
Regulation of extracellular vesicles for protein secretion in Aspergillus nidulans
This study reveals that Aspergillus nidulans boosts extracellular vesicle production when ER-trafficked enzymes are induced, uncovering how fungi remodel their secretome through vesicle-mediated secretion to adapt to changing environments and biofilm formation.

Transcriptomic response to different heme sources in Trypanosoma cruzi epimastigotes
This study uncovers how the Chagas disease parasite adapts to changes in heme, an essential molecule for its survival, providing transcriptional clues to heme metabolism and identifying a previously unreported heme-binding protein in T. cruzi.

Luminal acetylation of microtubules is not essential for Plasmodium berghei and Toxoplasma gondii survival
Acetylation of α-tubulin at lysine 40 is not essential for cytoskeletal stability in Plasmodium berghei or Toxoplasma gondii, suggesting redundancy and plasticity in microtubule regulation in these parasites.

The dual-site agonist for human M2 muscarinic receptors Iper-8-naphtalimide induces mitochondrial dysfunction in Saccharomyces cerevisiae
S. cerevisiae is a model to study human GPCRs. N-8-Iper, active against glioblastoma via M2 receptor, causes mitochondrial damage in yeast by binding Ste2, highlighting evolutionary conservation of GPCRs.

Integrative Omics reveals changes in the cellular landscape of peroxisome-deficient pex3 yeast cells
To uncover the consequences of peroxisome deficiency, we compared Saccharomyces cerevisiae wild-type with pex3 cells, which lack peroxisomes, employing quantitative proteomics and transcriptomics technologies.

The mechanism of Tat-dependent protein translocation
Brüser and SandersThis review integrates mechanistically relevant biochemical, molecular, and structural studies on Tat-dependent translocation of folded proteins into an in its molecular detail new comprehensive explanation of how the Tat system mediates protein transport.

TOR-dependent regulation of the yeast homolog of the juvenile Batten Disease-associated gene CLN3
Pillalamarri et al.This study identifies conditions and genes that induce BTN1 expression in yeast. We show that BTN1 expression is regulated by translational control and by the mTOR1 pathway. An understanding of when and why BTN1 expression will aid in understanding the expression of CLN3, which may be helpful in the treatment of this devastating disease.

Overcoming phagocytosis resistance of hypervirulent Klebsiella pneumoniae by directly targeting capsules
Tsubaki et al.This study highlights a promising strategy for disarming hypervirulent K. pneumoniae by directly targeting its key virulence factors and provides novel insights into antibacterial therapeutic approaches against this clinically significant pathogen.
Protein arginine methyltransferases in protozoan parasites: a new path for antiparasitic chemotherapy?
Campagnaro et al.This review discusses the activity and the relevance of arginine methyltransferases for the survival of pathogenic kinetoplastids, apicomplexans and amoebas, and how these enzymes could be exploited as drug targets.

VapA/Scs2 sustains polarized growth in Aspergillus nidulans by maintaining AP-2-mediated apical endocytosis
Georgiou et al.To explore the functional significance of ER–PM contact sites in filamentous fungi, we identified and genetically characterized all Aspergillus nidulans proteins homologous to Snc2/VAP, Ist2, or tricalbins.

Genetic make-up and regulation of the L-lysine biosynthesis pathway in Vibrio natriegens
Straube et al.This study analysed the make-up and regulation of the biosynthetic pathway for L-lysine and related L-aspartate family amino acids (AFAAs) in Vibrio natriegens DSM759 to provide a comprehensive basis for future metabolic engineering endeavours aiming at developing this strain into an amino acid overproducer.
Airborne bacteria in show caves from Southern Spain
This study analyzes the factors conditioning the diversity of airborne bacteria recorded in three Andalusian show caves, subjected to different managements.
Landscapes and bacterial signatures of mucosa-associated intestinal microbiota in Chilean and Spanish patients with inflammatory bowel disease
This study investigates the landscapes and alterations of mucosa-associated intestinal microbiota in patients with inflammatory bowel diseases, which cause chronic inflammation of the gut, including ulcerative colitis and Crohn’s disease.
Genome, transcriptome and secretome analyses of the antagonistic, yeast-like fungus Aureobasidium pullulans to identify potential biocontrol genes
This study highlights the value of a sequential approach starting with genome mining and consecutive transcriptome and secretome analyses in order to identify a limited number of potential target genes for detailed, functional analyses in Aureobasidium pullulans.
Proanthocyanidin-enriched cranberry extract induces resilient bacterial community dynamics in a gnotobiotic mouse model
This study investigates the effect of a water-soluble, proanthocyanidin-rich cranberry juice extract on the short-term dynamics of a human-derived bacterial community in a gnotobiotic mouse model.
Dry biocleaning of artwork: an innovative methodology for Cultural Heritage recovery?
This work proposes an innovative methodology based on applied biotechnology for the recovery of altered stonework: the “dry biocleaning”, which envisages the use of dehydrated microbial cells without the use of free water or gel-based matrices.
Aeration mitigates endoplasmic reticulum stress in Saccharomyces cerevisiae even without mitochondrial respiration
This work demonstrates a scenario, in which aeration acts beneficially on Saccharmyces cerevisiae cells even under fermentative conditions.
A novel BR-SMAD is required for larval development in barber’s pole worm Haemonchus contortus
The herein presented results show a BMP-like receptor-regulated SMAD in Haemonchus contortus that is required for larval differentiation and underscore an adaptive functional repurposing of BMP-signaling in parasitic worms.
Nutrient sensing and cAMP signaling in yeast: G-protein coupled receptor versus transceptor activation of PKA
The herein presented work supports a model, in which nutrient transceptors are evolutionary ancestors of GPCRs, employing a more primitive direct signaling mechanism compared to the indirect cAMP second-messenger signaling mechanism used by GPCRs for activation of PKA.
Longevity pathways and maintenance of the proteome: the role of autophagy and mitophagy during yeast ageing
Belém Sampaio-Marques et al.This review describes recent findings that shed light on how longevity pathways and metabolic status impact maintenance of the proteome in both yeast ageing paradigms. These findings demonstrate that yeast remain a powerful model system for elucidating these relationships and their influence on ageing regulation.
Secondary structures involving the poly(A) tail and other 3’ sequences are major determinants of mRNA isoform stability in yeast
Zarmik Moqtaderi at al.This article comments on work published by Geisberg et al. (Cell (2014), which points to an important role for mRNA structure at 3’ termini in governing transcript stability, likely by reducing the interaction of the mRNA with the degradation apparatus.
De novo peroxisome biogenesis revisited
Marten Veenhuis and Ida J. van der KleiThis article comments on work published by Knoops et al. (JCB, 2014), which describes an alternative peroxisome formation pathway in yeast pex3 and pex19 cells, which relies on the existence of small peroxisomal remnants that are present in these cells.
Transcriptional and genomic mayhem due to aging-induced nucleosome loss in budding yeast
Zheng Hu et al.This article comments on work published by Zheng et al. (Genes and Development, 2014), which investigated a loss of histones during replicative aging in budding yeast, which was also accompanied by a significantly-increased frequency of genomic instability including DNA breaks, chromosomal translocations, retrotransposition, and transfer of mitochondrial DNA into the nuclear genome.
The Parkinson’s disease-associated protein α-synuclein disrupts stress signaling – a possible implication for methamphetamine use?
Shaoxiao Wang and Stephan N. WittThis article comments on work published by Wang et al. (PNAS, 2012), which reported that human α-syn, at high expression levels, disrupts stress-activated signal transduction pathways in both yeast and human neuroblastoma cells. Disruption of these signaling pathways ultimately leads to vulnerability to stress and to cell death.
Massive gene swamping among cheese-making Penicillium fungi
Jeanne Ropars et al.This article comments on work published by Cheeseman et al. (Nat Comm, 2014), which indicates that horizontal gene transfer is a crucial mechanism of rapid adaptation, even among eukaryotes.
Genome-wide studies of telomere biology in budding yeast
Harari and KupiecIn the last decade, technical advances have allowed carrying out systematic genome-wide screens for mutants affecting various aspects of telomere biology. In this review we summarize these efforts, and the insights that this Systems Biology approach has produced so far.
Mnemons: encoding memory by protein super-assembly
Fabrice Caudron and Yves BarralThis article comments on work published by Caudron and Barral (Cell, 2013), which proposes that polyQ- and polyN-based elements, termed mnemons, act as cellular memory devices to encode previous environmental conditions.
Transceptors as a functional link of transporters and receptors
George DiallinasA relative newcomer in environment sensing are the so called transceptors, membrane proteins that possess both solute transport and receptor-like signaling activities. Now, the transceptor concept is further enlarged to include micronutrient sensing via the iron and zinc high-affinity transporters of Saccharomyces cerevisiae.
S. pombe placed on the prion map
Jacqueline HaylesThis article comments on work published by Sideri et al. (Microbial Cell, 2017), which identified the Ctr4 prion in S. pombe.
Using microbes as a key tool to unravel the mechanism of autophagy and the functions of the ATG proteins
Mario Mauthe and Fulvio ReggioriMicrobes have served to discover and characterize unconventional functions of the ATG proteins, which are uncoupled from their role in autophagy. In our recent study, we have taken advantage of viruses as a screening tool to determine the extent of the unconventional functions of the ATG proteome and characterize one of them.
Autophagy: one more Nobel Prize for yeast
Zimmermann et al.The recent announcement of the 2016 Nobel Prize in Physiology or Medicine, awarded to Yoshinori Ohsumifor the discoveries of mechanisms governing autophagy, underscores the importance of intracellular degradation and recycling. Here we provide a quick historical overview that mirrors both the importance of autophagy as a conserved and essential process for cellular life and death as well as the crucial role of yeast in its mechanistic characterization.
Physiology, phylogeny, and LUCA
Martin et al.Genomes record their own history. But if we want to look all the way back to life’s beginnings some 4 billion years ago, the record of microbial evolution that is preserved in prokaryotic genomes is not easy to read. The classical approach has been to look for genes that are universally distributed. Another approach is to make all trees for all genes, and sift out the trees where signals have been overwritten by lateral gene transfer. What is left ought to be ancient. If we do that, what do we find?
Sexually transmitted infections: old foes on the rise
Carmona-Gutierrez et al.Sexually transmitted infections (STIs) are commonly spread via sexual contact. It is estimated that one million STIs are acquired every day worldwide. Besides their impact on sexual, reproductive and neonatal health, they can cause disastrous and life-threatening complications if left untreated. In addition to this personal burden, STIs also represent a socioeconomic problem, deriving in treatment costs of tremendous proportions. Despite a substantial progress in diagnosis, treatment and prevention, the incidence of many common STIs is increasing, and STIs continue to represent a global public health problem and a major cause for morbidity and mortality. With this Special Issue, Microbial Cell provides an in-depth overview of the eight major STIs, covering all relevant features of each infection.
Can’t find what you’re looking for?
You can browse all our issues and published articles here.
FAQs
Peer-reviewed, open-access research using unicellular organisms (and multicellular microorganisms) to understand cellular responses and human disease.
The journal (founded in 2014) is led by its Editors-in-Chief Frank Madeo, Didac Carmona-Gutierrez, and Guido Kroemer
Microbial Cell has been publishing original scientific literature since 2014, and from the very beginning has been managed by active scientists through an independent Publishing House (Shared science Publishers). The journal was conceived as a platform to acknowledge the importance of unicellular organisms, both as model systems as well as in the biological context of human health and disease.
Ever since, Microbial Cell has very positively developed and strongly grown into a respected journal in the unicellular research community and even beyond. This scientific impact is reflected in the yearly number of citations obtained by articles published in Microbial Cell, as recorded by the Web of Science (Clarivate, formerly Thomson/Reuters):
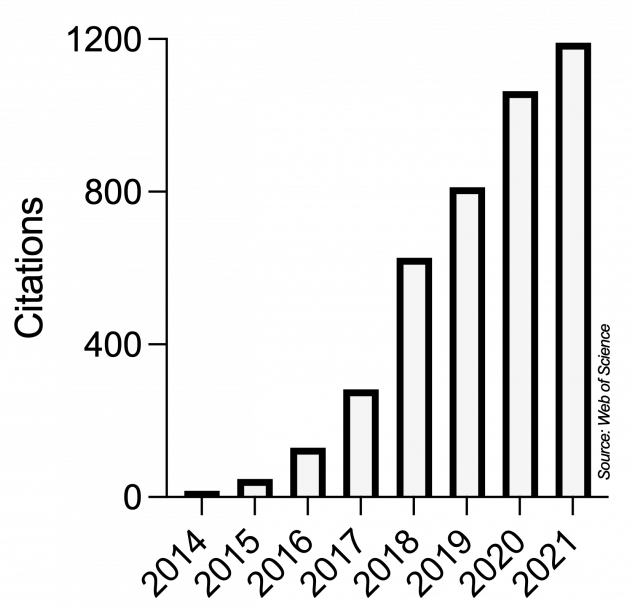
The scientific impact of Microbial Cell is also mirrored in a series of milestones:
2015: Microbial Cell is included in the Emerging Sources Citation Index (ESCI), a selection of developing journals drafted by Clarivate Analytics based on the candidate’s publishing standards, quality, editorial content, and citation data. Note: As an ESCI-selected journal, Microbial Cell is currently being evaluated in a rigorous and long process to determine an inclusion in the Science Citation Index Expanded (SCIE), which allows the official calculation of Clarivate Analytics’ impact factor.
2016: Microbial Cell is awarded the so-called DOAJ Seal by the selective Directory of Open Access Journals (DOAJ). The DOAJ Seal is an exclusive mark of certification for open access journals granted by DOAJ to journals that adhere to outstanding best practice and achieve an extra high and clear commitment to open access and high publishing standards.
2017: Microbial Cell is included in Pubmed Central (PMC), allowing the archiving of all the journal’s articles in PMC and PubMed.
2019: Microbial Cell is indexed in the prestigious abstract and citation database Scopus after a thorough selection process. This also means that Microbial Cell obtains, for the first time, an official Scopus CiteScore as well as an official journal ranking in the Scimago Journal and Country Ranking.
2022: Microbial Cell’s CiteScore reaches a value of 7.2 for the year 2021, positioning Microbial Cell among the top microbiology journals (previously available CiteScores: 2019: 5.4; 2020: 5.1).
2022: Microbial Cell is indexed in the highly selective Science Citation Index Expanded™, which covers approx. 9,500 of the world’s most impactful journals across 178 scientific disciplines. In their journal selection and curation process, Clarivate´s editors apply 24 ‘quality’ criteria and four ‘impact’ criteria to select the most influential journals in their respective fields. This selection is also a pre-requisite for inclusion in the JCR, which features the impact factor.
2022: Microbial Cell is listed in the Journal Citation Reports™ (JCR), and obtains its first official Journal Impact Factor™ (JIF) for the year 2021: 5.316.Check Article Types and Manuscript Preparation guidelines. Submit online via Scholastica.



Staphylococcus aureus type I signal peptidase: essential or not essential, that’s the question
Wouter L.W. Hazenbos et al.This article comments on work published by Morisaki et al. (mBio, 2016), which characterized a novel ABC transporter. This transporter apparently compensates for SpsB’s essential function by mediating alternative cleavage of a subset of proteins at a site distinct from the SpsB-cleavage site, leading to SpsB-independent secretion.