toxoplasma gondii survival
Acetylation of α-tubulin at lysine 40 is not essential for cytoskeletal stability in Plasmodium berghei or Toxoplasma gondii, suggesting redundancy and plasticity in microtubule regulation in these parasites.

The dual-site agonist for human M2 muscarinic receptors Iper-8-naphtalimide induces mitochondrial dysfunction in Saccharomyces cerevisiae
S. cerevisiae is a model to study human GPCRs. N-8-Iper, active against glioblastoma via M2 receptor, causes mitochondrial damage in yeast by binding Ste2, highlighting evolutionary conservation of GPCRs.

Integrative Omics reveals changes in the cellular landscape of peroxisome-deficient pex3 yeast cells
To uncover the consequences of peroxisome deficiency, we compared Saccharomyces cerevisiae wild-type with pex3 cells, which lack peroxisomes, employing quantitative proteomics and transcriptomics technologies.

The mechanism of Tat-dependent protein translocation
Brüser and SandersThis review integrates mechanistically relevant biochemical, molecular, and structural studies on Tat-dependent translocation of folded proteins into an in its molecular detail new comprehensive explanation of how the Tat system mediates protein transport.

TOR-dependent regulation of the yeast homolog of the juvenile Batten Disease-associated gene CLN3
Pillalamarri et al.This study identifies conditions and genes that induce BTN1 expression in yeast. We show that BTN1 expression is regulated by translational control and by the mTOR1 pathway. An understanding of when and why BTN1 expression will aid in understanding the expression of CLN3, which may be helpful in the treatment of this devastating disease.

Overcoming phagocytosis resistance of hypervirulent Klebsiella pneumoniae by directly targeting capsules
Tsubaki et al.This study highlights a promising strategy for disarming hypervirulent K. pneumoniae by directly targeting its key virulence factors and provides novel insights into antibacterial therapeutic approaches against this clinically significant pathogen.
Protein arginine methyltransferases in protozoan parasites: a new path for antiparasitic chemotherapy?
Campagnaro et al.This review discusses the activity and the relevance of arginine methyltransferases for the survival of pathogenic kinetoplastids, apicomplexans and amoebas, and how these enzymes could be exploited as drug targets.

VapA/Scs2 sustains polarized growth in Aspergillus nidulans by maintaining AP-2-mediated apical endocytosis
Georgiou et al.To explore the functional significance of ER–PM contact sites in filamentous fungi, we identified and genetically characterized all Aspergillus nidulans proteins homologous to Snc2/VAP, Ist2, or tricalbins.

Genetic make-up and regulation of the L-lysine biosynthesis pathway in Vibrio natriegens
Straube et al.This study analysed the make-up and regulation of the biosynthetic pathway for L-lysine and related L-aspartate family amino acids (AFAAs) in Vibrio natriegens DSM759 to provide a comprehensive basis for future metabolic engineering endeavours aiming at developing this strain into an amino acid overproducer.
Measurement of apoptosis by SCAN©, a system for counting and analysis of fluorescently labelled nuclei
This work reports on a system for analyses of apoptosis-like programmed cell death in fungal hyphae that is composed of several modules, which enable automatic quantification of nuclei with chromatin condensation and DNA strand break in large datasets according to nuclei-associated fluorescent markers.
Rewiring yeast acetate metabolism through MPC1 loss of function leads to mitochondrial damage and decreases chronological lifespan
This work shows that MPC1-deficient cells make up for their impairment in mitochondrial pyruvate with a metabolic rewiring which involves several intermediates of the mitochondrially localized TCA cycle and the cytosolic glyoxylate shunt but ultimately results in a pro-aging process.
Overexpression of the transcription factor Yap1 modifies intracellular redox conditions and enhances recombinant protein secretion
This article investigates the role of Yap1 during the production of recombinant secretory proteins in glucose based growth conditions in Pichia pastoris, and reports a novel role of Yap1 during ER-resident oxidative protein folding.
Functional analysis of lipid metabolism genes in wine yeasts during alcoholic fermentation at low temperature
This study confirms the importance of specific genes in growth and fermentation activity of Saccharomyces cerevisiae at low temperature.
Angiotensin II type 1 receptor blockers increase tolerance of cells to copper and cisplatin
This study reports the identification of the drug class of Angiotensin II Type 1 receptor blockers (ARBs) and shows that specific ARBs increase yeast tolerance to Cu and Cp, and affect markers of Cu-induced apoptosis. Likewise, this study finds that specific ARBs increase human cell line tolerance to Cu and decrease the prevalence of apoptotic markers.
An extensive endoplasmic reticulum-localised glycoprotein family in trypanosomatids
This work describes a novel family of type I membrane proteins (“invariant glycoproteins”) and proposes them as trypanosomatid-specific ER-localised glycoproteins, with potential contributions to life cycle progression and immunity, that utilise oligomerisation as an ER retention mechanism.
Time resolved DNA occupancy dynamics during the respiratory oscillation uncover a global reset point in the yeast growth program
Using multiple approaches, this work implies a nucleosome focusing event as a key step that resets transcription during the respiratory oscillation.
Cell wall dynamics modulate acetic acid-induced apoptotic cell death of Saccharomyces cerevisiae
This work characterizes the involvement of MAPK signaling pathways in cell death induced by acetic acid in Saccharomyces cerevisiae.
Extracellular calcium triggers unique transcriptional programs and modulates staurosporine-induced cell death in Neurospora crassa
The results presented here reveal that in Neurospora crassa, extracellular Ca2+ modulates cell death and the transcriptional alterations induced by staurosporine, and lead to the identification of two novel putative Ca2+-binding proteins, encoded by the NCU08524 and NCU06607 genes.
Intersubunit communications within KaiC hexamers contribute the robust rhythmicity of the cyanobacterial circadian clock
Yohko Kitayama et al.This article comments on work published by Kitayama et al. (Nat Comm, 2013), which suggests that intersubunit communication precisely synchronizes KaiC subunits to avoid dephasing, and contributes to the robustness of circadian rhythms in cyanobacteria.
Mitochondrial protein import under kinase surveillance
Magdalena Opalińska and Chris MeisingerThis article summarizes recent discoveries in the yeast Saccharomyces cerevisiae model system that point towards a vital role of reversible phosphorylation in regulation of mitochondrial protein import.
Building a flagellum in biological outer space
Lewis D. B. Evans et al.This article comments on work published by Evans et al. (Nature, 2013), which presents a simple and elegant transit mechanism in which growth is powered by the subunits themselves as they link head-to-tail in a chain that is pulled through the length of the growing structure to the tip. This new mechanism answers an old question and may have resonance in other assembly processes.
A novel mechanism involved in the coupling of mitochondrial biogenesis to oxidative phosphorylation
Jelena Ostojić et al.This article comments on a study by Ostojić et al. (Cell Metabolism, 2013), which has uncovered a regulatory loop by which the biogenesis of a major enzyme of the OXPHOS pathway, the respiratory complex III, is coupled to the energy producing activity of the mitochondria.
Identifying the assembly pathway of cyanophage inside the marine bacterium using electron cryo-tomography
Wei Dai et al.Thiswork comments on a study by Dai et al. (Nature 2013) that illustrates that electron cryo-tomography is an approach whereby one can capture directly structural snapshots of transient phage assembly intermediates during maturation process. Such analysis can be generalizable not only to human viruses in human cells but also various molecular machines undergoing biological processes.
The emerging role of complex modifications of tRNALysUUU in signaling pathways
Patrick C. Thiaville and Valérie de Crécy-LagardThis comment discusses the article “Loss of wobble uridine modification in tRNA anticodons interferes with TOR pathway signaling” by Scheidt et al (Microbial Cell, 2014).
Only functional localization is faithful localization
Roland LillThis article comments on work published by Peleh et al. (Microbial Cell 2014), which analyzes the localization of Dre2 in Saccharomyces cerevisiae.
One cell, one love: a journal for microbial research
Didac Carmona-Gutierrez et al.In this inaugural article of Microbial Cell, we highlight the importance of microbial research in general and the journal’s intention to serve as a publishing forum that supports and enfolds the scientific diversity in this area as it provides a unique, high-quality and universally accessible source of information and inspiration.
What’s the role of autophagy in trypanosomes?
Katherine Figarella and Néstor L. UzcáteguiThis article comments on Proto et al. (Microbial Cell, 2014), who report first insights into the molecular mechanism of autophagy in African trypanosomes by generating reporter bloodstream form cell lines.
Can’t find what you’re looking for?
You can browse all our issues and published articles here.
FAQs
Peer-reviewed, open-access research using unicellular organisms (and multicellular microorganisms) to understand cellular responses and human disease.
The journal (founded in 2014) is led by its Editors-in-Chief Frank Madeo, Didac Carmona-Gutierrez, and Guido Kroemer
Microbial Cell has been publishing original scientific literature since 2014, and from the very beginning has been managed by active scientists through an independent Publishing House (Shared science Publishers). The journal was conceived as a platform to acknowledge the importance of unicellular organisms, both as model systems as well as in the biological context of human health and disease.
Ever since, Microbial Cell has very positively developed and strongly grown into a respected journal in the unicellular research community and even beyond. This scientific impact is reflected in the yearly number of citations obtained by articles published in Microbial Cell, as recorded by the Web of Science (Clarivate, formerly Thomson/Reuters):
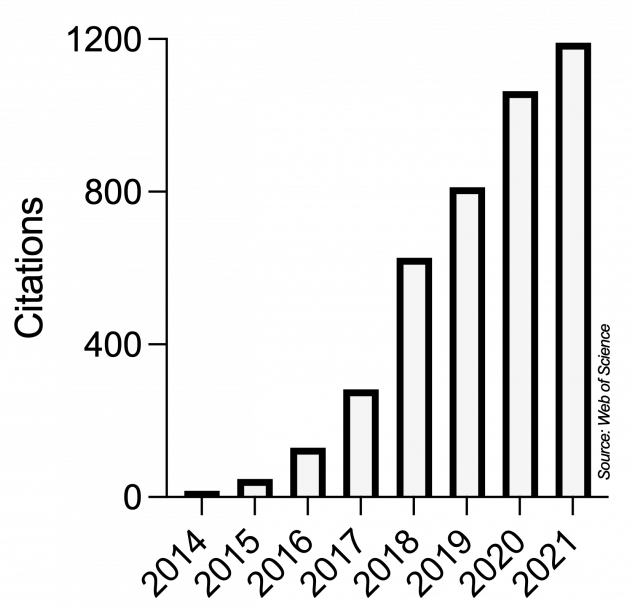
The scientific impact of Microbial Cell is also mirrored in a series of milestones:
2015: Microbial Cell is included in the Emerging Sources Citation Index (ESCI), a selection of developing journals drafted by Clarivate Analytics based on the candidate’s publishing standards, quality, editorial content, and citation data. Note: As an ESCI-selected journal, Microbial Cell is currently being evaluated in a rigorous and long process to determine an inclusion in the Science Citation Index Expanded (SCIE), which allows the official calculation of Clarivate Analytics’ impact factor.
2016: Microbial Cell is awarded the so-called DOAJ Seal by the selective Directory of Open Access Journals (DOAJ). The DOAJ Seal is an exclusive mark of certification for open access journals granted by DOAJ to journals that adhere to outstanding best practice and achieve an extra high and clear commitment to open access and high publishing standards.
2017: Microbial Cell is included in Pubmed Central (PMC), allowing the archiving of all the journal’s articles in PMC and PubMed.
2019: Microbial Cell is indexed in the prestigious abstract and citation database Scopus after a thorough selection process. This also means that Microbial Cell obtains, for the first time, an official Scopus CiteScore as well as an official journal ranking in the Scimago Journal and Country Ranking.
2022: Microbial Cell’s CiteScore reaches a value of 7.2 for the year 2021, positioning Microbial Cell among the top microbiology journals (previously available CiteScores: 2019: 5.4; 2020: 5.1).
2022: Microbial Cell is indexed in the highly selective Science Citation Index Expanded™, which covers approx. 9,500 of the world’s most impactful journals across 178 scientific disciplines. In their journal selection and curation process, Clarivate´s editors apply 24 ‘quality’ criteria and four ‘impact’ criteria to select the most influential journals in their respective fields. This selection is also a pre-requisite for inclusion in the JCR, which features the impact factor.
2022: Microbial Cell is listed in the Journal Citation Reports™ (JCR), and obtains its first official Journal Impact Factor™ (JIF) for the year 2021: 5.316.Check Article Types and Manuscript Preparation guidelines. Submit online via Scholastica.



Metabolic pathways further increase the complexity of cell size control in budding yeast
Jorrit M. EnserinkThis article comments on work published by Soma et al. (Microbial Cell, 2014), which teased apart the effect of metabolism and growth rate on setting of critical cell size in Saccharomyces cerevisiae.