Reviews:
Microbial Cell, Vol. 6, No. 8, pp. 335 - 355; doi: 10.15698/mic2019.08.686
Integrins in disguise – mechanosensors in Saccharomyces cerevisiae as functional integrin analogues
1 Dept. of Chemistry and Molecular Biology, Univ. of Gothenburg, Sweden.
Keywords: integrins, yeast, mechanosensors, stress-responses, starvation.
Abbreviations:
cAMP – cyclic AMP,
CWI – cell wall integrity,
ECM – extracellular matrix,
FAC – focal adhesion complex,
FAK – focal adhesion kinase,
MIDAS – metal ion-dependent adhesion site,
VWA – von Willebrand A.
Received originally: 23/12/2018 Received in revised form: 03/07/2019
Accepted: 08/07/2019
Published: 15/07/2019
Correspondence:
A. Blomberg, phone: +46 31 786 2589; anders.blomberg@cmb.gu.se
Conflict of interest statement: The authors declare no conflict of interests.
Please cite this article as: Tarek Elhasi and Anders Blomberg (2019). Integrins in disguise - mechanosensors in Saccharomyces cerevisiae as func-tional integrin analogues. Microbial Cell 6(8): 335-355. doi: 10.15698/mic2019.08.686
Abstract
The ability to sense external mechanical stimuli is vital for all organisms. Integrins are transmembrane receptors that mediate bidirectional signalling between the extracellular matrix (ECM) and the cytoskeleton in animals. Thus, integrins can sense changes in ECM mechanics and can translate these into internal biochemical responses through different signalling pathways. In the model yeast species Saccharomyces cerevisiae there are no proteins with sequence similarity to mammalian integrins. However, we here emphasise that the WSC-type (Wsc1, Wsc2, and Wsc3) and the MID-type (Mid2 and Mtl1) mechanosensors in yeast act as partial functional integrin analogues. Various environmental cues recognised by these mechanosensors are transmitted by a conserved signal transduction cascade commonly referred to as the PKC1-SLT1 cell wall integrity (CWI) pathway. We exemplify the WSC- and MID-type mechanosensors functional analogy to integrins with a number of studies where they resemble the integrins in terms of both mechanistic and molecular features as well as in the overall phenotypic consequences of their activity. In addition, many important components in integrin-dependent signalling in humans are conserved in yeast; for example, Sla1 and Sla2 are homologous to different parts of human talin, and we propose that they together might be functionally similar to talin. We also propose that the yeast cell wall is a prominent cellular feature involved in sensing a number of external factors and subsequently activating different signalling pathways. In a hypothetical model, we propose that nutrient limitations modulate cell wall elasticity, which is sensed by the mechanosensors and results in filamentous growth. We believe that mechanosensing is a somewhat neglected aspect of yeast biology, and we argue that the physiological and molecular consequences of signal transduction initiated at the cell wall deserve more attention.
INTRODUCTION
The yeast Saccharomyces cerevisiae is an excellent test bed for the study of human proteins, and in a recent systematic, large-scale study it was found that roughly 50% of human proteins with homology to yeast proteins could compensate for the corresponding gene deletion in yeast [1]. This systematic study of human proteins in yeast revealed a high degree of conserved functions; in particular for genes involved in specific functional classes like central metabolism and lipid and sterol biosynthesis. Human proteins with no clear homology to yeast proteins can also be studied in this model system, and several important disease models of human genes have been developed and utilised for functional studies in yeast (the following reviews provide excellent overviews and a more complete list of the use of yeast in surrogate genetics related to human disease [2][3][4]). Thus, S. cerevisiae has proven its worth as a powerful tool in the discovery and study of mechanisms involved in a wide array of human diseases.
–
The strategy in yeast surrogate genetics is to learn about human proteins by studying their functions in yeast. However, in this review we will explore the opposite logic – to start with knowledge about a human protein and then seek out functionally equivalent analogues in yeast. We focus on integrins and integrin-signalling in humans and review the importance and functions of mechanosensor analogues in yeast. Integrin-mediated adhesion and signalling are probably the most important cell adhesion mechanisms for metazoan development. However, many of the components of the integrin adhesome evolved well before the origin of metazoans and fungi [5], and it has been suggested that several key components of the integrin adhesome have been lost independently in fungi and choanoflagellates. In this context, it is interesting to note that genome sequences from unicellular metazoan-related lineages have pushed the time of origin of gene families earlier believed to be metazoan-specific back into the Proterozoic period, e.g. tyrosine kinases [6], transcription factors [7], and cadherins [8].
–
We will start with a short overview of the function of integrins in humans and the evolution of integrin and integrin-like systems. We will then make connections to the mechanosensors in yeast and the various aspects of their molecular functions and biological roles, highlighting where we think they are similar to integrins. Finally, we will outline several hypotheses in relation to the cell wall in yeast and the role of the mechanosensors in transmitting cell wall changes to intracellular signalling. We think yeast mechanosensors deserve more attention and that future studies will most likely reveal novel biological activities, and it is possible that the knowledge generated in yeast will feed back to the human system to provide a better understanding of the complexity of mechanosensing and the function of integrins in humans.
BACKGROUND TO INTEGRINS
Integrins in mammals
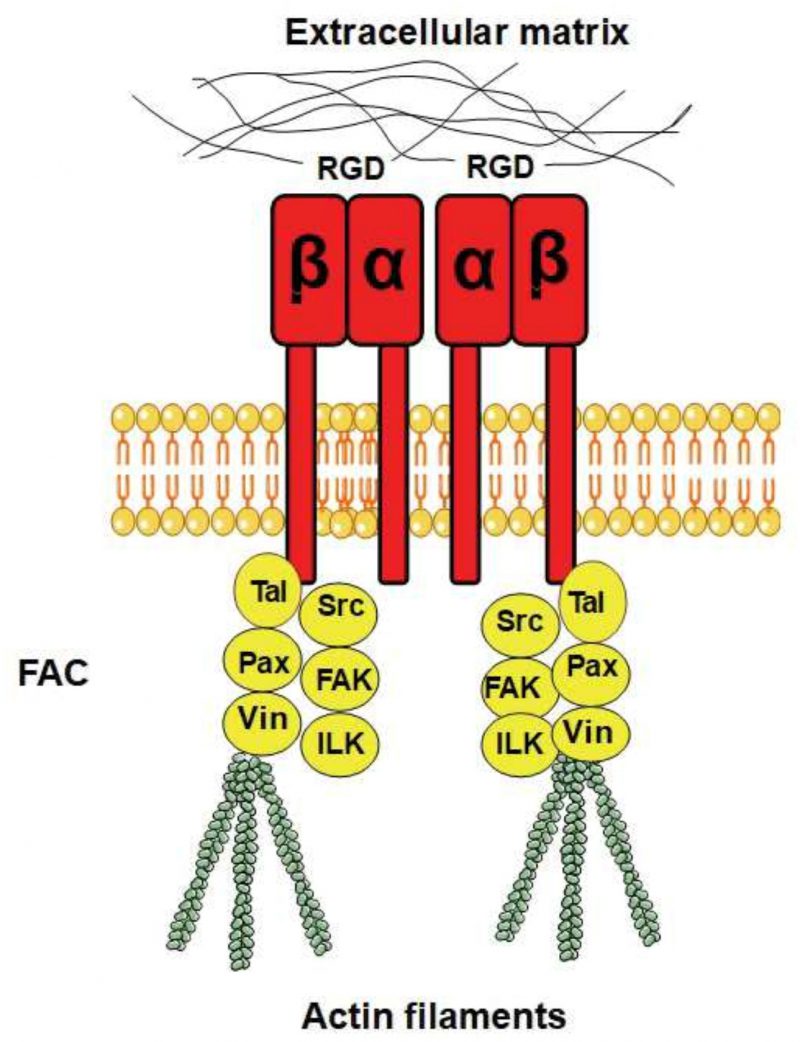
The ability to sense mechanical stimuli is vital for all organisms. In mammals, mechanical stimuli are mainly sensed by transmembrane adhesion receptors called integrins that mediate cell adhesion to the extracellular matrix (ECM) or to neighbouring cells [9]. The extracellular domain of integrin interacts with extracellular protein ligands that bind specifically to integrin, while the intracellular domain acts as an attachment site for the internal cytoskeleton through actin-associated proteins, forming the focal adhesion complex (FAC) [10][11]. In this way integrins physically link the ECM to the cytoskeleton in the interior of the cell [11]. In fact, the name “integrin” was given to these cell receptors because of their importance in the integration of the ECM and the cytoskeleton [12][13]. Integrins are highly important proteins involved in a wide variety of processes such as the development of the immune system [14], apoptosis [15], and wound healing [16]. They are also functionally involved in many diseases such as cancer [17], diabetes [18], and Alzheimer's [19] as well as in aging [20].
–
Integrins are heterodimers composed of α and β subunits, which are non-covalently bound to each other [12]. In humans, there are 18 α subunits and eight β subunits that physically interact and form 24 different heterodimers [21]. Integrins are localised to the plasma membrane and are composed of a rather extensive extracellular domain that is about 20 nm long [22], one transmembrane domain with a single membrane-spanning helix, and a shorter intracellular domain (Fig. 1). Integrin β subunits contain a number of rather well-studied domains, including the von Willebrand A (VWA) domain, an N-terminal PSI domain, and repeated EGF-like domains [23]. Many VWA domains bind metal ions via a sequence motif called the metal ion-dependent adhesion site (MIDAS) with the consensus sequence DXSX, and a number of human diseases arise from mutations in the VWA domain [24]. The majority of VWA domains are found in cell adhesion and ECM proteins [25], as well as in certain classes of intracellular proteins [24]. A plausible hypothesis for the function of the VWA domain is that it supports protein–protein interactions through the joint coordination of a metal ion with the MIDAS motif [26]. Although integrins are heterodimers, it has been suggested that β integrin is the principle subunit for the intracellular binding to the cytoskeleton and the extracellular binding to signalling molecules, while α integrin has a regulatory role in maintaining the proper conformation of β integrin for binding to its ligands [27].
–
–
Integrin ligands
The different combinations of α and β subunits create different kinds of integrins with different ligand specificities, with some integrins binding to ECM proteins and others binding to cell surface receptors. Some integrins recognise the RGD motif on their ligands as an active site for binding (Fig. 1), while other integrins recognise the LDV motif. The ECM ligands with RGD as the active integrin binding site include vitronectin, fibronectin, and fibrinogen, where vitronectin and fibronectin also contain an LDV domain. Integrins also bind to proteins with alternative binding motifs; for example, collagen has the GFOGER motif (where O is hydroxyproline) as its recognition site while the recognition site of laminin is not yet known [27]. Synthetic peptides that contain an RGD motif can inhibit integrin signalling by outcompeting RGD-containing ligands, and these peptides are widely used as integrin inhibitors and have been used to study integrin signalling in different organisms (see below).
–
ECM and integrin signalling in mammals
In mammals, the nature of the ECM is of great importance, and integrins mediate the response to ECM stiffness that determines cell fate [28]. Cells remodel the ECM in response to the force applied to integrin-mediated adhesions. For example, when fibroblasts are stretched they activate the expression of genes coding for collagen, fibronectin, and metalloproteins and form a dense ECM that is rich in collagen [29]. This regulation occurs in many cases through signalling pathways downstream of integrins [30]. Integrin expression also changes according to ECM elasticity [31], and this elasticity probably causes conformational changes in integrins – and thus changes their ligand-binding properties – or exposes buried domains to their interacting proteins [32].
–
Integrins and actin foci in mammals
FACs are specialised sites where clustered integrin receptors interact with the ECM on the outside of cells and with the actin cytoskeleton on the inside of cells (Fig. 1). The FACs provide adhesion to the ECM and transmit mechanical tension generated within cells across the plasma membrane to the external environment. In addition, they act as scaffolds for many signalling pathways that are triggered by ligands binding to the integrins or by mechanical forces exerted on the cells [32][33]. FACs are distributed in the cell in a punctate pattern [34], and the most important components of FACs are talin (an adapter protein), focal adhesion kinase (FAK, a non-receptor tyrosine kinase), paxilin (an adapter protein), Src-family kinase (another tyrosine kinase), integrin-linked kinase, and vinculin.
–
Talin was found to be instrumental for the activation of integrins and to enhance the association between integrins and the ECM [35]. It has been shown that the stretching of talin exposes multiple cryptic vinculin-binding sites that are normally (without the tension) buried within talin [36]. Given the prominence of talin and vinculin in FACs, this tension-induced interaction is likely a key factor for FAC assembly, growth, and function.
–
Integrin signalling in mammals
When bound to an extracellular ligand, integrins bind intracellularly to many other proteins to form the FAC. When the FAC is formed, it activates diverse signalling components like the tyrosine kinase c-Jun N-terminal kinase, phosphatidylinositol 3-kinase (AKT), the GTPases of the RHO family [37], and the mitogen activated protein kinases (MAPKs) ERK and p38 [38].
–
The interactions between integrins and the ECM are important in order for the cell to adjust its intracellular activities in response to the surrounding microenvironment, and these interactions occur through two different mechanisms [39]. First, there are signals from the ECM to the cell (outside-in signalling). Here integrins sense a change in the ECM tension (if ECM is loose or stiff) and sense the dimensional changes in the ECM (increases in depth/volume). In addition to the response to the ECM dynamics, integrins also receive signals from antagonists like growth factor receptors, and they interact with other membrane proteins like tetraspanins that function as adaptors in recruiting other integrin interactors [40]. In the second type of interaction (inside-out signalling), cellular signals bind to the integrin cytopasmic domain and cause a conformational change that result in integrin activation [41].
EVOLUTION OF INTEGRINS
The integrin α and β subunits exist throughout the metazoans. It is believed that integrins have an early origin, preceding the first metazoans, with some integrin domains identifiable even in bacteria [42][43]. For example, proteins containing VWA domains are present throughout the eukaryotes (metazoa, fungi, plants, and protists) as well as in eubacteria and archaea. However, both integrin α and β, as well as several other components of the integrin adhesion complex, are absent from choanoflagellates and fungi and were presumably lost independently in these two lineages. However, integrin-like functions exerted by integrin functional analogues seem conserved even if integrin sequence homologues might be missing.
–
Integrin-like proteins in plants
The cell wall is considered the ECM in plants [44], and similar to the animal ECM the plant cell wall contains mechanosensors that sense mechanical changes imposed on the cell wall from the surrounding environment. There are no integrin sequence homologues in plants, but experimental evidence – mainly based on immunoblotting techniques – suggests the existence of what are called “integrin-like proteins” and “integrin-like signalling” in plants. For example, monoclonal antibodies against chicken β1 integrin, human vitronectin, and human fibronectin recognised proteins on the surface of protoplasts isolated from onion epidermis and tobacco root cap [45][46]. In addition, polyclonal antibodies against chicken β1 integrin detected proteins in the Arabidopsis thaliana plasma membrane [47].
–
Integrin inhibitor peptides containing RGD sequences have been exploited in plant studies to study integrin-like mechanisms. Arabidopsis protoplasts agglutinate when expressing ProNectin, a genetically engineered protein that contains 13 RGD sequences, and RGD peptides disrupt this adhesion [48]. In another study, application of RGD peptides to peas disrupted the cell wall-plasma membrane connection [49]. Overall, applying RGD-containing peptides to plant cells interferes with many physiological processes such as development [50], cytoplasmic streaming [51], gravisensing [52], root induction [53], and defence responses [54]. The RGD peptides seem to act on proteins at the plasma membrane-cell wall interface [54]. It has been proposed that these integrin-like proteins likely act as mechanosensors as part of the defence response, and that their attachment to receptors in the cell wall or changes in the mechanical property of the cell wall might warn the host plant about pathogen invasion [55]. In A. thaliana, the predicted structure of NDR1, a membrane protein involved in the defence response, showed that the core of NDR1 has strong 3-dimensional structural similarity with β integrin [56]. In addition, the A. thaliana genome codes for five proteins in the integrin-linked kinase family. Another important protein in FACs, FAK, has also been detected in plants [57]. Moreover, proteins with sequence similarity to filamin, α-actinin, integrin-linked kinase, and proteins with a LIM domain (which is important for protein association to the FAC) have been found in green algae [58]. In summary, it is likely that integrin-like proteins and integrin-like signalling exist throughout the plant kingdom.
–
Integrin-like proteins in unicellular organisms
In immunoblotting experiments, integrin-like proteins have also been detected in oomycetes [59] and in the amoebae Naegleria fowleri and Naegleria lovaniensis [60]. Furthermore, the integrin-like protein β1EhFNR was detected in Entamoeba hystolytica based on immunoblotting [61]. Despite the lack of sequence similarity between β1EhFNR and mammalian integrin, β1EhFNR shows functional similarity to integrins because it forms intracellular FACs. These complexes contain homologues to many mammalian FAC proteins such as actin, myosin I, myosin II, alpha actinin, and FAK [62] as well as paxilin and vinculin [63]. The E. hystolytica β1EhFNR does not form a heterodimer like the mammalian integrins do, suggesting that integrin-like signalling can be mediated solely by the equivalent to the integrin β subunit.
–
Integrin-like proteins in fungi and yeast
It was shown by applying RGD-containing peptides that integrin-like proteins in fungi are important for fungal germination during the pre-penetration stage [64]. Based on immunoblotting assays, integrin-like proteins were detected in the yeast form of Histoplasma capsulatum [65], and RGD peptides inhibited conidial adhesion and appressorium formation in Magnaporthe oryzae [66]. The conditional yeast pathogen Candida albicans binds to hyphae of human lymphocyte cells. This binding was shown to be inhibited by a variety of RGD-containing peptides; thus, indicating that some integrin-like proteins on Candida are involved in this yeast-human cell binding [67]. Furthermore, the adhesion of the C. albicans germ tube to purified human vitronectin was inhibited by monoclonal antibodies against integrins α5 and β3, thus showing that Candida contains some integrin-like proteins [68].
–
The protein INT1/BUD4 in C. albicans has integrin-like motifs that are involved in integrin-ligand binding, and this protein localises to the cell surface [69]. It was also reported that the ascomycete fungus Pneumocystis carinii (a pneumonia-causing pathogen) expresses a protein with considerable structural similarity to INT1 [70] (Fig. 2). The 1,006 amino acid Pneumocystis protein (pcINT1) has homology to the C-terminal region of C. albicans INT1 (31% sequence identity) and contains four MIDAS motifs (DXSX) required for adhesion and for coordination of divalent cations, a membrane spanning region, an internal RGD motif, and a specific tyrosine residue found in the cytoplasmic tails of some human integrin receptors and in INT1 from C. albicans. The internal RGD motif in INT1 might well just be a coincidence, because RGD is not found to be functional in human integrins. Furthermore, heterologous expression of pcINT1 in S. cerevisiae showed that pcINT1 was localised to the surface and that yeast cells subsequently gained the ability to bind to human fibronectin in a cation-dependent fashion. However, despite the presence of several integrin-like motifs none of the INT1 proteins are good sequence homologs to human integrins; the caINT1 and the human integrins have only roughly 20% overall sequence similarity (global alignment).
–
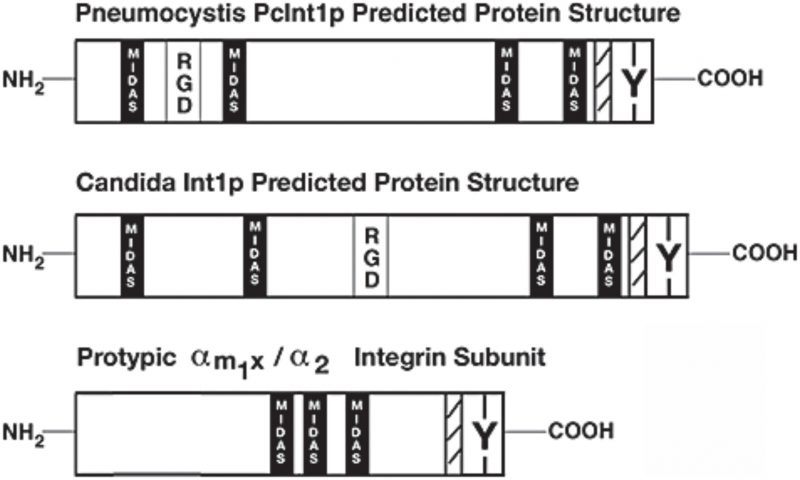 | FIGURE 2: Integrin-like proteins in fungi share some of the overall structure and motifs with mammalian integrins. Schematic representation of integrin-like INT1 from Candida albicans and Pneumocystis carinii. Indicated are the approximate locations of some motifs – RGD (the integrin binding site), MIDAS (the metal ion-dependent adhesion site), Y (the conserved tyrosine-residue in the C-terminal part), and hatched marking (the transmembrane domain). A prototypical human integrin subunit is shown at the bottom. This figure is from [70] and is published with permission from John Wiley and Sons. Copyright © 2008, American Society for Microbiology. |
MECHANOSENSORS IN SACCHAROMYCES CEREVISIAE – INTEGRINS IN DISGUISE
No integrin homologues in S. cerevisiae
There are no sequence homologues to human integrins in the yeast S. cerevisiae. Blast searches with any of the 26 human integrin α or integrin β proteins do not yield any hits (E-value threshold 10−5) in the S. cerevisiae genome. However, the integrin-like INT1 proteins from Candida or Pneumocystis both have rather strong matches to the S. cerevisiae sequence homologue Bud4 (E-value of 10−47 and 10−20, respectively), but with homology only to a very distinct C-terminal region. Bud4 is an anillin-like protein that is involved in bud-site selection and the formation and disassembly of the double septin ring structure in S. cerevisiae. However, it has no membrane-spanning domain and consequently no extracellular region, and therefore does not have an overall integrin-like structure like INT1. Bud4 in S. cerevisiae is thus not believed to be functionally analogous to INT1 or to animal integrins.
–
Mechanosensors in S. cerevisiae as functional integrin analogues
There is a group of proteins that act as mechanosensors in yeast and that activate different intracellular signalling pathways upon external perturbations of either the cell wall or the plasma membrane in much the same way as mammalian integrins. Various environmental cues recognised by these mechanosensors are transmitted by a conserved signal transduction cascade commonly referred to as the PKC1-SLT1 CWI pathway [71]. Similar to integrins, these mechanosensors in yeast consist of a long extracellular domain that is anchored to the ECM (i.e. the cell wall in the case of yeast), a single transmembrane domain that anchors these proteins in the plasma membrane, and a short C-terminal cytoplasmic tail that interacts with and activates intracellular signalling components (Fig. 3) [72]. Another similarity is that both human integrins and the yeast Wsc1 exhibit bidirectional signalling with the actin cytoskeleton. In mammals, actin cytoskeleton regulates integrin clustering, while actin dynamics is regulated by integrin-mediated adhesion [73]. Similarly, in yeast Wsc1 localization is mediated through actin cytoskeleton, while actin depolarization caused by cell wall damage is mediated through Wsc1 [74].
–
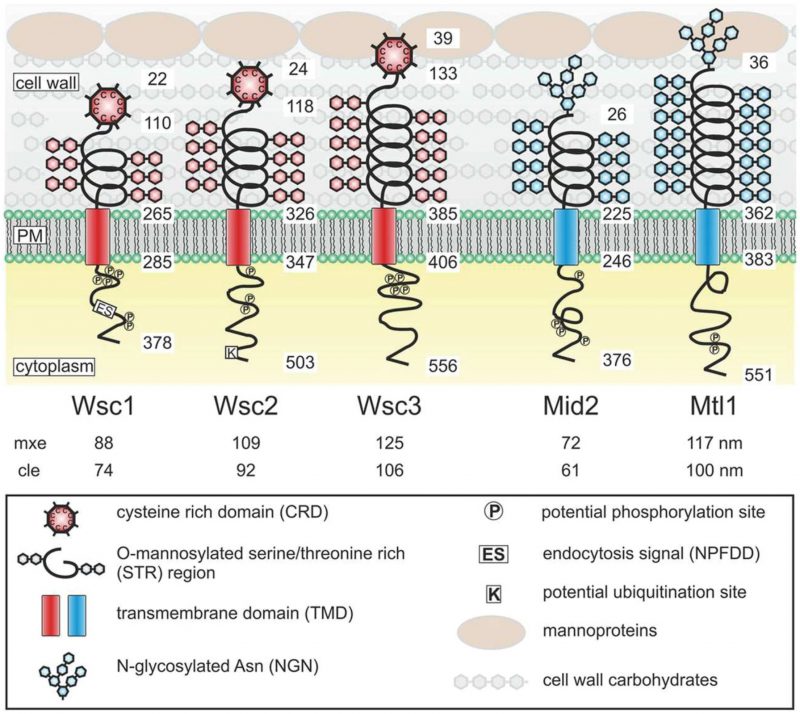 | FIGURE 3: Structures and proposed functional domains of yeast mechanosensors. Protein domains are indicated in the legend box. PM, plasma membrane. Numbers indicate residues in the respective proteins counting from the N-termini. Numbers below the protein names indicate the estimated length of the extracellular sensor region. mxe, maximal possible extension. cle, computed live-cell extension. This figure is from [72] and is published with permission from the American Society for Microbiology. Copyright © 2015, American Society for Microbiology. |
–
These cell wall mechanosensors in S. cerevisiae can be divided into two types – the WSC type that includes the genes WSC1/SLG1 (SLG1 is its standard gene name, but we will here use WSC1 for consistency with the other WSC genes), WSC2, and WSC3 and the MID type that includes the genes MID2 and MTL1. The functional similarity between Wsc1 and integrins has been described in several studies [74][75][76]. The two mechanosensor types in yeast differ in the nature of their extracellular head groups, where the head group is a cysteine-rich domain in the WSC-type sensors but is an N-glycosylated asparagine in the MID-type sensors (Fig. 3). The head group of the WSC-type sensors is presumed to be in contact with the glucan layer, while the head group of the MID-type sensors is presumed to be in contact with both the glucan and the mannoprotein layers in the cell wall [72]. The presumed physical interaction with the mannoprotein layer might give the MID-type mechanosensors the capacity to sense the thickness and elastic properties of the outer mannoprotein layer, as proposed by Heinisch and colleagues [72]. Because both sensor types can bind to the mannoprotein layer of the cell wall, they are believed to be redundant in sensing the thickness and elastic properties of the glucan layer.
–
Both types of yeast mechanosensors possess a cytoplasmic tail that communicates with downstream signalling components and thus allows cells to respond rapidly to external mechanical stress. The proposed model is that the mechanosensors' intracellular C-terminal tails interact with the GDP/GTP exchange factor Rom2 that subsequently activates the small GTPase Rho1 [77]. RHO1 encodes an essential small GTPase in the Rho/Rac subfamily of Ras-like GTPases, and in S. cerevisiae Rho GTPases are also encoded by RHO2, RHO3, RHO4, RHO5, and CDC42, and of these at least Rho5 has also been shown to be activated by Wsc1 [78]. Rho1 regulates protein kinase C (encoded by PKC1), which is a protein serine/threonine kinase that is essential for cell wall remodelling during growth. Pkc1 is a homolog of the alpha, beta, and gamma isoforms of mammalian protein kinase C (PKC). Pkc1 then activates a basic three-protein kinase signalling module that integrates the MAPKKK Bck1, the redundant MAPKKs Mkk1 and Mkk2, and the MAPK Slt2.
–
The cellular roles of the mechanosensors in yeast have many similarities with integrins. Integrins in humans are involved in the establishment of cell polarity [79], the response to osmotic stress [80], actin cytoskeleton organisation [81], the response to heat stress [82], regulation of autophagy [83], aging [20], and the positive regulation of endocytosis [84]. In a similar manner, Wsc1 in S. cerevisiae is also involved in the establishment of cell polarity [74][85], the response to osmotic stress [86], actin cytoskeleton organisation [74], the positive regulation of endocytosis [85][87], the response to heat [74][88], and the regulation of autophagy [89]. The mechanosensors also have an impact on chronological life span in yeast [90][91]. In addition, both human integrins and yeast mechanosensors are members of the RHO-dependent signal transduction pathway that communicates downstream with the MAPK signalling pathway [37][74][77]. Wsc1 also has a dynamic localisation that resembles the polarised distribution of integrins in mammalian cells [76], where the polarised distribution for some of the mechanosensors is mediated through their endocytosis and recycling [92]. Moreover, human integrins protect cells from the genotoxic effects of bleomycin (a drug that degrades DNA) [93], and wsc1 mutant yeast are hypersensitive to bleomycin [94]. Wsc1 acts upstream of Ste20 [95], which is required for pheromone-induced apoptosis [96], and upstream of Rho5, which plays a role in oxidative stress-induced apoptosis [97], and both of these observations suggest that Wsc1 might play a role in apoptosis in a similar manner as human integrins [98]. Taken together, S. cerevisiae possesses mechanosensors connected to the cell wall that in many ways are functional analogues to human integrins.
THE YEAST CELL WALL IS A HIGHLY DYNAMIC STRUCTURE
The cell wall in yeast resembles the animal ECM, and similar to the ECM changes in the cell wall in yeast trigger different biological responses. The yeast cell wall determines cell shape, provides rigidity that counteracts the outward turgor pressure on the plasma membrane, protects the cells against external environmental injury, and acts as a diffusion barrier delimiting the periplasmic space. The yeast cell wall is a prominent structure that makes up 15–30% of the cellular dry weight [99][100], has a thickness of 110–200 nm [101][102][103], and is mainly built from glucans (β-1,3-glucan and β-1,6-glucan), proteins (many of them being mannoproteins containing mannose), and chitin (N-acetylglucosamine polymers) [101]. An internal skeletal framework in the yeast cell wall forms a three-dimensional network of β-1,3-glucan molecules that surrounds the whole cell. This layer is primarily responsible for the mechanical strength of the cell wall. The β-1,3-glucan molecules are branched and they therefore have multiple nonreducing ends that can act as attachment sites for other components of the cell wall. Some of the mechanosensors impact on cell wall synthesis, i.e. deletion of WSC1 results in a drastic decrease in the synthesis of β-1,3-glucan at the bud [104]. The skeletal glucan framework is strengthened close to the plasma membrane by chitin chains [105]. Chitin represents a minor component of the cell wall (only 1–3% of its biomass), but chitin is important for the proper function of the cell wall and is essential for fungal viability [106]. It has been shown that either overexpression of the mechanosensor Mid2 [88] or mutations that result in a defective cell wall [107][108] cause hyper-accumulation of chitin. Chitin polymers are covalently cross-linked to the β-glucan network and are thought to contribute to the rigidity and physical strength of the cell wall [109]. On the outside of the β-1,3-glucan skeletal framework are mainly found β-1,6-glucan molecules that interconnect cell wall proteins with the central β-1,3-glucan framework [100].
–
The yeast cell wall is a highly dynamic structure, and its elasticity depends on a number of external and internal factors like nutrient source, temperature, and certain mutations; for example, cell wall elasticity can be increased by mutations that inactivate the CRH family of cell wall cross-linking chitin transglycosylases or can be decreased by overexpression of CRH family members [110]. It has also been observed that a sudden decrease in cell volume as a result of osmotic shock drives rapid increases in cell wall thickness. However, these changes in cell wall thickness are reversible, and quite rapidly the cell diameter and cell wall thickness return to normal when the osmotic stress is relieved. This behaviour seems analogous to springs being oriented parallel to the plasma membrane that are compressed and relaxed (Fig. 4). Several molecular factors contribute to the elasticity of the C. albicans cell wall, and it was suggested that the helical conformation of β-glucans might permit their expansion and contraction [111]. In addition, slippage of locally aligned polymers over one another could promote the rapid changes that are observed in the dimensions of the glucan and chitin layers in the cell wall. An important factor for this elasticity is that the helical compression and polymer slippage are constrained by covalent crosslinks between cell wall polymers (chitin and glucans) as well as between these polymers and the cell wall proteins.
–
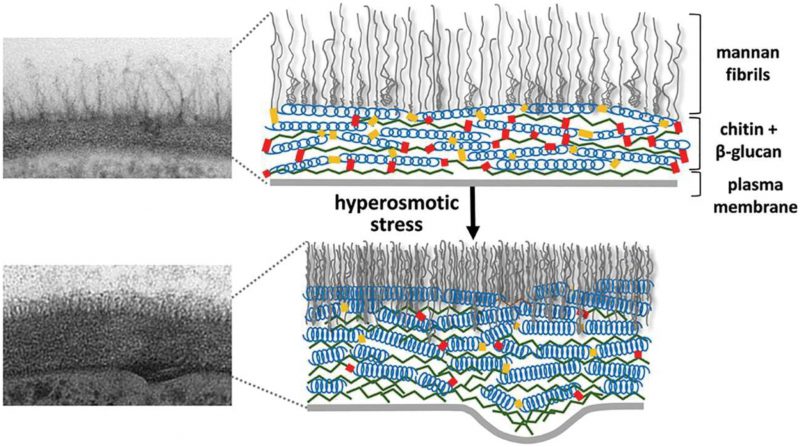 | FIGURE 4: Cell wall structure before and after hyper-osmotic stress. The cell wall is shown as a transmission electron microscopy image (left) and as a schematic drawing (right). After osmotic dehydration (hyper-osmotic shock) the cell wall becomes much more compact and thicker. In the schematic drawing, this is indicated as the β-glucan helical structures acting like springs being compressed. This figure is from [150] and is published with permission from the American Society for Microbiology. Copyright © 2015, American Society for Microbiology. |
–
Inhibition of enzymes involved in the biosynthesis of cell wall components also leads to altered cell wall properties. For example, calcofluor white is a fluorescent agent that strongly binds to chitin [112] and interferes with chitin synthesis [112][113]. The ATP analogue caffeine is also considered a drug that imposes cell wall stress [114]. Caffeine has been shown to inhibit the Tor1 kinase [115][116] which results in activation of the CWI pathway leading to changes in the cell wall (see below). Caspofungin is an antifungal from the echinocandin family that inhibits β-1,3-glucan synthesis [117] and thus causes an increase in the β-1,6-glucan polysaccharide fraction and a partial reduction of β-1,3-glucan, both in S. cerevisiae and C. albicans [118]. All of these agents are frequently used in studies of the impact on cellular physiology and intracellular signalling from changes in the cell wall structure.
THE YEAST CELL WALL IS CONNECTED TO THE ACTIN CYTOSKELETON
Mechanosensors in S. cerevisiae and the link to actin
There is strong evidence that the cell wall and the actin cytoskeleton regulate/influence each other in yeast, and enzymatic removal of the cell wall (to form protoplasts) causes the actin cables to disappear and the actin patches to become evenly distributed over the cell surface [119]. The cell wall of S. cerevisiae displays nano-mechanical oscillating movements that are suggested to be caused by molecular motors in the cell [120], and such movement means that the cytoskeleton needs to be connected to the cell wall to allow force transfer to the cell wall [121]. Control of the actin cytoskeleton is also an important element in the compensatory response to weakening of the cell wall by altered external conditions, for example, by heat shock [71].
–
Like mammalian integrins, the yeast mechanosensor Wsc1 both controls and responds to the actin cytoskeleton [74]. Wsc1 acts as a regulator of rearrangements of the actin network during conditions of cell wall expansion and membrane stretching, and it is also a component of the cap, a complex structure located at the bud tip that is essential for the regulation of polarity of the actin cytoskeleton [122]. The actin patches are redistributed from the polarised growth sites to the cell periphery during cell wall stress, and this depolarisation is mediated by Wsc1 [74]. The actin network can also undergo depolarisation following hypo-osmotic shock, but at longer times of hypo-osmotic conditions repolarisation of actin occurs. Interestingly, wsc1 mutants show a marked delay in the repolarisation of the actin cytoskeleton after an osmotic shift, and about 40% of wsc1 mutant cells are still depolarised at 2 h after a hypo-osmotic shift [71]. The fact that loss of Wsc1 affects actin repolarisation suggests that Wsc1 might be an essential anchor site for actin. However, the Wsc1 protein does not appear to bind directly to actin, so how then is Wsc1 linked to the actin cytoskeleton?
–
Talin homologues in yeast connect Wsc1 to actin
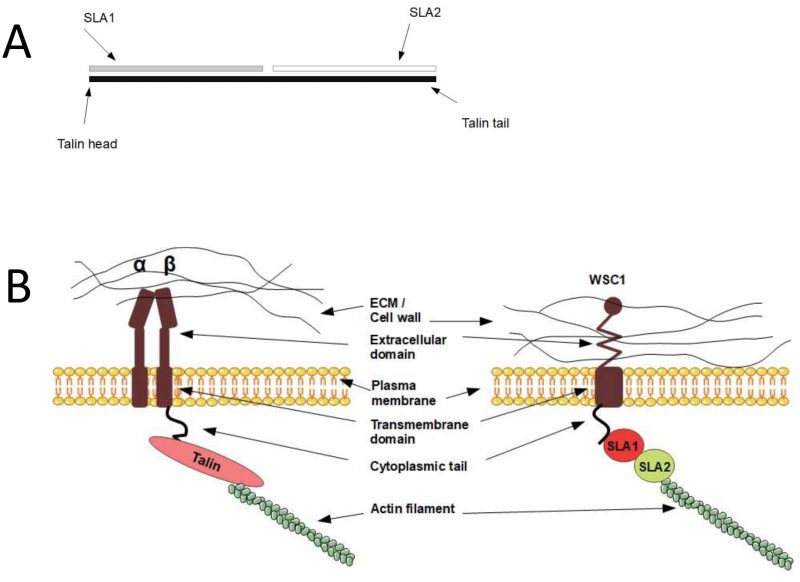
In mammals, integrin binds to the actin cytoskeleton via adapter proteins, of which talin is the most prominent (Fig. 1). The talin head binds to the cytoplasmic domain of integrin [123], while its tail binds the actin cytoskeleton [124]. In S. cerevisiae, the adapter protein Sla2 has been reported to possess a domain with talin homology [125]. Similarly to mammalian talin, Sla2 binds to actin via an I/LWEQ domain, and the deletion of this domain in Sla2 leads to defects in actin cytoskeleton morphology [126]. In contrast, overexpression of Sla2 is lethal and leads to the formation of thick actin cables [127]. Different studies suggest that Sla2 functions between the actin cytoskeleton and the endocytosis machinery [127][128][129][130]. In addition, Sla2 was identified together with Sla1 as synthetic-lethal with the actin-binding protein Abp1 [125], indicating related functions of Sla1 and Sla2. The protein Sla1 is important for proper localisation of Sla2 [131], and Sla1 can physically interact via its SHD1 domain with Wsc1 [76]. Sla1 binds Wsc1 to direct it into endocytosis [76], and this function of Sla1 is reminiscent of talin, which binds to integrin to keep it in an active conformation during endocytosis [132]. The fact that Sla1 and Sla2 physically interact [133][134] and function together suggests that they might act as a heterodimer that functions as an adapter between the actin cytoskeleton and Wsc1 in an analogous manner to mammalian talin. Surprisingly, by aligning the protein sequences of Sla1, Sla2, and human talin, we found that Sla1 aligns to the N-terminal part of human talin, while Sla2 aligns to the C-terminal part (Fig. 5A). Even though the sequence similarity is rather weak, this observation suggests that human talin is a fusion gene, i.e. a “Rosetta stone”, of Sla1 and Sla2. Taken together, yeast Sla1 and Sla2 might function together in a similar manner to human talin and physically connect Wsc1 to the actin cytoskeleton (Fig. 5B).
–
MECHANOSENSORS AND INTRACELLULAR SIGNALLING
Many components of integrin-dependent signalling in humans are conserved in yeast. For example, Sla1 and Sla2 are homologues of human talin (see above), the yeast Pxl1 is a homologue of human paxilin [135], Pfy1 is a homologue of human profilin [136], Tep1 is a homologue of PTEN [137], and Las17 is a homologue of WASP [138]. In mammals, the LIM domain of paxilin targets the protein to FACs [139], while in yeast it has been shown that the LIM domain in the paxilin-homolog Pxl1 targets the protein to sites of polarised growth [135]. Las17, Sla1, and Sla2 are components of the actin cortical patch in yeast [131][140], which suggests that the actin cortical patch might act as a yeast functional analogue to the human FAC.
–
Mammalian down-stream signalling pathways that receive input from integrins contain signalling components that have homologues in yeast. For example, the human ERK2 MAPK is a homolog of Kss1 in yeast, p38 MAPK is a homolog of Hog1, and ERK5 is a homolog of Slt2. The CWI pathway, which is the main signalling pathway responsible for maintaining cell wall homeostasis, has been extensively studied in S. cerevisiae [114]. As indicated above, the mechanosensors act as upstream sensors for the CWI pathway. A key component in the CWI pathway is the MAPK Slt2, which activates transcriptional responses to counteract cell wall stress mainly through phosphorylation of the Rlm1 transcription factor. Activated Slt2 also has a non-catalytic domain with which it forms a complex with Swi4/Swi6 and activates the FKS2 promoter (Fks2 is the catalytic subunit of β-1,3-glucan synthase) [141].
THE YEAST CELL WALL AND MECHANOSENSORS AS MEDIATORS OF VARIOUS EXTERNAL CONDITIONS
How yeast cells sense and respond to external perturbations are vital fields of investigation. Here we will give some examples in which the cell wall and the integrin functional analogues, the mechanosensors, play important roles in initiating intracellular signalling in response to changes in the external conditions, and we will provide connections to integrin-linked responses in mammals.
–
The heat shock response
S. cerevisiae responds to changes in temperature through drastic modifications of gene expression, and roughly 10% of all genes display significant changes in expression in response to elevated temperature [142]. Central to this transcriptional response in yeast is the heat shock transcription factor Hsf1. In addition to the effects of heat shock on internal cellular processes, thermal stress also appears to impact the cell surface. It has been shown that heat shock induces structural weakness in the cell wall that leads to activation of signalling via a putative cell wall elasticity/stiffness sensor [75][143]. In line with this, studies have reported on the importance of the yeast mechanosensors in the response to heat [74][88]. It was found that Wsc1, Wsc2, and Mid2 – and the GDP/GTP exchange factor Rom2 that interacts with these cell wall mechanosensors – are all multicopy suppressors of a temperature-sensitive hsf1 mutation [144]. Activation of the Pkc1 protein was furthermore shown to be necessary for suppression of the hsf1 mutation. This indicates that Hsf1 may act in parallel with the mechanosensors and Pkc1 to regulate cell wall remodelling in response to heat shock. In addition, overexpression of several other enzymes involved in cell wall organisation, like exo-1,3-β-glucanase (Exg1) and β-1,6-glucan synthase (Kre6), were also shown to suppress the temperature sensitivity of the hsf1 mutant. In addition, the hsf1 mutant was also suppressed in sorbitol medium that provides osmotic support, further highlighting the presence of a cell wall defect in the hsf1 mutant. It has also been reported that the mechanosensor triple mutant wsc1Δwsc2Δwsc3Δ is more sensitive to heat shock compared to wild type [145]. Taken together, there is ample genetic evidence that heat shock affects the organisation of the cell wall and that a proper cellular response to heat is dependent on the mechanosensors and down-stream signalling via Pkc1.
–
Heat shock has been shown to modulate several important properties of the yeast cell wall [146][147][148]. A shift from 30°C to 42°C induces the formation of a concentric circular ring on the yeast cell wall that appears after 20 minutes and reaches a diameter of 2–3 μm roughly 1 h after the temperature shift [146]. It has been shown that active cell growth is essential for the formation of the circular structure, and its appearance is accompanied by an increase of chitin in the cell wall and an increase in cell wall stiffness [146]. Heat shock also causes an increase in the β-1,6-glucan polysaccharide fraction and a partial reduction of β-1,3-glucan, both in S. cerevisiae and C. albicans [118]. All of these heat-induced cell wall changes might be detected by the mechanosensors and transmitted to intracellular signalling pathways, ultimately leading to drastic changes in gene expression and cell physiology/heat tolerance. We thus propose that the heat shock response in yeast is partly sensed by the mechanosensors, detecting heat-induced changes in the cell wall properties. Two possibilities arise: the first one is that heat affects membrane fluidity and that this change can be detected by the mechanosensors; membrane fluidity changes lead to a change in cell wall elasticity [149]. The second possibility is that heat affects the activity of pre-existing wall-editing enzymes, which leads to a change in cell wall elasticity. An analogy to this hypothesis is that osmostress causes an immediate change in cell wall elasticity by affecting wall editing enzymes [150]. The model is that upon heat shock and subsequent changes to the cell wall, Wsc1, Wsc2, and Mid2 activate Rom2 to promote GTP loading of Rho1, which in turn activates Pkc1 in the CWI pathway and leads to gene expression changes and cell wall remodelling.
–
The osmotic response
There is a large body of literature regarding osmostress on yeast, with extensive studies on several of the signalling pathways involved [151] and the multitude of genes and proteins that take part in the resulting physiological changes [152][153][154][155]. However, less attention has been devoted to the immediate morphological effects of the yeast cell wall in response to hyper-osmotic shock [156].
–
The yeast cell wall structure is modulated by osmostress. In C. albicans, cell wall remodelling enzymes change the cell wall elasticity, and a more elastic cell wall can be generated within seconds after osmotic stress [111]. Cells shifted to a hyper-osmotic culture medium exhibit fluorescent patches distributed irregularly over the cell surface when stained with cell-wall specific markers (e.g. calcofluor). These patches are proposed to be the result of redistribution of the pre-existing cell wall material into plasma membrane invaginations as a result of plasmolysis caused by osmotic dehydration [156]. Interestingly, this immediate osmo-response is not dependent on the HOG signalling pathway because hog1 and pbs2 mutant cells respond with similar cell wall patchiness as wild type cells. Yeasts growing on different carbon sources are differently sensitive to hyper-osmotic shock [157]. For example, lactate-grown yeast cells are more resistant than glucose-grown cells to osmotically induced volume changes, potentially due to structural differences in their respective cell walls [150]. The elevated resistance of lactate-grown cells correlates with reduced cell wall elasticity, as reflected in slower changes in cell volume following hyper-osmotic shock, and this is believed to lead to enhanced survival. These osmostress-induced changes in cell wall elasticity have also been confirmed by atomic force microscopy measurements of wild type and mutant cells and have been shown to reflect changes in the Young's modulus [150].
–
The MAPK Hog1 is the final component in the main signalling pathway that responds to hyper-osmotic stress in yeast. The human Hog1 orthologue p38 MAPK can functionally replace Hog1 in yeast [151], and the activation of p38 MAPK in response to mechanical stress is mediated through integrin [158]. In addition, cell swelling caused by starvation in human cells is sensed by integrin, which activates p38 MAPK [83][159]. So, could it be that the upstream activator of Hog1 in S. cerevisiae might be an integrin functional analogue? Indeed, the HOG pathway in yeast is activated by a large number of external stressors, including osmostress [160], low pH [161], zymolyase digestion of the cell wall [162], and microgravity [163], all of which affect the mechanical properties of the cell wall or the membrane-cell wall interaction. The HOG pathway in yeast is activated by two branches; the first is via Sln1 and the second is via Sho1, Hkr1, and Msb2 [164]. In the latter, the membrane protein Sho1 functions as an adapter in recruiting the kinases responsible for signalling in response to osmostress [165]. It has been shown in high-throughput studies that resistance to hyper-osmotic stress is decreased in WSC1 deletion mutants [166] and WSC2 deletion mutants [167]. The CWI pathway is activated and plays its main role in response to hypo-osmotic shock [168]. However, there are also reports of cooperative crosstalk between the HOG and CWI pathways suggesting that they jointly regulate the cell's response to zymolyase-induced cell wall damage. The HOG pathway shares some functions with the CWI pathway, and both are involved in cell wall biogenesis and are activated by hyper-osmotic shock, heat shock, oxidative stress, and zymolyase stress [169].
–
In conclusion, the cell wall and the mechanosensors appear to play important roles in several aspects of the osmo-response and to exert their effects through the HOG and CWI signalling pathways.
–
The starvation response
There are different signalling pathways in yeast that sense nutrient levels. There are several plasma membrane components involved in these processes, including receptors (Gpr1 and its associated down-stream Gα protein Gpa2), transporter-like sensors (Snf3 and Rgt2), and combined transporters and receptors (transceptors, e.g. Gap1). These nutrient sensors transmit their signals through various signal transduction pathways that involve central protein kinases like PKA, TORC1, and Sch9 [170]. In addition, both cyclic AMP (cAMP) and protons act as second messengers, and there are internal proteins, like the hexokinase Hxk2 and the AMP-activated S/T protein kinase Snf1, for intracellular sensing and signalling of the availability of glucose. The deletion of SNF1 causes sensitivity to cell wall stress agents and causes reduction in cell wall thickness in S. cervisiae [171]. Thus, the network of molecular players in the response to nutrients is rather complex. We will not cover all of the details of these pathways, but refer the reader to several recent extensive reviews that provide excellent overviews of this interesting field [170][172][173]. Here we will focus on additional aspects of nutrient signalling where the cell wall and the mechanosensors appear to be involved.
–
In mammals, integrins mediate cAMP signalling through a G-protein dependent pathway [174] and activate the cAMP pathway through the Gαs protein at FACs [175]. Gpa2 is the Gαs homologue in S. cerevisiae, and overexpression of Gpa2 in yeast increases cAMP levels [176]. Thus, Gpa2 in yeast can activate the cAMP pathway similarly to its mammalian homologue. An integrin-like mechanistic connection between the mechanosensors and cAMP production in yeast seems plausible, and it has been reported that Gpa2 physically interacts with the mechanosensors Wsc2, Wsc3, and Mid2 [133]. The central component in the glucose-sensing pathway in yeast is the RAS/PKA system [177]. cAMP is synthesised through adenylyl cyclase, the activity of which is stimulated by the GTP-bound Ras1 and Ras2 proteins. However, despite extensive studies regarding the above-mentioned receptors and signalling components, the mechanism by which glucose activates the RAS proteins is still not fully understood [172][173]. A potentially critical protein in glucose signalling is Gpa2, which along with its upstream receptor Gpr1 participates in the activation of PKA through the stimulation of adenylyl cyclase activity and the subsequent increase in cAMP production. However, the importance of Gpr1 and Gpa2 in glucose sensing has also been strongly questioned [173]. Several lines of evidence indicate that Gpr1 does not serve as a primary mediator in the acute response of cells to glucose; for example, deletion of Gpr1 or Gpa2 has no effect on the transcriptional response to glucose addition [177]. To further complicate things, it is clear that different mechanisms are involved in short-term and long-term responses to nutrient limitations, but the specifics of these mechanisms are not fully understood [172].
–
We propose here that the cell wall and the yeast mechanosensors might be part of yet another sensory/signalling pathway, compared to the ones described above, that is activated during nutrient starvation. The Mtl1 mechanosensor appears to be important in this respect, and it has been shown that Mtl1 activates the CWI pathway in response to glucose starvation [178]. Recent data indicate that Rho5, which is down-stream of the mechanosensors, might be involved in the response to glucose starvation [179], which gives further support for the involvement of the mechanosensors in nutrient sensing. Further mechanistic insights into this nutrient-sensing hypothesis comes from the fact that Wsc1, Wsc2, Wsc3, and Mtl1 have all been identified as inhibitors of the RAS2-cAMP pathway [145][180]. Petkova et al. showed that the mtl1 deletion mutant is deficient in the transcriptional induction of specific stress genes upon glucose deprivation [178]. These authors also observed that higher levels of the stress-induced transcription factor Msn2 are sufficient to rescue this glucose-dependent induction defect in the mtl1 deletion mutant in response to glucose starvation. Because the nuclear localisation of Msn2 is controlled by the RAS/PKA pathway and because mtl1 mutant cells show higher levels of cAMP compared to wild type, it was suggested that Mtl1 impacts on Msn2 activity by controlling cAMP levels through the down-regulation of Ras2. In line with this, Verna et al. found that deletion of RAS2 could rescue the heat shock-sensitive phenotype of the wsc1Δ strain, i.e. RAS2 and WSC1 showed a genetic interaction indicating that they are functionally linked [145]. Based on various experiments, including overexpression of Wsc1 in different mutant strains, the authors concluded that Wsc1 acts downstream of Ras1/2, perhaps modulating the activity of adenylate cyclase or the activity of PKA [145]. Thus, even if there are somewhat different working hypotheses for the mechanistic connection between the mechanosensors and cAMP/PKA, i.e. directly via inhibition of RAS proteins or of Gpa2, it is clear that the mechanosensors affect cAMP levels and PKA activity. Future experiments are needed to conclude which of the two mechanisms is the most prominent or if both connections exist and are equally important.
–
Another interesting observation is that the antifungal drug caspofungin, which inhibits the β-1,3-glucan synthase and in that way affects the cell wall, activates the CWI pathway involving Wsc1, Slt2, and Rom2 and inhibits the RAS/PKA pathway, the latter by decreasing cAMP levels (the decrease in cAMP levels was substantial and close to the cAMP decrease observed in a ras2Δ mutant [108]). The effects on RAS/PKA from caspofungin treatment were independent of the MAPK Slt2, but were dependent on Wsc1. This indicates that Wsc1 may transduce the change in the cell wall made by caspofungin through RAS/PKA, and that the effect on RAS/PKA is not an indirect consequence of MAPK signalling. Garcia et al. found that caspofungin treatment reduced levels of Ras2-GTP (the active form of Ras2) to similar levels as seen under conditions of glucose starvation [108]. Taken together, all of these results strongly argue in favour of a mechanistic connection between the mechanosensors and the RAS proteins leading to the de-activation of the RAS/PKA pathway in yeast during nutrient-poor conditions.
–
Further support for the involvement of the yeast cell wall in nutrient sensing comes from the fact that the chemical composition of the cell wall is modulated under various nutritional regimes, leading to changes in cell wall properties. When grown under NH4 limitation, the S. cerevisiae cell wall contains only half as much protein as cell walls from cells grown under glucose-limiting conditions [181]. In addition, under nitrogen limitation the cell walls are rigid and their inner layers look spongy under an electron microscope, while under glucose limitation cell walls are more elastic and appear more compact. The yeast Kluyveromyces lactis has a cell wall thickness of 64 nm when the carbon source is glucose, but it is 105 nm thick when the carbon source is ethanol. In addition, the cell walls of K. lactis grown on ethanol have a chemical structure that is more sensitive to breakdown by zymolyase [182]. Also, in C. albicans there are reports of changes in cell wall thickness and elasticity in response to changes in carbon source, and the thickness of the inner cell wall layer decreased from 75 nm to 22 nm when the carbon source was changed from glucose to lactate [111]. It has also been shown that cells grown on lactate display higher elasticity than cells grown on glucose [150]. We propose that these nutrient-induced changes in cell wall properties like thickness and elasticity are sensed by the mechanosensors that together with the cell wall form a sensory system for nutrient availability. In our model, the mechanosensors sense these nutrient-imposed cell wall changes and transmit signals intracellularly to activate various signalling pathways, in particular the RAS/PKA pathway.
–
The filamentous growth pathway
External factors can also induce developmental programs in yeast. In particular, starvation for some nutrients stimulates different developmental responses – nitrogen starvation induces mating [183], carbon starvation induces haploid and diploid yeast to enter the quiescent state [184], carbon starvation can stimulate invasive growth in haploid yeast [185], in diploid cells nitrogen starvation induces pseudohyphal growth, and starvation for both nitrogen and carbon stimulates sporulation [186][187]. However, it is still not completely clear how nutrient limitations cause all these developmental responses.
–
Filamentous growth is a regulated developmental response characterised by cell elongation, unipolar budding, physical attachment of mother and daughter cells, and increased adhesion to and invasion of growth substrates. Filamentous growth is a complex process which is regulated by many different signalling pathways and transcription factors [188]. The current model of filamentous growth signalling suggests the two mucins Msb2 and Hkr1 as the upstream components in the filamentous growth MAPK (FG-MAPK) pathway, that transmit the signal via Sho1 and through a signalling cascade involving Cdc24, Cdc42, Ste20, Ste50, Ste11 and Ste7 to reach the MAPK Kss1, which activity leads to the transcription of filamentous growth genes. In S. cerevisiae, filamentous differentiation is also positively correlated with the activity of the cAMP-PKA pathway, and increased intracellular cAMP levels or PKA activity result in increased filamentous growth [188]. Both the cAMP-PKA pathway and the FG-MAPK pathway are capable of inducing pseudohyphal growth in S. cerevisiae, and many important transcription factors and other targets are regulated in parallel by both pathways. Both cAMP-PKA signalling and the FG-MAPK pathway are regulated by Ras2, but it is not fully clear how Ras2 is activated by upstream components that respond to nutrient availability. Filamentous growth is also regulated by Tor1, and the Tor1 inhibitor rapamycin inhibits filamentous growth. This response is mediated through TAP42, a phosphatase in the TOR pathway, and the overexpression of Tap42 in cells treated with rapamycin restored filamentous growth [189]. In addition to the factors mentioned above, both the AMP-activated S/T protein kinase Snf1 and the transcription factor Rim101 take part in regulating filamentous growth [188].
–
A connection between filamentous growth and the mechanosensors has been established [190], and overexpression of mechanosensor genes was shown to induce expression of a filamentous growth-reporter (the transcription factor Fus1). In addition, overexpression of MID2 and WSC2 increased invasive growth, and deletion mutants of the mechanosensors (wsc1Δ, wsc2Δ, and to a lesser degree mid2Δ) exhibited defects in invasive growth. Because nutrient limitation alters the cell wall thickness and elasticity (see above), these changes in the cell wall might be part of the stimulus for the induction of filamentous growth. Haploid and diploid cells differ in terms of cell volume [191] and the thickness of their cell walls (4.18 µm and 5.3 μm, respectively) [191][192]. The difference in cell wall thickness and cell volume might have an impact on the distribution and the binding affinity between the wall mechanosensors and the cell wall and thus trigger the developmental changes. This hypothesis thus suggests that filamentous growth might also be affected by any factor that perturbs the cell wall dynamics independent of nutrient starvation. Indeed, in S. cerevisiae grown in rich medium with an adequate supply of nitrogen and other nutrients, both pseudohyphal formation and invasive growth (both of which are aspects of filamentous growth) were stimulated under certain stress conditions that affect the cell wall structure, such as increased temperature (37°C), osmotic stress (1 M NaCl), and drugs that perturb the glucan layer of the cell wall (i.e. Congo red) [187]. Conversely, it has also been shown that invasive growth is reduced in starved yeast cells under microgravity conditions, where microgravity decreases the mechanical stress on the cell wall [193].
–
In mammals, integrin regulates the activation of the ERK pathway in response to mechanical stress [194], and it has been shown that integrin-ECM interactions regulate ERK1/2 activity [195]. In S. cerevisiae, pseudohyphal growth is mediated by the KSS1 pathway [196], where the MAPK Kss1 has strong sequence similarity with human ERK1. A constitutively active form of human ERK1 induced pseudohyphal growth in S. cerevisiae [197].
–
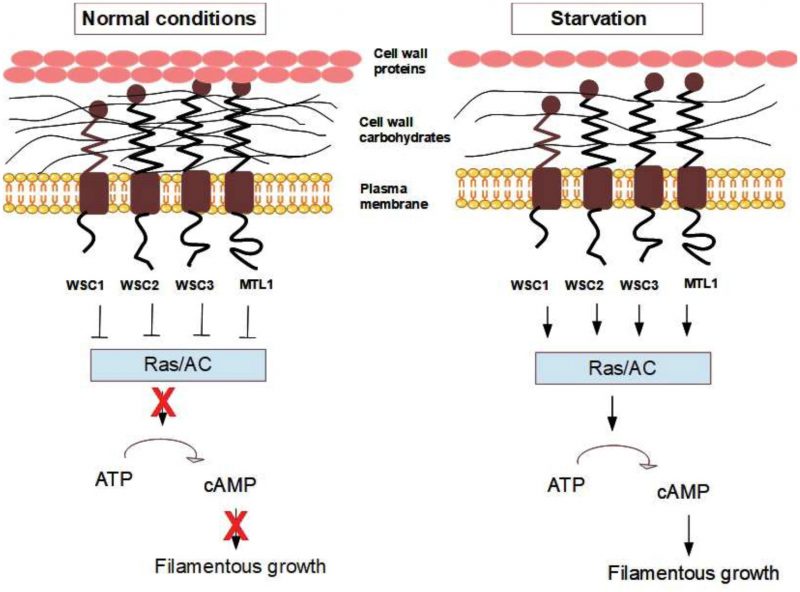
The current model states that the above-mentioned sensors and signalling mechanisms, e.g. Msb2, FG-MAPK, RAS/PKA and TOR, respond to nutrient limitations and trigger filamentous growth and in parallel change the cell wall [188]. However, we here propose an alternative model for nutrient-limitation signalling via cell wall changes and the mechanosensors during the initiation of filamentous growth. Yeast cells undergo filamentous growth when cAMP levels and PKA activity are high [198]. As indicated above, cAMP is produced by the enzyme adenylate cyclase, which is activated either by Gpa2 or Ras2. It has been shown that Gpa2 controls cell differentiation into filamentous growth in response to glucose starvation [199], and it has been reported that Gpa2 physically interacts with the mechanosensors Wsc2, Wsc3, and Mid2 [133]. Both the Wsc2 and Mid2 mechanosensors were identified as regulators of filamentous growth [190]. Moreover, overexpression of either Wsc1 and Wsc3 increased filamentous growth [200]. Together, these studies suggest that pseudohyphal growth in S. cerevisiae is probably at least partly regulated by an integrin-analogous signal in response to structural changes in the cell wall. In our hypothetical model (Fig. 6), the mechanosensors Wsc1, Wsc2, Wsc3, and Mtl1 (the ones identified as inhibitors of the RAS2-cAMP pathway [145][180]) inhibit RAS/adenylate cyclase under normal conditions, while changes in cell wall elasticity caused by starvation or mechanical stress might change the conformation of the cell wall sensors and remove this inhibition. In our model, this leads to the activation of the adenylate cyclase enzyme, increased cAMP levels, enhanced PKA activity, and the activation of the filamentous growth cascade.
–
CONCLUSIONS AND FUTURE CHALLENGES
In this review we discuss a general principle that seems adopted by all forms of life – stretching of either the cell wall or the plasma membrane generates a mechanical force on the extracellular part of a mechanosensor, and this stimulus is transmitted into a conformational change within the intracellular, cytoplasmic tail of the sensor and a subsequent activation of downstream signalling pathways and re-orientation of the actin cytoskeleton [201]. This is the main functional theme of the mammalian integrins. It is clear from sequence similarity that integrins exist throughout the metazoans, as well as in some unicellular organisms. Integrin-like proteins have been detected in fungi, but it should be stressed that these integrin-like proteins in fungi have no clear overall sequence similarity to their mammalian counterparts but possess shorter integrin-like motifs. They do have an integrin-like design, and thus they appear not to be evolutionarily related to integrins but to be functional analogues and to contain parts that have integrin-like features.
–
In the model yeast species S. cerevisiae there is no protein with sequence similarity to mammalian integrins. However, we propose here, as some other authors before us have done, that the mechanosensors in yeast are integrins in disguise – they are integrin functional analogues. We exemplify this with a number of experimental studies where the WSC-type and MID-type mechanosensors in yeast have been shown to resemble the integrins, both in terms of their mechanistic features and in their overall phenotypic consequences. We also propose that the cell wall is a prominent cellular feature of yeast that is involved in sensing/signalling pathways for a number of external factors. We believe that this is a generally neglected aspect of yeast physiology, and we propose that more in-depth studies are needed of the impact of the cell wall in relation to all aspects of yeast biology. In a hypothetical model, we propose that nutrient limitations modulate cell wall elasticity, which is sensed by the mechanosensors and results in developmental changes such as the induction of filamentous growth.
–
We also propose that the general importance of the mechanosensors in a number of central responses in yeast might have been missed because of functional redundancy among these proteins. Overall, there are five different genes encoding these mechanosensors in S. cerevisiae and few, if any, studies have really explored their functions in isolation. In this respect, it would be valuable to make penta-deletion mutants studying one mechanosensor at the time by reintroducing them individually. In the context of redundancy, it is interesting to note that other yeast species also harbour multiple redundant mechanosensor that complicates their functional characterisation, e.g. the milk yeast K. lactis has three mechanosensor paralogues [202]. In addition, the functions of the paralogues will most likely also be partly unique, where the different lengths of their extracellular domains might provide specificity in sensing different types of changes in cell wall elasticity and thickness. Finally, we would like to emphasise that the models proposed here, e.g. sensing glucose limitations, do not exclude the importance of other already well-known signalling pathways, but rather that they act in parallel and/or cooperate with each other.
–
We thus propose that there is still a lack of understanding of the cellular roles and molecular functions of the yeast mechanosensors. There is a long list of studies that can be proposed, and we will here only indicate some important ones. The Sla1 and Sla2 connection to human talin is interesting. Why is the long human talin protein split into two genes in yeast? Can the human talin protein compensate for the double mutant sla1Δsla2Δ in yeast? What would happen if one makes a fusion protein of SLA1 and SLA2 to form a protein more in line with human talin? Will that fusion hamper some of the functionality that these two proteins individually take part in in yeast? In line with this, we also believe it would be interesting to examine the dynamics of the actin cytoskeleton in response to mutations in SLA1 and SLA2. In particular, it would be revealing to study if the oscillatory movements observed by Pelling et al. [120] persist in SLA1 and SLA2 mutations or in WSC/MID gene deletions. These studies could either be conducted on the individual single deletion mutants of the mechanosensors or in the penta-deletion background expressing one of them at the time. These oscillatory studies would be interesting in order to get more mechanistic and functional information regarding the mechanosensor-actin connections and because the oscillatory phenomenon is still not fully understood.
–
Finally, another interesting research avenue would be to study the mechanosensors in relation to the cell cycle, and the presence of a “cell wall integrity checkpoint” to control cell cycle progression in relation to cell wall perturbations has been described (reviewed in [203]). A main feature of pseudohyphal growth is the delay in the G2/M phase, which allows daughter cells to grow to the same size as the mother cell before septum formation and cell separation [204]. We propose that this response might be regulated by the cell wall mechanosensors. Interestingly, defects in cell wall synthesis cause G2/M arrest, and this arrest in the cell cycle requires Wsc1 [205]. In mammals, integrin-mediated cell adhesion to the ECM regulates G2/M arrest [206]. We foresee that studies along these lines will deepen our mechanistic understanding of the connections between the cell cycle and the mechanosensors in yeast and will also have relevance for the understanding of the connection between integrins and the cell cycle in humans.
REFERENCES
- Kachroo AH, Laurent JM, Yellman CM, Meyer AG, Wilke CO, Marcotte EM (2015). Evolution. Systematic humanization of yeast genes reveals conserved functions and genetic modularity. Science 348(6237): 921-925. doi: 10.1126/science.aaa0769
- Mager WH, Winderickx J (2005). Yeast as a model for medical and medicinal research. Trends Pharmacol Sci 26(5): 265-273. doi: 10.1016/j.tips.2005.03.004
- Khurana V, Lindquist S (2010). Modelling neurodegeneration in Saccharomyces cerevisiae: why cook with baker's yeast? Nat Rev Neurosci 11(6): 436-449. doi: 10.1038/nrn2809
- McWhite CD, Liebeskind BJ, Marcotte EM (2015). Applications of comparative evolution to human disease genetics. Curr Opin Genet Dev 35: 16-24. doi: 10.1016/j.gde.2015.08.004
- Sebe-Pedros A, Roger AJ, Lang FB, King N, Ruiz-Trillo I (2010). Ancient origin of the integrin-mediated adhesion and signaling machinery. Proc Natl Acad Sci U S A 107(22): 10142-10147. doi: 10.1073/pnas.1002257107
- Pincus D, Letunic I, Bork P, Lim WA (2008). Evolution of the phospho-tyrosine signaling machinery in premetazoan lineages. Proc Natl Acad Sci U S A 105(28): 9680-9684. doi: 10.1073/pnas.0803161105
- Degnan BM, Vervoort M, Larroux C, Richards GS (2009). Early evolution of metazoan transcription factors. Curr Opin Genet Dev 19(6): 591-599. doi: 10.1016/j.gde.2009.09.008
- Abedin M, King N (2008). The premetazoan ancestry of cadherins. Science 319(5865): 946-948. doi: 10.1126/science.1151084
- Butcher EC, Picker LJ (1996). Lymphocyte homing and homeostasis. Science 272(5258): 60-66. doi: 10.1126/science.272.5258.60
- Luo BH, Carman CV, Springer TA (2007). Structural basis of integrin regulation and signaling. Annu Rev Immunol 25: 619-647. doi: 10.1146/annurev.immunol.25.022106.141618
- Ingber D (1991). Integrins as mechanochemical transducers. Curr Opin Cell Biol 3(5): 841-848. doi: 10.1016/0955-0674(91)90058-7
- Hynes RO (2004). The emergence of integrins: a personal and historical perspective. Matrix Biol 23(6): 333-340. doi: 10.1016/j.matbio.2004.08.001
- Tamkun JW, DeSimone DW, Fonda D, Patel RS, Buck C, Horwitz AF, Hynes RO (1986). Structure of integrin, a glycoprotein involved in the transmembrane linkage between fibronectin and actin. Cell 46(2): 271-282. doi: 10.1016/0092-8674(86)90744-0
- Laudanna C, Bolomini-Vittori M (2009). Integrin activation in the immune system. Wiley Interdiscip Rev Syst Biol Med 1(1): 116-127. doi: 10.1002/wsbm.9
- Cheresh DA, Stupack DG (2002). Integrin-mediated death: an explanation of the integrin-knockout phenotype? Nat Med 8(3): 193-194. doi: 10.1038/nm0302-193
- Koivisto L, Heino J, Hakkinen L, Larjava H (2014). Integrins in Wound Healing. Adv Wound Care 3(12): 762-783. doi: 10.1089/wound.2013.0436
- Desgrosellier JS, Cheresh DA (2010). Integrins in cancer: biological implications and therapeutic opportunities. Nat Rev Cancer 10(1): 9-22. doi: 10.1038/nrc2748
- Michie SA, Sytwu HK, McDevitt JO, Yang XD (1998). The roles of alpha 4-integrins in the development of insulin-dependent diabetes mellitus. Curr Top Microbiol Immunol 231: 65-83. doi: 10.1007/978-3-642-71987-5_5
- Caltagarone J, Jing Z, Bowser R (2007). Focal adhesions regulate Abeta signaling and cell death in Alzheimer's disease. Biochim Biophys Acta 1772(4): 438-445. doi: 10.1016/j.bbadis.2006.11.007
- Nishimura M, Kumsta C, Kaushik G, Diop SB, Ding Y, Bisharat-Kernizan J, Catan H, Cammarato A, Ross RS, Engler AJ, Bodmer R, Hansen M, Ocorr K (2014). A dual role for integrin-linked kinase and beta1-integrin in modulating cardiac aging. Aging Cell 13(3): 431-440. doi: 10.1111/acel.12193
- Takada Y, Ye X, Simon S (2007). The integrins. Genome Biol 8(5): 215. doi: 10.1186/gb-2007-8-5-215
- Paszek MJ, DuFort CC, Rossier O, Bainer R, Mouw JK, Godula K, Hudak JE, Lakins JN, Wijekoon AC, Cassereau L, Rubashkin MG, Magbanua MJ, Thorn KS, Davidson MW, Rugo HS, Park JW, Hammer DA, Giannone G, Bertozzi CR, Weaver VM (2014). The cancer glycocalyx mechanically primes integrin-mediated growth and survival. Nature 511(7509): 319-325. doi: 10.1038/nature13535
- Beglova N, Blacklow SC, Takagi J, Springer TA (2002). Cysteine-rich module structure reveals a fulcrum for integrin rearrangement upon activation. Nat Struct Biol 9(4): 282-287. doi: 10.1038/nsb779
- Whittaker CA, Hynes RO (2002). Distribution and evolution of von Willebrand/integrin A domains: widely dispersed domains with roles in cell adhesion and elsewhere. Mol Biol Cell 13(10): 3369-3387. doi: 10.1091/mbc.E02-05-0259
- Tuckwell D (1999). Evolution of von Willebrand factor A (VWA) domains. Biochem Soc Trans 27(6): 835-840. doi: 10.1042/bst0270835
- Emsley J, Knight CG, Farndale RW, Barnes MJ, Liddington RC (2000). Structural basis of collagen recognition by integrin alpha2beta1. Cell 101(1): 47-56. doi: 10.1016/S0092-8674(00)80622-4
- Humphries JD, Byron A, Humphries MJ (2006). Integrin ligands at a glance. J Cell Sci 119(Pt 19): 3901-3903. doi: 10.1242/jcs.03098
- Kim SH, Turnbull J, Guimond S (2011). Extracellular matrix and cell signalling: the dynamic cooperation of integrin, proteoglycan and growth factor receptor. J Endocrinol 209(2): 139-151. doi: 10.1530/JOE-10-0377
- Chiquet M, Renedo AS, Huber F, Fluck M (2003). How do fibroblasts translate mechanical signals into changes in extracellular matrix production? Matrix Biol 22(1): 73-80. doi: 10.1016/s0945-053x(03)00004-0
- Katsumi A, Orr AW, Tzima E, Schwartz MA (2004). Integrins in mechanotransduction. J Biol Chem 279(13): 12001-12004. doi: 10.1074/jbc.R300038200
- Shih YR, Tseng KF, Lai HY, Lin CH, Lee OK (2011). Matrix stiffness regulation of integrin-mediated mechanotransduction during osteogenic differentiation of human mesenchymal stem cells. J Bone Miner Res 26(4): 730-738. doi: 10.1002/jbmr.278
- Burridge K (2017). Focal adhesions: a personal perspective on a half century of progress. FEBS J 284(20): 3355-3361. doi: 10.1111/febs.14195
- Burridge K, Chrzanowska-Wodnicka M (1996). Focal adhesions, contractility, and signaling. Annu Rev Cell Dev Biol 12: 463-518. doi: 10.1146/annurev.cellbio.12.1.463
- Choi CK, Vicente-Manzanares M, Zareno J, Whitmore LA, Mogilner A, Horwitz AR (2008). Actin and alpha-actinin orchestrate the assembly and maturation of nascent adhesions in a myosin II motor-independent manner. Nat Cell Biol 10(9): 1039-1050. doi: 10.1038/ncb1763
- Calderwood DA, Campbell ID, Critchley DR (2013). Talins and kindlins: partners in integrin-mediated adhesion. Nat Rev Mol Cell Biol 14(8): 503-517. doi: 10.1038/nrm3624
- Case LB, Baird MA, Shtengel G, Campbell SL, Hess HF, Davidson MW, Waterman CM (2015). Molecular mechanism of vinculin activation and nanoscale spatial organization in focal adhesions. Nat Cell Biol 17(7): 880-892. doi: 10.1038/ncb3180
- Harburger DS, Calderwood DA (2009). Integrin signalling at a glance. J Cell Sci 122(Pt 2): 159-163. doi: 10.1242/jcs.018093
- Yee KL, Weaver VM, Hammer DA (2008). Integrin-mediated signalling through the MAP-kinase pathway. IET Syst Biol 2(1): 8-15. doi: 10.1049/iet-syb:20060058
- Shen B, Delaney MK, Du X (2012). Inside-out, outside-in, and inside-outside-in: G protein signaling in integrin-mediated cell adhesion, spreading, and retraction. Curr Opin Cell Biol 24(5): 600-606. doi: 10.1016/j.ceb.2012.08.011
- Berditchevski F (2001). Complexes of tetraspanins with integrins: more than meets the eye. J Cell Sci 114(Pt 23): 4143-4151. PMID: 11739647
- Qin J, Vinogradova O, Plow EF (2004). Integrin bidirectional signaling: a molecular view. PLoS Biol 2(6): e169. doi: 10.1371/journal.pbio.0020169
- Johnson MS, Lu N, Denessiouk K, Heino J, Gullberg D (2009). Integrins during evolution: evolutionary trees and model organisms. Biochim Biophys Acta 1788(4): 779-789. doi: 10.1016/j.bbamem.2008.12.013
- Chouhan B, Denesyuk A, Heino J, Johnson MS, Denessiouk K (2011). Conservation of the human integrin-type beta-propeller domain in bacteria. Plos One 6(10): e25069. doi: 10.1371/journal.pone.0025069
- Roberts K (1989). The plant extracellular matrix. Curr Opin Cell Biol 1(5): 1020-1027. PMID: 2697288
- Gens JS, Reuzeau C, Doolittle KW, McNally JG, Pickard BG (1996). Covisualization by computational optical-sectioning microscopy of integrin and associated proteins at the cell membrane of living onion protoplasts. Protoplasma 194(3-4): 215-230. doi: 10.1007/bf01882029
- Lynch TM, Lintilhac PM, Domozych D (1998). Mechanotransduction molecules in the plant gravisensory response: amyloplast/statolith membranes contain a beta 1 integrin-like protein. Protoplasma 201(1-2): 92-100. doi: 10.1007/bf01280715
- Swatzell LJ, Edelmann RE, Makaroff CA, Kiss JZ (1999). Integrin-like proteins are localized to plasma membrane fractions, not plastids, in Arabidopsis. Plant Cell Physiol 40(2): 173-183. doi: 10.1093/oxfordjournals.pcp.a029525
- Canut H, Carrasco A, Galaud JP, Cassan C, Bouyssou H, Vita N, Ferrara P, Pont-Lezica R (1998). High affinity RGD-binding sites at the plasma membrane of Arabidopsis thaliana links the cell wall. Plant J 16(1): 63-71. doi: 10.1046/j.1365-313x.1998.00276.x
- Mellersh DG, Heath MC (2001). Plasma membrane-cell wall adhesion is required for expression of plant defense responses during fungal penetration. Plant Cell 13(2): 413-424. doi: 10.2307/3871285
- Schindler M, Meiners S, Cheresh DA (1989). RGD-dependent linkage between plant cell wall and plasma membrane: consequences for growth. J Cell Biol 108(5): 1955-1965. doi: 10.1083/jcb.108.5.1955
- Hayashi T, Takagi S (2003). Ca2+-dependent cessation of cytoplasmic streaming induced by hypertonic treatment in Vallisneria mesophyll cells: possible role of cell wall-plasma membrane adhesion. Plant Cell Physiol 44(10): 1027-1036. doi: 10.1093/pcp/pcg123
- Wayne R, Staves MP, Leopold AC (1992). The contribution of the extracellular matrix to gravisensing in characean cells. J Cell Sci 101 (Pt 3): 611-623. PMID: 1522145
- Diaz-Sala C, Garrido G, Sabater B (2002). Age-related loss of rooting capability in Arabidopsis thaliana and its reversal by peptides containing the Arg-Gly-Asp (RGD) motif. Physiol Plant 114(4): 601-607. doi: 10.1034/j.1399-3054.2002.1140414.x
- Senchou V, Weide R, Carrasco A, Bouyssou H, Pont-Lezica R, Govers F, Canut H (2004). High affinity recognition of a Phytophthora protein by Arabidopsis via an RGD motif. Cell Mol Life Sci 61(4): 502-509. doi: 10.1007/s00018-003-3394-z
- Nuhse TS (2012). Cell wall integrity signaling and innate immunity in plants. Front Plant Sci 3: 280. doi: 10.3389/fpls.2012.00280
- Knepper C, Savory EA, Day B (2011). Arabidopsis NDR1 is an integrin-like protein with a role in fluid loss and plasma membrane-cell wall adhesion. Plant Physiol 156(1): 286-300. doi: 10.1104/pp.110.169656
- Mithoe SC, Boersema PJ, Berke L, Snel B, Heck AJ, Menke FL (2012). Targeted quantitative phosphoproteomics approach for the detection of phospho-tyrosine signaling in plants. J Proteome Res 11(1): 438-448. doi: 10.1021/pr200893k
- Becker B, Doan JM, Wustman B, Carpenter EJ, Chen L, Zhang Y, Wong GK, Melkonian M (2015). The Origin and Evolution of the Plant Cell Surface: Algal Integrin-Associated Proteins and a New Family of Integrin-Like Cytoskeleton-ECM Linker Proteins. Genome Biol Evol 7(6): 1580-1589. doi: 10.1093/gbe/evv089
- Chitcholtan K, Garrill A (2005). A beta4 integrin-like protein co-localises with a phosphotyrosine containing protein in the oomycete Achlya bisexualis: inhibition of tyrosine phosphorylation slows tip growth. Fungal Genet Biol 42(6): 534-545. doi: 10.1016/j.fgb.2005.03.010
- Jamerson M, da Rocha-Azevedo B, Cabral GA, Marciano-Cabral F (2012). Pathogenic Naegleria fowleri and non-pathogenic Naegleria lovaniensis exhibit differential adhesion to, and invasion of, extracellular matrix proteins. Microbiol 158(Pt 3): 791-803. doi: 10.1099/mic.0.055020-0
- Talamas-Rohana P, Hernandez-Ramirez VI, Perez-Garcia JN, Ventura-Juarez J (1998). Entamoeba histolytica contains a beta 1 integrin-like molecule similar to fibronectin receptors from eukaryotic cells. J Eukaryot Microbiol 45(3): 356-360. doi: 10.1111/j.1550-7408.1998.tb04549.x
- Vazquez J, Franco E, Reyes G, Meza I (1995). Characterization of adhesion plates induced by the interaction of Entamoeba histolytica trophozoites with fibronectin. Cell Motil Cytoskeleton 32(1): 37-45. doi: 10.1002/cm.970320105
- Flores-Robles D, Rosales C, Rosales-Encina JL, Talamas-Rohana P (2003). Entamoeba histolytica: a beta 1 integrin-like fibronectin receptor assembles a signaling complex similar to those of mammalian cells. Exp Parasitol 103(1-2): 8-15. doi: 10.1016/s0014-4894(03)00062-6
- Tucker SL, Talbot NJ (2001). Surface attachment and pre-penetration stage development by plant pathogenic fungi. Annu Rev Phytopathol 39(385-417. doi: 10.1146/annurev.phyto.39.1.385
- Tagliari L, Toledo MS, Lacerda TG, Suzuki E, Straus AH, Takahashi HK (2012). Membrane microdomain components of Histoplasma capsulatum yeast forms, and their role in alveolar macrophage infectivity. Biochim Biophys Acta 1818(3): 458-466. doi: 10.1016/j.bbamem.2011.12.008
- Bae CY, Kim S, Choi WB, Lee YH (2007). Involvement of extracellular matrix and integrin-like proteins on conidial adhesion and appressorium differentiation in Magnaporthe oryzae. J Microbiol Biotechnol 17(7): 1198-1203. PMID: 18051333
- Forsyth CB, Mathews HL (1996). Lymphocytes utilize CD11b/CD18 for adhesion to Candida albicans. Cell Immunol 170(1): 91-100. doi: 10.1006/cimm.1996.0138
- Santoni G, Spreghini E, Lucciarini R, Amantini C, Piccoli M (2001). Involvement of alpha(v)beta3 integrin-like receptor and glycosaminoglycans in Candida albicans germ tube adhesion to vitronectin and to a human endothelial cell line. Microb Pathog 31(4): 159-172. doi: 10.1006/mpat.2001.0459
- Gale C, Finkel D, Tao N, Meinke M, McClellan M, Olson J, Kendrick K, Hostetter M (1996). Cloning and expression of a gene encoding an integrin-like protein in Candida albicans. Proc Natl Acad Sci U S A 93(1): 357-361. doi: 10.1073/pnas.93.1.357
- Kottom TJ, Kennedy CC, Limper AH (2008). Pneumocystis PCINT1, a molecule with integrin-like features that mediates organism adhesion to fibronectin. Mol Microbiol 67(4): 747-761. doi: 10.1111/j.1365-2958.2007.06093.x
- Gualtieri T, Ragni E, Mizzi L, Fascio U, Popolo L (2004). The cell wall sensor Wsc1p is involved in reorganization of actin cytoskeleton in response to hypo-osmotic shock in Saccharomyces cerevisiae. Yeast 21(13): 1107-1120. doi: 10.1002/yea.1155
- Kock C, Dufrene YF, Heinisch JJ (2015). Up against the wall: is yeast cell wall integrity ensured by mechanosensing in plasma membrane microdomains? Appl Environ Microbiol 81(3): 806-811. doi: 10.1128/AEM.03273-14
- Schoenwaelder SM, Burridge K (1999). Bidirectional signaling between the cytoskeleton and integrins. Curr Opin Cell Biol 11(2): 274-286. doi: 10.1016/s0955-0674(99)80037-4
- Delley PA, Hall MN (1999). Cell wall stress depolarizes cell growth via hyperactivation of RHO1. J Cell Biol 147(1): 163-174. doi: 10.1083/jcb.147.1.163
- Bickle M, Delley PA, Schmidt A, Hall MN (1998). Cell wall integrity modulates RHO1 activity via the exchange factor ROM2. EMBO J 17(8): 2235-2245. doi: 10.1093/emboj/17.8.2235
- Piao HL, Machado IM, Payne GS (2007). NPFXD-mediated endocytosis is required for polarity and function of a yeast cell wall stress sensor. Mol Biol Cell 18(1): 57-65. doi: 10.1091/mbc.E06-08-0721
- Heinisch JJ, Rodicio R (2018). Protein kinase C in fungi-more than just cell wall integrity. FEMS Microbiol Rev 42(1). doi: 10.1093/femsre/fux051
- Schmitz HP, Huppert S, Lorberg A, Heinisch JJ (2002). Rho5p downregulates the yeast cell integrity pathway. J Cell Sci 115(Pt 15): 3139-3148. PMID: 12118069
- Fernandez-Minan A, Cobreros L, Gonzalez-Reyes A, Martin-Bermudo MD (2008). Integrins contribute to the establishment and maintenance of cell polarity in the follicular epithelium of the Drosophila ovary. Int J Dev Biol 52(7): 925-932. doi: 10.1387/ijdb.072418af
- Zhu P, Liu X, Labelle EF, Freedman BD (2005). Mechanisms of hypotonicity-induced calcium signaling and integrin activation by arachidonic acid-derived inflammatory mediators in B cells. J Immunol 175(8): 4981-4989. doi: 10.4049/jimmunol.175.8.4981
- Defilippi P, Olivo C, Venturino M, Dolce L, Silengo L, Tarone G (1999). Actin cytoskeleton organization in response to integrin-mediated adhesion. Microsc Res Tech 47(1): 67-78. doi: Wei H, Vander Heide RS (2010). Ischemic preconditioning and heat shock activate Akt via a focal adhesion kinase-mediated pathway in Langendorff-perfused adult rat hearts. Am J Physiol Heart Circ Physiol 298(1): H152-157. doi: 10.1152/ajpheart.00613.2009
- vom Dahl S, Schliess F, Reissmann R, Gorg B, Weiergraber O, Kocalkova M, Dombrowski F, Haussinger D (2003). Involvement of integrins in osmosensing and signaling toward autophagic proteolysis in rat liver. J Biol Chem 278(29): 27088-27095. doi: 10.1074/jbc.M210699200
- Marre ML, Petnicki-Ocwieja T, DeFrancesco AS, Darcy CT, Hu LT (2010). Human integrin alpha(3)beta(1) regulates TLR2 recognition of lipopeptides from endosomal compartments. Plos One 5(9): e12871. doi: 10.1371/journal.pone.0012871
- Ueno K, Saito M, Nagashima M, Kojima A, Nishinoaki S, Toshima JY, Toshima J (2014). V-ATPase-dependent luminal acidification is required for endocytic recycling of a yeast cell wall stress sensor, Wsc1p. Biochem Biophys Res Commun 443(2): 549-555. doi: 10.1016/j.bbrc.2013.12.008
- Merchan S, Bernal D, Serrano R, Yenush L (2004). Response of the Saccharomyces cerevisiae Mpk1 mitogen-activated protein kinase pathway to increases in internal turgor pressure caused by loss of Ppz protein phosphatases. Eukaryot Cell 3(1): 100-107. doi: 10.1128/ec.3.1.100-107.2004
- deHart AK, Schnell JD, Allen DA, Tsai JY, Hicke L (2003). Receptor internalization in yeast requires the Tor2-Rho1 signaling pathway. Mol Biol Cell 14(11): 4676-4684. doi: 10.1091/mbc.E03-05-0323
- Ketela T, Green R, Bussey H (1999). Saccharomyces cerevisiae mid2p is a potential cell wall stress sensor and upstream activator of the PKC1-MPK1 cell integrity pathway. J Bacteriol 181(11): 3330-3340. PMID: 10348843
- Mao K, Klionsky DJ (2011). MAPKs regulate mitophagy in Saccharomyces cerevisiae. Autophagy 7(12): 1564-1565. doi: 10.4161/auto.7.12.17971
- Burtner CR, Murakami CJ, Olsen B, Kennedy BK, Kaeberlein M (2011). A genomic analysis of chronological longevity factors in budding yeast. Cell Cycle 10(9): 1385-1396. doi: 10.4161/cc.10.9.15464
- Garay E, Campos SE, Gonzalez de la Cruz J, Gaspar AP, Jinich A, Deluna A (2014). High-resolution profiling of stationary-phase survival reveals yeast longevity factors and their genetic interactions. PLoS Genet 10(2): e1004168. doi: 10.1371/journal.pgen.1004168
- Wilk S, Wittland J, Thywissen A, Schmitz HP, Heinisch JJ (2010). A block of endocytosis of the yeast cell wall integrity sensors Wsc1 and Wsc2 results in reduced fitness in vivo. Mol Genet Genomics 284(3): 217-229. doi: 10.1007/s00438-010-0563-2
- Hoyt DG, Rizzo M, Gerritsen ME, Pitt BR, Lazo JS (1997). Integrin activation protects pulmonary endothelial cells from the genotoxic effects of bleomycin. Am J Physiol 273(3 Pt 1): L612-617. doi: 10.1152/ajplung.1997.273.3.L612
- Kapitzky L, Beltrao P, Berens TJ, Gassner N, Zhou C, Wuster A, Wu J, Babu MM, Elledge SJ, Toczyski D, Lokey RS, Krogan NJ (2010). Cross-species chemogenomic profiling reveals evolutionarily conserved drug mode of action. Mol Syst Biol 6: 451. doi: 10.1038/msb.2010.107
- Jacoby JJ, Nilius SM, Heinisch JJ (1998). A screen for upstream components of the yeast protein kinase C signal transduction pathway identifies the product of the SLG1 gene. Mol Gen Genet 258(1-2): 148-155. doi: 10.1007/s004380050717
- Severin FF, Hyman AA (2002). Pheromone induces programmed cell death in S. cerevisiae. Curr Biol 12(7): R233-235. doi: 10.1016/s0960-9822(02)00776-5
- Schmitz HP, Jendretzki A, Wittland J, Wiechert J, Heinisch JJ (2015). Identification of Dck1 and Lmo1 as upstream regulators of the small GTPase Rho5 in Saccharomyces cerevisiae. Mol Microbiol 96(2): 306-324. doi: 10.1111/mmi.12937
- Ruoslahti E, Reed JC (1994). Anchorage dependence, integrins, and apoptosis. Cell 77(4): 477-478. doi: 10.1016/0092-8674(94)90209-7
- Smits GJ, Kapteyn JC, van den Ende H, Klis FM (1999). Cell wall dynamics in yeast. Curr Opin Microbiol 2(4): 348-352. PMID: 10458981
- Kapteyn JC, Van Den Ende H, Klis FM (1999). The contribution of cell wall proteins to the organization of the yeast cell wall. Biochim Biophys Acta 1426(2): 373-383. doi: 10.1016/s0304-4165(98)00137-8
- Orlean P (2012). Architecture and biosynthesis of the Saccharomyces cerevisiae cell wall. Genetics 192(3): 775-818. doi: 10.1534/genetics.112.144485
- Dupres V, Dufrene YF, Heinisch JJ (2010). Measuring cell wall thickness in living yeast cells using single molecular rulers. ACS Nano 4(9): 5498-5504. doi: 10.1021/nn101598v
- Yamaguchi M, Namiki Y, Okada H, Mori Y, Furukawa H, Wang J, Ohkusu M, Kawamoto S (2011). Structome of Saccharomyces cerevisiae determined by freeze-substitution and serial ultrathin-sectioning electron microscopy. J Electron Microsc (Tokyo) 60(5): 321-335. doi: 10.1093/jmicro/dfr052
- Sekiya-Kawasaki M, Abe M, Saka A, Watanabe D, Kono K, Minemura-Asakawa M, Ishihara S, Watanabe T, Ohya Y (2002). Dissection of upstream regulatory components of the Rho1p effector, 1,3-beta-glucan synthase, in Saccharomyces cerevisiae. Genetics 162(2): 663-676. PMID: 12399379
- Kollar R, Petrakova E, Ashwell G, Robbins PW, Cabib E (1995). Architecture of the yeast cell wall. The linkage between chitin and beta(1–>3)-glucan. J Biol Chem 270(3): 1170-1178. doi: 10.1074/jbc.270.3.1170
- Munro CA, Winter K, Buchan A, Henry K, Becker JM, Brown AJ, Bulawa CE, Gow NA (2001). Chs1 of Candida albicans is an essential chitin synthase required for synthesis of the septum and for cell integrity. Mol Microbiol 39(5): 1414-1426. doi: 10.1046/j.1365-2958.2001.02347.x
- Kapteyn JC, Ram AF, Groos EM, Kollar R, Montijn RC, Van Den Ende H, Llobell A, Cabib E, Klis FM (1997). Altered extent of cross-linking of beta1,6-glucosylated mannoproteins to chitin in Saccharomyces cerevisiae mutants with reduced cell wall beta1,3-glucan content. J Bacteriol 179(20): 6279-6284. doi: 10.1128/jb.179.20.6279-6284.1997
- Garcia R, Bravo E, Diez-Muniz S, Nombela C, Rodriguez-Pena JM, Arroyo J (2017). A novel connection between the Cell Wall Integrity and the PKA pathways regulates cell wall stress response in yeast. Sci Rep 7(1): 5703. doi: 10.1038/s41598-017-06001-9
- Bowman SM, Free SJ (2006). The structure and synthesis of the fungal cell wall. Bioessays 28(8): 799-808. doi: 10.1002/bies.20441
- Arroyo J, Farkas V, Sanz AB, Cabib E (2016). 'Strengthening the fungal cell wall through chitin-glucan cross-links: effects on morphogenesis and cell integrity'. Cell Microbiol 18(9): 1239-1250. doi: 10.1111/cmi.12615
- Ene IV, Heilmann CJ, Sorgo AG, Walker LA, de Koster CG, Munro CA, Klis FM, Brown AJ (2012). Carbon source-induced reprogramming of the cell wall proteome and secretome modulates the adherence and drug resistance of the fungal pathogen Candida albicans. Proteomics 12(21): 3164-3179. doi: 10.1002/pmic.201200228
- Roncero C, Duran A (1985). Effect of Calcofluor white and Congo red on fungal cell wall morphogenesis: in vivo activation of chitin polymerization. J Bacteriol 163(3): 1180-1185. PMID: 3897187
- Roncero C, Valdivieso MH, Ribas JC, Duran A (1988). Effect of calcofluor white on chitin synthases from Saccharomyces cerevisiae. J Bacteriol 170(4): 1945-1949. doi: 10.1128/jb.170.4.1945-1949.1988
- Levin DE (2011). Regulation of cell wall biogenesis in Saccharomyces cerevisiae: the cell wall integrity signaling pathway. Genetics 189(4): 1145-1175. doi: 10.1534/genetics.111.128264
- Reinke A, Chen JC, Aronova S, Powers T (2006). Caffeine targets TOR complex I and provides evidence for a regulatory link between the FRB and kinase domains of Tor1p. J Biol Chem 281(42): 31616-31626. doi: 10.1074/jbc.M603107200
- Kuranda K, Leberre V, Sokol S, Palamarczyk G, Francois J (2006). Investigating the caffeine effects in the yeast Saccharomyces cerevisiae brings new insights into the connection between TOR, PKC and Ras/cAMP signalling pathways. Mol Microbiol 61(5): 1147-1166. doi: 10.1111/j.1365-2958.2006.05300.x
- Deresinski SC, Stevens DA (2003). Caspofungin. Clin Infect Dis 36(11): 1445-1457. doi: 10.1086/375080
- Schiavone M, Vax A, Formosa C, Martin-Yken H, Dague E, Francois JM (2014). A combined chemical and enzymatic method to determine quantitatively the polysaccharide components in the cell wall of yeasts. FEMS Yeast Res 14(6): 933-947. doi: 10.1111/1567-1364.12182
- Gabriel M, Kopecká M, Svoboda A (1992). Structural rearrangement of the actin cytoskeleton in regenerating protoplasts of budding yeasts. J Gen Microbiol 138: 2229-2234. doi: 10.1099/00221287-138-10-2229
- Pelling AE, Sehati S, Gralla EB, Valentine JS, Gimzewski JK (2004). Local nanomechanical motion of the cell wall of Saccharomyces cerevisiae. Science 305(5687): 1147-1150. doi: 10.1126/science.1097640
- Kumamoto CA (2008). Molecular mechanisms of mechanosensing and their roles in fungal contact sensing. Nat Rev Microbiol 6(9): 667-673. doi: 10.1038/nrmicro1960
- Pruyne D, Bretscher A (2000). Polarization of cell growth in yeast. J Cell Sci 113 (Pt 4): 571-585. PMID: 10652251
- Calderwood DA, Zent R, Grant R, Rees DJ, Hynes RO, Ginsberg MH (1999). The Talin head domain binds to integrin beta subunit cytoplasmic tails and regulates integrin activation. J Biol Chem 274(40): 28071-28074. doi: 10.1074/jbc.274.40.28071
- Goldmann WH, Guttenberg Z, Kaufmann S, Hess D, Ezzell RM, Isenberg G (1997). Examining F-actin interaction with intact talin and talin head and tail fragment using static and dynamic light scattering. Eur J Biochem 250(2): 447-450. doi: 10.1111/j.1432-1033.1997.0447a.x
- Holtzman DA, Yang S, Drubin DG (1993). Synthetic-lethal interactions identify two novel genes, SLA1 and SLA2, that control membrane cytoskeleton assembly in Saccharomyces cerevisiae. J Cell Biol 122(3): 635-644. doi: 10.1083/jcb.122.3.635
- McCann RO, Craig SW (1999). Functional genomic analysis reveals the utility of the I/LWEQ module as a predictor of protein:actin interaction. Biochem Biophys Res Commun 266(1): 135-140. doi: 10.1006/bbrc.1999.1776
- Yang S, Cope MJ, Drubin DG (1999). Sla2p is associated with the yeast cortical actin cytoskeleton via redundant localization signals. Mol Biol Cell 10(7): 2265-2283. doi: 10.1091/mbc.10.7.2265
- McCann RO, Craig SW (1997). The I/LWEQ module: a conserved sequence that signifies F-actin binding in functionally diverse proteins from yeast to mammals. Proc Natl Acad Sci U S A 94(11): 5679-5684. doi: 10.1073/pnas.94.11.5679
- Wesp A, Hicke L, Palecek J, Lombardi R, Aust T, Munn AL, Riezman H (1997). End4p/Sla2p interacts with actin-associated proteins for endocytosis in Saccharomyces cerevisiae. Mol Biol Cell 8(11): 2291-2306. doi: 10.1091/mbc.8.11.2291
- Baggett JJ, D'Aquino KE, Wendland B (2003). The Sla2p talin domain plays a role in endocytosis in Saccharomyces cerevisiae. Genetics 165(4): 1661-1674. PMID: 14704157
- Ayscough KR, Eby JJ, Lila T, Dewar H, Kozminski KG, Drubin DG (1999). Sla1p is a functionally modular component of the yeast cortical actin cytoskeleton required for correct localization of both Rho1p-GTPase and Sla2p, a protein with talin homology. Mol Biol Cell 10(4): 1061-1075. doi: 10.1091/mbc.10.4.1061
- Nader GP, Ezratty EJ, Gundersen GG (2016). FAK, talin and PIPKIgamma regulate endocytosed integrin activation to polarize focal adhesion assembly. Nat Cell Biol 18(5): 491-503. doi: 10.1038/ncb3333
- Tarassov K, Messier V, Landry CR, Radinovic S, Serna Molina MM, Shames I, Malitskaya Y, Vogel J, Bussey H, Michnick SW (2008). An in vivo map of the yeast protein interactome. Science 320(5882): 1465-1470. doi: 10.1126/science.1153878
- Gourlay CW, Dewar H, Warren DT, Costa R, Satish N, Ayscough KR (2003). An interaction between Sla1p and Sla2p plays a role in regulating actin dynamics and endocytosis in budding yeast. J Cell Sci 116(Pt 12): 2551-2564. doi: 10.1242/jcs.00454
- Mackin NA, Sousou TJ, Erdman SE (2004). The PXL1 gene of Saccharomyces cerevisiae encodes a paxillin-like protein functioning in polarized cell growth. Mol Biol Cell 15(4): 1904-1917. doi: 10.1091/mbc.E04-01-0004
- Ostrander DB, Ernst EG, Lavoie TB, Gorman JA (1999). Polyproline binding is an essential function of human profilin in yeast. Eur J Biochem 262(1): 26-35. doi: 10.1046/j.1432-1327.1999.00354.x
- Heymont J, Berenfeld L, Collins J, Kaganovich A, Maynes B, Moulin A, Ratskovskaya I, Poon PP, Johnston GC, Kamenetsky M, DeSilva J, Sun H, Petsko GA, Engebrecht J (2000). TEP1, the yeast homolog of the human tumor suppressor gene PTEN/MMAC1/TEP1, is linked to the phosphatidylinositol pathway and plays a role in the developmental process of sporulation. Proc Natl Acad Sci U S A 97(23): 12672-12677. doi: 10.1073/pnas.97.23.12672
- Urbanek AN, Smith AP, Allwood EG, Booth WI, Ayscough KR (2013). A novel actin-binding motif in Las17/WASP nucleates actin filaments independently of Arp2/3. Curr Biol 23(3): 196-203. doi: 10.1016/j.cub.2012.12.024
- Brown MC, Perrotta JA, Turner CE (1996). Identification of LIM3 as the principal determinant of paxillin focal adhesion localization and characterization of a novel motif on paxillin directing vinculin and focal adhesion kinase binding. J Cell Biol 135(4): 1109-1123. doi: 10.1083/jcb.135.4.1109
- Galletta BJ, Chuang DY, Cooper JA (2008). Distinct roles for Arp2/3 regulators in actin assembly and endocytosis. PLoS biology 6(1): e1. doi: 10.1371/journal.pbio.0060001
- Kim KY, Truman AW, Levin DE (2008). Yeast Mpk1 mitogen-activated protein kinase activates transcription through Swi4/Swi6 by a noncatalytic mechanism that requires upstream signal. Mol Cell Biol 28(8): 2579-2589. doi: 10.1128/MCB.01795-07
- Gasch AP, Spellman PT, Kao CM, Carmel-Harel O, Eisen MB, Storz G, Botstein D, Brown PO (2000). Genomic expression programs in the response of yeast cells to environmental changes. Mol Cell Biol 11(12): 4241-4257. doi: 10.1091/mbc.11.12.4241
- Kamada Y, Jung US, Piotrowski J, Levin DE (1995). The protein kinase C-activated MAP kinase pathway of Saccharomyces cerevisiae mediates a novel aspect of the heat shock response. Genes Dev 9(13): 1559-1571. doi: 10.1101/gad.9.13.1559
- Imazu H, Sakurai H (2005). Saccharomyces cerevisiae heat shock transcription factor regulates cell wall remodeling in response to heat shock. Eukaryot Cell 4(6): 1050-1056. doi: 10.1128/EC.4.6.1050-1056.2005
- Verna J, Lodder A, Lee K, Vagts A, Ballester R (1997). A family of genes required for maintenance of cell wall integrity and for the stress response in Saccharomyces cerevisiae. Proc Natl Acad Sci U S A 94(25): 13804-13809. doi: 10.1073/pnas.94.25.13804
- Pillet F, Lemonier S, Schiavone M, Formosa C, Martin-Yken H, Francois JM, Dague E (2014). Uncovering by atomic force microscopy of an original circular structure at the yeast cell surface in response to heat shock. BMC Biol 12: 6. doi: 10.1186/1741-7007-12-6
- Dague E, Jauvert E, Laplatine L, Viallet B, Thibault C, Ressier L (2011). Assembly of live micro-organisms on microstructured PDMS stamps by convective/capillary deposition for AFM bio-experiments. Nanotechnology 22(39): 395102. doi: 10.1088/0957-4484/22/39/395102
- Francois JM, Formosa C, Schiavone M, Pillet F, Martin-Yken H, Dague E (2013). Use of atomic force microscopy (AFM) to explore cell wall properties and response to stress in the yeast Saccharomyces cerevisiae. Curr Genet 59(4): 187-196. doi: 10.1007/s00294-013-0411-0
- Schiavone M, Formosa-Dague C, Elsztein C, Teste MA, Martin-Yken H, De Morais MA, Jr., Dague E, Francois JM (2016). Evidence for a Role for the Plasma Membrane in the Nanomechanical Properties of the Cell Wall as Revealed by an Atomic Force Microscopy Study of the Response of Saccharomyces cerevisiae to Ethanol Stress. Appl Environ Microbiol 82(15): 4789-4801. doi: 10.1128/AEM.01213-16
- Ene IV, Walker LA, Schiavone M, Lee KK, Martin-Yken H, Dague E, Gow NA, Munro CA, Brown AJ (2015). Cell Wall Remodeling Enzymes Modulate Fungal Cell Wall Elasticity and Osmotic Stress Resistance. MBio 6(4): e00986. doi: 10.1128/mBio.00986-15
- Saito H, Posas F (2012). Response to hyperosmotic stress. Genetics 192(2): 289-318. doi: 10.1534/genetics.112.140863
- Blomberg A, Adler L (1992). Physiology of osmotolerance in fungi. Adv Microbial Phys 33: 145-212. doi: 10.1016/s0065-2911(08)60217-9
- Hohmann S (2002). Osmotic stress signalling and osmoadaptation in yeasts. Microbiol Mol Biol Rev 66: 300-372. doi: 10.1128/mmbr.66.2.300-372.2002
- Warringer J, Ericson E, Fernandez L, Nerman O, Blomberg A (2003). High-resolution yeast phenomics resolves different physiological features in the saline response. Proc Natl Acad Sci U S A 100(26): 15724-15729. doi: 10.1073/pnas.2435976100
- Blomberg A (1995). Global changes in protein synthesis during adaptation of the yeast Saccharomyces cerevisiae to 0.7 M NaCl. J Bacteriol 177(12): 3563-3572. doi: 10.1128/jb.177.12.3563-3572.1995
- Slaninova I, Sestak S, Svoboda A, Farkas V (2000). Cell wall and cytoskeleton reorganization as the response to hyperosmotic shock in Saccharomyces cerevisiae. Arch Microbiol 173(4): 245-252. doi: 10.1007/s002030000136
- Blomberg A (1997). The osmotic hypersensitivity of the yeast Saccharomyces cerevisiae is strain and growth media dependent: quantitative aspects of the phenomenon. Yeast 13(6): 529-539. doi: Aikawa R, Nagai T, Kudoh S, Zou Y, Tanaka M, Tamura M, Akazawa H, Takano H, Nagai R, Komuro I (2002). Integrins play a critical role in mechanical stress-induced p38 MAPK activation. Hypertension 39(2): 233-238. doi: 10.1161/hy0202.102699
- Schliess F, Reissmann R, Reinehr R, vom Dahl S, Haussinger D (2004). Involvement of integrins and Src in insulin signaling toward autophagic proteolysis in rat liver. J Biol Chem 279(20): 21294-21301. doi: 10.1074/jbc.M313901200
- Schaber J, Adrover MA, Eriksson E, Pelet S, Petelenz-Kurdziel E, Klein D, Posas F, Goksor M, Peter M, Hohmann S, Klipp E (2010). Biophysical properties of Saccharomyces cerevisiae and their relationship with HOG pathway activation. Eur Biophys J 39(11): 1547-1556. doi: 10.1007/s00249-010-0612-0
- Kapteyn JC, ter Riet B, Vink E, Blad S, De Nobel H, Van Den Ende H, Klis FM (2001). Low external pH induces HOG1-dependent changes in the organization of the Saccharomyces cerevisiae cell wall. Mol Microbiol 39(2): 469-479. doi: 10.1046/j.1365-2958.2001.02242.x
- Bermejo C, Rodriguez E, Garcia R, Rodriguez-Pena JM, Rodriguez de la Concepcion ML, Rivas C, Arias P, Nombela C, Posas F, Arroyo J (2008). The sequential activation of the yeast HOG and SLT2 pathways is required for cell survival to cell wall stress. Mol Biol Cell 19(3): 1113-1124. doi: 10.1091/mbc.E07-08-0742
- Bradamante S, Villa A, Versari S, Barenghi L, Orlandi I, Vai M (2010). Oxidative stress and alterations in actin cytoskeleton trigger glutathione efflux in Saccharomyces cerevisiae. Biochim Biophys Acta 1803(12): 1376-1385. doi: 10.1016/j.bbamcr.2010.07.007
- Tatebayashi K, Tanaka K, Yang HY, Yamamoto K, Matsushita Y, Tomida T, Imai M, Saito H (2007). Transmembrane mucins Hkr1 and Msb2 are putative osmosensors in the SHO1 branch of yeast HOG pathway. EMBO J 26(15): 3521-3533. doi: 10.1038/sj.emboj.7601796
- Roman E, Nombela C, Pla J (2005). The Sho1 adaptor protein links oxidative stress to morphogenesis and cell wall biosynthesis in the fungal pathogen Candida albicans. Mol Cell Biol 25(23): 10611-10627. doi: 10.1128/MCB.25.23.10611-10627.2005
- Dudley AM, Janse DM, Tanay A, Shamir R, Church GM (2005). A global view of pleiotropy and phenotypically derived gene function in yeast. Mol Syst Biol 1: 2005.0001. doi: 10.1038/msb4100004
- Yoshikawa K, Tanaka T, Furusawa C, Nagahisa K, Hirasawa T, Shimizu H (2009). Comprehensive phenotypic analysis for identification of genes affecting growth under ethanol stress in Saccharomyces cerevisiae. FEMS Yeast Res 9(1): 32-44. doi: 10.1111/j.1567-1364.2008.00456.x
- Davenport KR, Sohaskey M, Kamada Y, Levins DE, Gustin MC (1995). A second osmosensing signal transduction pathway in yeast: hypotonic shock activates the PKC1 protein kinase-regulated cell integrety pathway. J Biol Chem 270 (59): 30157-30161. doi: 10.1074/jbc.270.50.30157
- Rodriguez-Pena JM, Garcia R, Nombela C, Arroyo J (2010). The high-osmolarity glycerol (HOG) and cell wall integrity (CWI) signalling pathways interplay: a yeast dialogue between MAPK routes. Yeast 27(8): 495-502. doi: 10.1002/yea.1792
- Deprez MA, Eskes E, Wilms T, Ludovico P, Winderickx J (2018). pH homeostasis links the nutrient sensing PKA/TORC1/Sch9 menage-a-trois to stress tolerance and longevity. Microbial Cell 5(3): 119-136. doi: 10.15698/mic2018.03.618
- Backhaus K, Rippert D, Heilmann CJ, Sorgo AG, de Koster CG, Klis FM, Rodicio R, Heinisch JJ (2013). Mutations in SNF1 complex genes affect yeast cell wall strength. Eur J Cell Biol 92(12): 383-395. doi: 10.1016/j.ejcb.2014.01.001
- Conrad M, Schothorst J, Kankipati HN, Van Zeebroeck G, Rubio-Texeira M, Thevelein JM (2014). Nutrient sensing and signaling in the yeast Saccharomyces cerevisiae. FEMS Microbiol Rev 38(2): 254-299. doi: 10.1111/1574-6976.12065
- Broach JR (2012). Nutritional control of growth and development in yeast. Genetics 192(1): 73-105. doi: 10.1534/genetics.111.135731
- Meyer CJ, Alenghat FJ, Rim P, Fong JH, Fabry B, Ingber DE (2000). Mechanical control of cyclic AMP signalling and gene transcription through integrins. Nat Cell Biol 2(9): 666-668. doi: 10.1038/35023621
- Alenghat FJ, Tytell JD, Thodeti CK, Derrien A, Ingber DE (2009). Mechanical control of cAMP signaling through integrins is mediated by the heterotrimeric Galphas protein. J Cell Biochem 106(4): 529-538. doi: 10.1002/jcb.22001
- Nakafuku M, Obara T, Kaibuchi K, Miyajima I, Miyajima A, Itoh H, Nakamura S, Arai K, Matsumoto K, Kaziro Y (1988). Isolation of a second yeast Saccharomyces cerevisiae gene (GPA2) coding for guanine nucleotide-binding regulatory protein: studies on its structure and possible functions. Proc Natl Acad Sci U S A 85(5): 1374-1378. doi: 10.1073/pnas.85.5.1374
- Zaman S, Lippman SI, Schneper L, Slonim N, Broach JR (2009). Glucose regulates transcription in yeast through a network of signaling pathways. Mol Syst Biol 5: 245. doi: 10.1038/msb.2009.2
- Petkova MI, Pujol-Carrion N, Arroyo J, Garcia-Cantalejo J, Angeles de la Torre-Ruiz M (2010). Mtl1 is required to activate general stress response through Tor1 and Ras2 inhibition under conditions of glucose starvation and oxidative stress. J Biol Chem 285(25): 19521-19531. doi: 10.1074/jbc.M109.085282
- Schmitz HP, Jendretzki A, Sterk C, Heinisch JJ (2018). The Small Yeast GTPase Rho5 and Its Dimeric GEF Dck1/Lmo1 Respond to Glucose Starvation. Int J Mol Sci 19(8). doi: 10.3390/ijms19082186
- Petkova MI, Pujol-Carrion N, de la Torre-Ruiz MA (2010). Signal flow between CWI/TOR and CWI/RAS in budding yeast under conditions of oxidative stress and glucose starvation. Commun Integr Biol 3(6): 555-557. doi: 10.4161/cib.3.6.12974
- McMurrough I, Rose AH (1967). Effect of growth rate and substrate limitation on the composition and structure of the cell wall of Saccharomyces cerevisiae. Biochem J 105(1): 189-203. doi: 10.1042/bj1050189
- Backhaus K, Heilmann CJ, Sorgo AG, Purschke G, de Koster CG, Klis FM, Heinisch JJ (2010). A systematic study of the cell wall composition of Kluyveromyces lactis. Yeast 27(8): 647-660. doi: 10.1002/yea.1781
- Merlini L, Dudin O, Martin SG (2013). Mate and fuse: how yeast cells do it. Open Biol 3(3): 130008. doi: 10.1098/rsob.130008
- Werner-Washburne M, Braun EL, Crawford ME, Peck VM (1996). Stationary phase in Saccharomyces cerevisiae. Mol Microbiol 19(6): 1159-1166. PMID: 8730858
- Cullen PJ, Sprague GF, Jr. (2000). Glucose depletion causes haploid invasive growth in yeast. Proc Natl Acad Sci U S A 97(25): 13619-13624. doi: 10.1073/pnas.240345197
- Pan X, Harashima T, Heitman J (2000). Signal transduction cascades regulating pseudohyphal differentiation of Saccharomyces cerevisiae. Curr Opin Microbiol 3(6): 567-572. doi: 10.1016/s1369-5274(00)00142-9
- Zaragoza O, Gancedo JM (2000). Pseudohyphal growth is induced in Saccharomyces cerevisiae by a combination of stress and cAMP signalling. Antonie van Leeuwenhoek 78(2): 187-194. PMID: 11204770
- Cullen PJ, Sprague GF, Jr. (2012). The regulation of filamentous growth in yeast. Genetics 190(1): 23-49. doi: 10.1534/genetics.111.127456
- Cutler NS, Pan X, Heitman J, Cardenas ME (2001). The TOR signal transduction cascade controls cellular differentiation in response to nutrients. Mol Biol Cell 12(12): 4103-4113. doi: 10.1091/mbc.12.12.4103
- Birkaya B, Maddi A, Joshi J, Free SJ, Cullen PJ (2009). Role of the cell wall integrity and filamentous growth mitogen-activated protein kinase pathways in cell wall remodeling during filamentous growth. Eukaryot Cell 8(8): 1118-1133. doi: 10.1128/EC.00006-09
- Mortimer RK (1958). Radiobiological and genetic studies on a polyploid series (haploid to hexaploid) of Saccharomyces cerevisiae. Radiat Res 9(3): 312-326. doi: 10.2307/3570795
- Soll DR, Herman MA (1983). Growth and the inducibility of mycelium formation in Candida albicans: a single-cell analysis using a perfusion chamber. J Gen Microbiol 129(9): 2809-2824. doi: 10.1099/00221287-129-9-2809
- Van Mulders SE, Stassen C, Daenen L, Devreese B, Siewers V, van Eijsden RG, Nielsen J, Delvaux FR, Willaert R (2011). The influence of microgravity on invasive growth in Saccharomyces cerevisiae. Astrobiology 11(1): 45-55. doi: 10.1089/ast.2010.0518
- Yan YX, Gong YW, Guo Y, Lv Q, Guo C, Zhuang Y, Zhang Y, Li R, Zhang XZ (2012). Mechanical strain regulates osteoblast proliferation through integrin-mediated ERK activation. Plos One 7(4): e35709. doi: 10.1371/journal.pone.0035709
- Chen Q, Kinch MS, Lin TH, Burridge K, Juliano RL (1994). Integrin-mediated cell adhesion activates mitogen-activated protein kinases. J Biol Chem 269(43): 26602-26605. PMID: 7929388
- Cook JG, Bardwell L, Thorner J (1997). Inhibitory and activating functions for MAPK Kss1 in the S. cerevisiae filamentous-growth signalling pathway. Nature 390(6655): 85-88. doi: 10.1038/36355
- Atienza JM, Suh M, Xenarios I, Landgraf R, Colicelli J (2000). Human ERK1 induces filamentous growth and cell wall remodeling pathways in Saccharomyces cerevisiae. J Biol Chem 275(27): 20638-20646. doi: 10.1074/jbc.M910024199
- Mosch HU, Roberts RL, Fink GR (1996). Ras2 signals via the Cdc42/Ste20/mitogen-activated protein kinase module to induce filamentous growth in Saccharomyces cerevisiae. Proc Natl Acad Sci U S A 93(11): 5352-5356. doi: 10.1073/pnas.93.11.5352
- Harashima T, Heitman J (2002). The Galpha protein Gpa2 controls yeast differentiation by interacting with kelch repeat proteins that mimic Gbeta subunits. Mol Cell 10(1): 163-173. doi: 10.1016/s1097-2765(02)00569-5
- Shively CA, Eckwahl MJ, Dobry CJ, Mellacheruvu D, Nesvizhskii A, Kumar A (2013). Genetic networks inducing invasive growth in Saccharomyces cerevisiae identified through systematic genome-wide overexpression. Genetics 193(4): 1297-1310. doi: 10.1534/genetics.112.147876
- Jendretzki A, Wittland J, Wilk S, Straede A, Heinisch JJ (2011). How do I begin? Sensing extracellular stress to maintain yeast cell wall integrity. Eur J Cell Biol 90(9): 740-744. doi: 10.1016/j.ejcb.2011.04.006
- Rodicio R, Buchwald U, Schmitz HP, Heinisch JJ (2008). Dissecting sensor functions in cell wall integrity signaling in Kluyveromyces lactis. Fungal Genet Biol 45(4): 422-435. doi: 10.1016/j.fgb.2007.07.009
- Negishi T, Ohya Y (2010). The cell wall integrity checkpoint: coordination between cell wall synthesis and the cell cycle. Yeast 27(8): 513-519. doi: 10.1002/yea.1795
- Gancedo JM (2001). Control of pseudohyphae formation in Saccharomyces cerevisiae. FEMS Microbiol Rev 25(1): 107-123. doi: 10.1016/s0168-6445(00)00056-5
- Suzuki M, Igarashi R, Sekiya M, Utsugi T, Morishita S, Yukawa M, Ohya Y (2004). Dynactin is involved in a checkpoint to monitor cell wall synthesis in Saccharomyces cerevisiae. Nat Cell Biol 6(9): 861-871. doi: 10.1038/ncb1162
- Cordes N, van Beuningen D (2003). Cell adhesion to the extracellular matrix protein fibronectin modulates radiation-dependent G2 phase arrest involving integrin-linked kinase (ILK) and glycogen synthase kinase-3beta (GSK-3beta) in vitro. Br J Cancer 88(9): 1470-1479. doi: 10.1038/sj.bjc.6600912
ACKNOWLEDGMENTS
Funding for this project was provided to AB by the Swedish Research Council (VR, #2017-04559). We would also like to thank two anonymous reviewers for their insightful comments that certainly improved our manuscript.
COPYRIGHT
© 2019

Integrins in disguise – mechanosensors in Saccharomyces cerevisiae as functional integrin analogues by Elhasi and Blomberg is licensed under a Creative Commons Attribution 4.0 International License.