
Regulation of extracellular vesicles for protein secretion in Aspergillus nidulans
This study reveals that Aspergillus nidulans boosts extracellular vesicle production when ER-trafficked enzymes are induced, uncovering how fungi remodel their secretome through vesicle-mediated secretion to adapt to changing environments and biofilm formation.

Transcriptomic response to different heme sources in Trypanosoma cruzi epimastigotes
This study uncovers how the Chagas disease parasite adapts to changes in heme, an essential molecule for its survival, providing transcriptional clues to heme metabolism and identifying a previously unreported heme-binding protein in T. cruzi.

Luminal acetylation of microtubules is not essential for Plasmodium berghei and Toxoplasma gondii survival
Acetylation of α-tubulin at lysine 40 is not essential for cytoskeletal stability in Plasmodium berghei or Toxoplasma gondii, suggesting redundancy and plasticity in microtubule regulation in these parasites.

The dual-site agonist for human M2 muscarinic receptors Iper-8-naphtalimide induces mitochondrial dysfunction in Saccharomyces cerevisiae
S. cerevisiae is a model to study human GPCRs. N-8-Iper, active against glioblastoma via M2 receptor, causes mitochondrial damage in yeast by binding Ste2, highlighting evolutionary conservation of GPCRs.

Integrative Omics reveals changes in the cellular landscape of peroxisome-deficient pex3 yeast cells
To uncover the consequences of peroxisome deficiency, we compared Saccharomyces cerevisiae wild-type with pex3 cells, which lack peroxisomes, employing quantitative proteomics and transcriptomics technologies.

Regulation of extracellular vesicles for protein secretion in Aspergillus nidulans
Rebekkah E. Pope1, Patrick Ballmann2, Lisa Whitworth3 and Rolf A. Prade1,*
This study reveals that Aspergillus nidulans boosts extracellular vesicle production when ER-trafficked enzymes are induced, uncovering how fungi remodel their secretome through vesicle-mediated secretion to adapt to changing environments and biofilm formation.

Transcriptomic response to different heme sources in Trypanosoma cruzi epimastigotes
Evelyn Tevere1,a, María G. Mediavilla1,a, Cecilia B. Di Capua1, Marcelo L. Merli1, Carlos Robello2,3, Luisa Berná2,4 and Julia A. Cricco
This study uncovers how the Chagas disease parasite adapts to changes in heme, an essential molecule for its survival, providing transcriptional clues to heme metabolism and identifying a previously unreported heme-binding protein in T. cruzi.
Sir2 regulates selective autophagy in stationary-phase yeast cells
Ji-In Ryua, Juhye Junga, and Jeong-Yoon Kim
This study establishes Sir2 as a previously unrecognized regulator of selective autophagy during the stationary phase and highlight how cells dynamically control organelle degradation.

Unresolved mystery of cyclic nucleotide second messengers, periplasmic acid phosphatases and bacterial natural competence
Kristina Kronborg and Yong Everett Zhang
In this study we aimed to identify the promotors responsible for the expression of the non-specific acid phosphatase AphA during different starvation conditions, to confirm the requirement of the cAMP-dependent CRP regulon for aphA expression, and to finally identify regulators of its expression.

Polyadenylated versions of small non-coding RNAs in Saccharomyces cerevisiae are degraded by Rrp6p/Rrp47p independent of the core nuclear exosome
Anusha Chaudhuri1,#, Soumita Paul2,#, Mayukh Banerjea2 and Biswadip Das2
In this investigation, we unveiled a novel functional role of the major nuclear 3′→5′ exoribonuclease, Rrp6p, and its cofactor Rrp47p in the degradation of polyadenylated versions of several mature sncRNAs, including 5S, 5.8S rRNAs, all sn- and some select snoRNAs in the baker’s yeast S. cerevisiae.
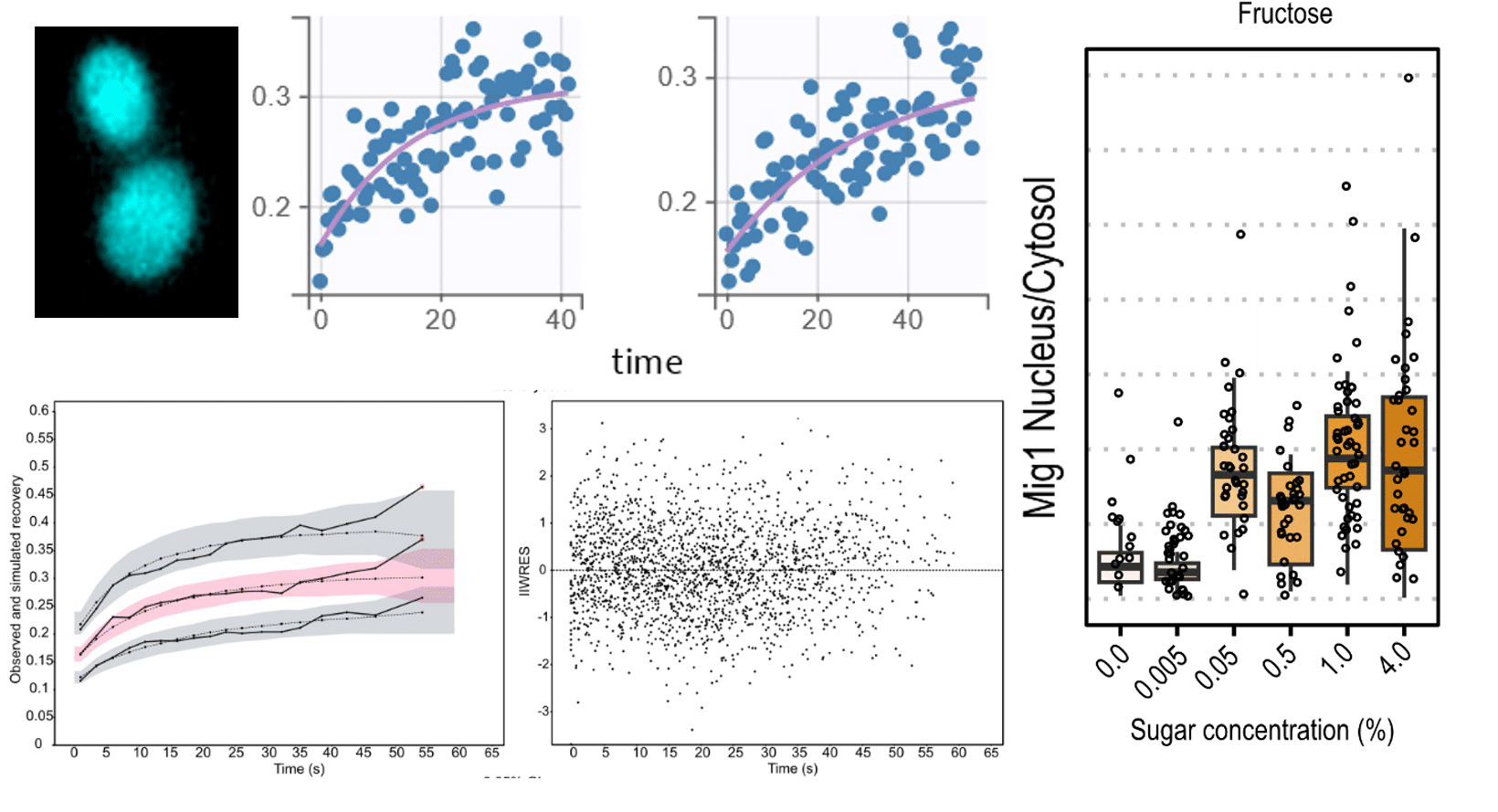
Exploring carbon source related localization and phosphorylation in the Snf1/Mig1 network using population and single cell-based approaches
Svenja Braam1, Farida Tripodi2, Linnea Österberg1,3, Sebastian Persson1, Niek Welkenhuysen1, Paola Coccetti2 and Marija Cvijovic1
In this work we set out to explore the relationship between the subcellular localization and regulation of kinases in the context of carbon source signaling. The data presented in this paper reinforce the notion that not only the activation/inactivation of kinases but also their subcellular localization and that of their targets influence fate decisions in response to environmental changes.

A Modular Cloning Toolkit for the production of recombinant proteins in Leishmania tarentolae
Katrin Hieronimus1,2,#, Tabea Donauer1,2,#, Jonas Klein1,#, Bastian Hinkel1,#, Julia Vanessa Spänle1,#, Anna Probst1,#, Justus Niemeyer1,#, Salina Kibrom1, Anna Maria Kiefer1, Luzia Schneider2, Britta Husemann2, Eileen Bischoff2, Sophie Möhring2, Nicolas Bayer1, Dorothée Klein1, Adrian Engels1, Benjamin Gustav Ziehmer2, Julian Stieß3, Pavlo Moroka1, Michael Schroda1, and Marcel Deponte2
Modular Cloning (MoClo) is based on libraries of standardized genetic parts that can be directionally assembled via Golden Gate cloning in one-pot reactions into transcription units and multigene constructs. We established a MoClo toolkit and exemplified its application for the production of recombinant proteins in L. tarentolae.
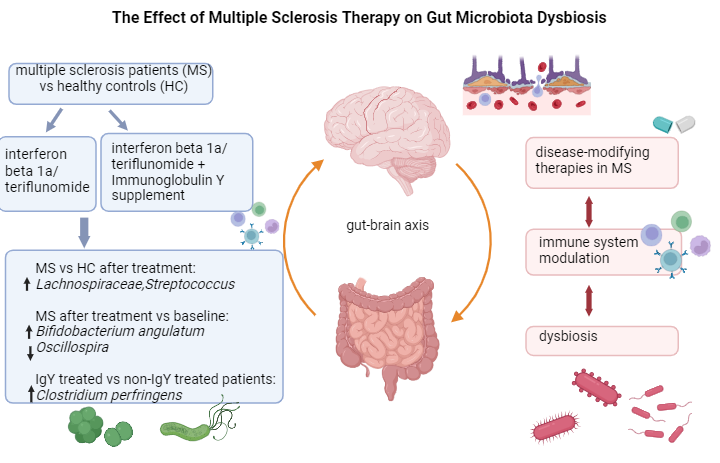
The effect of multiple sclerosis therapy on gut microbiota dysbiosis: a longitudinal prospective study
Andreea-Cristina Paraschiv1,a, Vitalie Vacaras1,2,a, Cristina Nistor1,2, Cristiana Vacaras3, Stefan Strilciuc1 and Dafin F Muresanu1,2
The gut microbiota, a complex ecosystem with various immune functions, plays a significant role in MS, and its response to different treatments is highlighted in this study. In clinical practice, maintaining a healthy microbiota is crucial for individuals with MS.
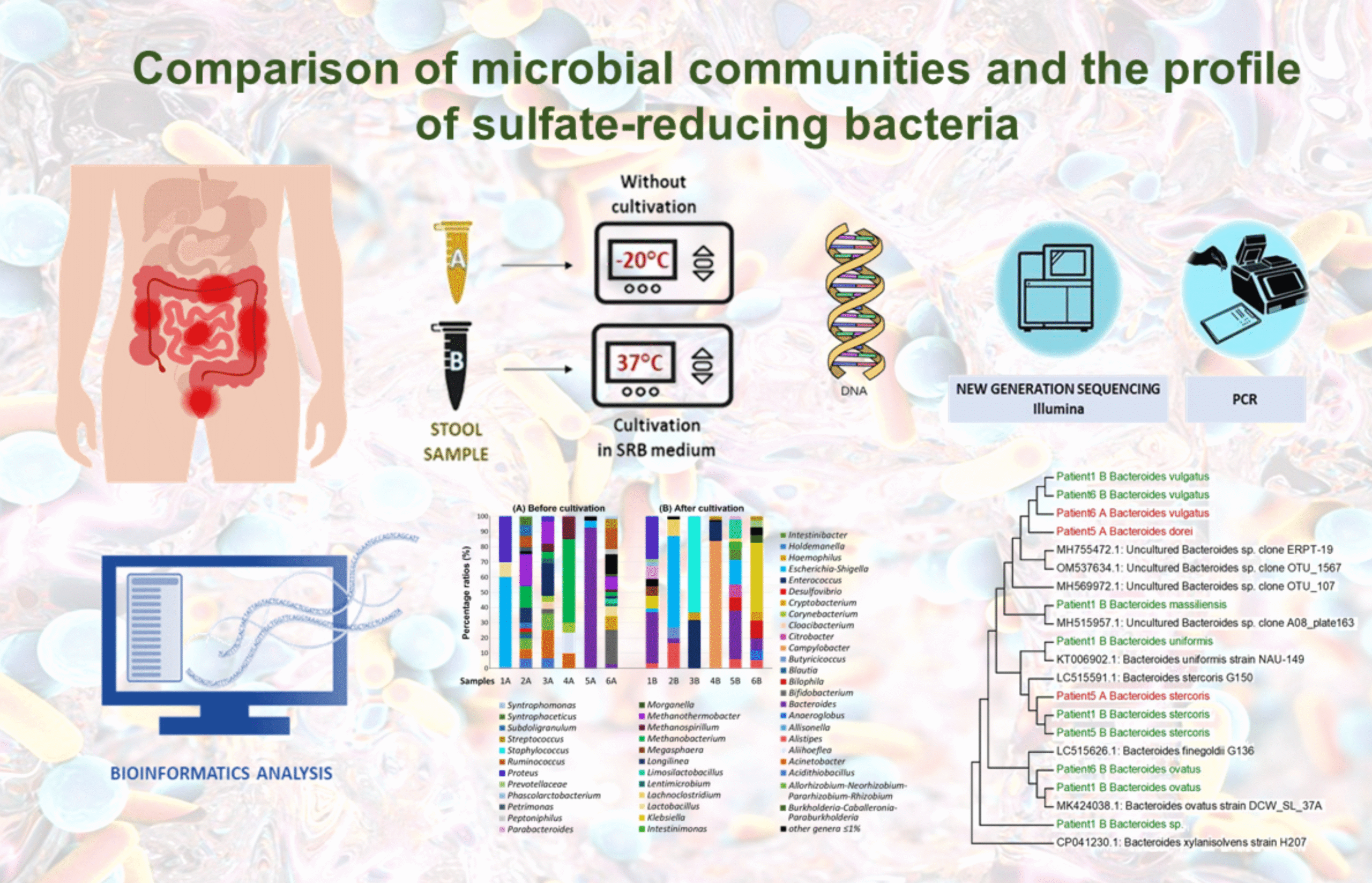
Comparison of microbial communities and the profile of sulfate-reducing bacteria in patients with ulcerative colitis and their association with bowel diseases: a pilot study
Ivan Kushkevych1, Kristýna Martínková1, Lenka Mráková1, Francesco Giudici2, Simone Baldi2, David Novak3, Márió Gajdács4, Monika Vítězová1, Dani Dordevic5, Amedeo Amedei2 and Simon K.-M. R. Rittmann6
Considerable evidence has accumulated regarding the molecular relationship between gut microbiota (GM) composition and the onset (clinical presentation and prognosis) of ulcerative colitis UC. Our findings highlight, among other observations, significant variations in the gut microbial composition among patients with varying disease severity and activity.

Replicative aging in yeast involves dynamic intron retention patterns associated with mRNA processing/export and protein ubiquitination
Jesús Gómez-Montalvo1, Alvaro de Obeso Fernández del Valle1, Luis Fernando De la Cruz Gutiérrez1, Jose Mario Gonzalez-Meljem1 and Christian Quintus Scheckhuber1
Saccharomyces cerevisiae has yielded relevant insights into some of the basic mechanisms of organismal aging. Among these are genomic instability, oxidative stress, caloric restriction and mitochondrial dysfunction. Our work uncovers a previously unexplored layer of the transcriptional program of yeast aging and, more generally, expands the knowledge on the occurrence of alternative splicing in baker´s yeast.
A pseudokinase couples signaling pathways to enable asymmetric cell division in a bacterium
W. Seth Childers and Lucy Shapiro
In this article, the authors comment on the study “Cell fate regulation governed by a repurposed bacterial histidine kinase” by Childers et al., PLoS Biol. 2014 Oct 28;12(10):e1001979.
Targeting of chromatin readers: a novel strategy used by the Shigella flexneri virulence effector OspF to reprogram transcription
Habiba Harouz, Christophe Rachez, Benoit Meijer, Christian Muchardt, Laurence Arbibe.
In this microreview, the authors discuss the article “Shigella flexneri targets the HP1γ subcode through the phosphothreoninelyase OspF” by Harouz et al. (2014), EMBO J, 22 : 2606-2622.
Plasmodium spp. membrane glutathione S-transferases: detoxification units and drug targets
Andreas Martin Lisewski
This article comments on work published by Lisewski et al. (Cell, 2014), which reported the first examples of membrane-associated proteins in eicosanoid and glutathione metabolism members among Plasmodium spp.
Proline cis-trans isomerization is influenced by local lysine acetylation-deacetylation
Françoise S. Howe and Jane Mellor
This article comments on work published by Howe et al. (Mol Cell, 2014), which shows that local lysine acetylation and deacetylation modulate proline cis-trans isomerization in Saccharomyces cerevisiae.
On the link between cell cycle and infection of the Alphaproteobacterium Brucella abortus
Michaël Deghelt, Jean-Jacques Letesson, Xavier De Bolle
This article comments on work published by Deghelt et al. (Nat Comm, 2014), which describe a cell cycle arrest and resume during the Brucella abortus trafficking in host cell, suggesting that like the model Alphaproteobacterium Caulobacter crescentus, these bacteria are able to block their cell cycle at the G1 phase when starvation is sensed.
Divide and conquer: processive transport enables multidrug transporters to tackle challenging drugs
Nir Fluman and Eitan Bibi
This article comments on work published by Fluman et al. (Nat Comm, 2014), which describes the ability of bacterial multidrug transporters to move long molecules through the membrane in a processive manner.
The dual role of cyclin C connects stress regulated gene expression to mitochondrial dynamics
Randy Strich and Katrina F. Cooper
This work summarizes the role cyclin C plays in regulating stress-responsive transcription in the budding yeast Saccharomyces cerevisiae, including mitochondrial fission and regulated cell death.
Combinatorial stress responses: direct coupling of two major stress responses in Escherichia coli
Daniel R. Brown, Geraint Barton, Zhensheng Pan, Martin Buck and Sivaramesh Wigneshweraraj
This article comments on work published by Brown et al. (Nat Comm, 2014), which showed that the transcription of relA is activated by NtrC during nitrogen starvation, revealing that in E. coli and related bacteria, NtrC functions in combinatorial stress and serves to couple two major stress responses, the Ntr response and stringent response.
The replication timing program in the hands of two HDACs
Kazumasa Yoshida1,2, Armelle Lengronne1 and Philippe Pasero1
This article comments on work published by Yoshida et al. (Mol Cell, 2014), which performed a systematic analysis of the role of histone deacetylases (HDACs) in the regulation of origin activity in budding yeast, finding that the epigenetic regulation of repetitive sequences is a key determinant of the DNA replication program.
Means of intracellular communication: touching, kissing, fusing
Anne Spang1
This work highlights different aspects of communication between organelles, including the importance of organellar contact sites.
Neuropathogenesis caused by Trypanosoma brucei, still an enigma to be unveiled
Katherine Figarella1
This Editorial addresses the meningo-encephalitic stage of Trypanosoma brucei infection and the resultig neuropathogenesis as well as the impact that the application of tools developed in the last years in the field of neuroscience will have on the study of neglected tropical diseases.
Lichens – growing greenhouses en miniature
Martin Grube1
This commentary article provides an overview on different aspects of lichen biology and the remarkable symbiotic association between fungi and algae.
Regulation of the mitochondrial permeability transition pore and its effects on aging
Damiano Pellegrino-Coppola1
Aging is linked to mitochondrial function, with the mitochondrial permeability transition pore (mPTP) playing a key role. Yeast is a useful model for studying how mPTP affects cell survival, aging, and related diseases.
Fungal infections in humans: the silent crisis
Katharina Kainz1, Maria A. Bauer1, Frank Madeo1-3 and Didac Carmona-Gutierrez1
This article highlights the growing global threat of fungal infections – exacerbated by rising drug resistance and medical practices – and emphasizes the urgent need for intensified research to develop more effective antifungal strategies.
Digesting the crisis: autophagy and coronaviruses
Didac Carmona-Gutierrez1, Maria A. Bauer1, Andreas Zimmermann1,2, Katharina Kainz1,
Sebastian J. Hofer1, Guido Kroemer3-7 and Frank Madeo1,2,8
This article reviews the multifaceted role of autophagy in antiviral defense and highlights how coronaviruses, including SARS-CoV-2, interact with this pathway, raising the possibility that targeting autophagy could offer novel therapeutic strategies against COVID-19.
Microbial Cell
is an open-access, peer-reviewed journal that publishes exceptionally relevant research works that implement the use of unicellular organisms (and multicellular microorganisms) to understand cellular responses to internal and external stimuli and/or human diseases.
you can trust
Can’t find what you’re looking for?
You can browse all our issues and published articles here.
FAQs
Peer-reviewed, open-access research using unicellular organisms (and multicellular microorganisms) to understand cellular responses and human disease.
The journal (founded in 2014) is led by its Editors-in-Chief Frank Madeo, Didac Carmona-Gutierrez, and Guido Kroemer
Microbial Cell has been publishing original scientific literature since 2014, and from the very beginning has been managed by active scientists through an independent Publishing House (Shared science Publishers). The journal was conceived as a platform to acknowledge the importance of unicellular organisms, both as model systems as well as in the biological context of human health and disease.
Ever since, Microbial Cell has very positively developed and strongly grown into a respected journal in the unicellular research community and even beyond. This scientific impact is reflected in the yearly number of citations obtained by articles published in Microbial Cell, as recorded by the Web of Science (Clarivate, formerly Thomson/Reuters):

The scientific impact of Microbial Cell is also mirrored in a series of milestones:
2015: Microbial Cell is included in the Emerging Sources Citation Index (ESCI), a selection of developing journals drafted by Clarivate Analytics based on the candidate’s publishing standards, quality, editorial content, and citation data. Note: As an ESCI-selected journal, Microbial Cell is currently being evaluated in a rigorous and long process to determine an inclusion in the Science Citation Index Expanded (SCIE), which allows the official calculation of Clarivate Analytics’ impact factor.
2016: Microbial Cell is awarded the so-called DOAJ Seal by the selective Directory of Open Access Journals (DOAJ). The DOAJ Seal is an exclusive mark of certification for open access journals granted by DOAJ to journals that adhere to outstanding best practice and achieve an extra high and clear commitment to open access and high publishing standards.
2017: Microbial Cell is included in Pubmed Central (PMC), allowing the archiving of all the journal’s articles in PMC and PubMed.
2019: Microbial Cell is indexed in the prestigious abstract and citation database Scopus after a thorough selection process. This also means that Microbial Cell obtains, for the first time, an official Scopus CiteScore as well as an official journal ranking in the Scimago Journal and Country Ranking.
2022: Microbial Cell’s CiteScore reaches a value of 7.2 for the year 2021, positioning Microbial Cell among the top microbiology journals (previously available CiteScores: 2019: 5.4; 2020: 5.1).
2022: Microbial Cell is indexed in the highly selective Science Citation Index Expanded™, which covers approx. 9,500 of the world’s most impactful journals across 178 scientific disciplines. In their journal selection and curation process, Clarivate´s editors apply 24 ‘quality’ criteria and four ‘impact’ criteria to select the most influential journals in their respective fields. This selection is also a pre-requisite for inclusion in the JCR, which features the impact factor.
2022: Microbial Cell is listed in the Journal Citation Reports™ (JCR), and obtains its first official Journal Impact Factor™ (JIF) for the year 2021: 5.316.Check Article Types and Manuscript Preparation guidelines. Submit online via Scholastica.



The long and winding road of reverse genetics in Trypanosoma cruzi
Miguel A. Chiurillo1 and Noelia Lander1
This Editorial provides a brief historic overview that highlights the strengths and weaknesses of the molecular strategies that have been developed to genetically modify Trypanosoma cruzi, emphasizing the future directions of the field.