
Regulation of extracellular vesicles for protein secretion in Aspergillus nidulans
This study reveals that Aspergillus nidulans boosts extracellular vesicle production when ER-trafficked enzymes are induced, uncovering how fungi remodel their secretome through vesicle-mediated secretion to adapt to changing environments and biofilm formation.

Transcriptomic response to different heme sources in Trypanosoma cruzi epimastigotes
This study uncovers how the Chagas disease parasite adapts to changes in heme, an essential molecule for its survival, providing transcriptional clues to heme metabolism and identifying a previously unreported heme-binding protein in T. cruzi.

Luminal acetylation of microtubules is not essential for Plasmodium berghei and Toxoplasma gondii survival
Acetylation of α-tubulin at lysine 40 is not essential for cytoskeletal stability in Plasmodium berghei or Toxoplasma gondii, suggesting redundancy and plasticity in microtubule regulation in these parasites.

The dual-site agonist for human M2 muscarinic receptors Iper-8-naphtalimide induces mitochondrial dysfunction in Saccharomyces cerevisiae
S. cerevisiae is a model to study human GPCRs. N-8-Iper, active against glioblastoma via M2 receptor, causes mitochondrial damage in yeast by binding Ste2, highlighting evolutionary conservation of GPCRs.

Integrative Omics reveals changes in the cellular landscape of peroxisome-deficient pex3 yeast cells
To uncover the consequences of peroxisome deficiency, we compared Saccharomyces cerevisiae wild-type with pex3 cells, which lack peroxisomes, employing quantitative proteomics and transcriptomics technologies.

The mechanism of Tat-dependent protein translocation
Brüser and SandersThis review integrates mechanistically relevant biochemical, molecular, and structural studies on Tat-dependent translocation of folded proteins into an in its molecular detail new comprehensive explanation of how the Tat system mediates protein transport.

TOR-dependent regulation of the yeast homolog of the juvenile Batten Disease-associated gene CLN3
Pillalamarri et al.This study identifies conditions and genes that induce BTN1 expression in yeast. We show that BTN1 expression is regulated by translational control and by the mTOR1 pathway. An understanding of when and why BTN1 expression will aid in understanding the expression of CLN3, which may be helpful in the treatment of this devastating disease.

Overcoming phagocytosis resistance of hypervirulent Klebsiella pneumoniae by directly targeting capsules
Tsubaki et al.This study highlights a promising strategy for disarming hypervirulent K. pneumoniae by directly targeting its key virulence factors and provides novel insights into antibacterial therapeutic approaches against this clinically significant pathogen.
Protein arginine methyltransferases in protozoan parasites: a new path for antiparasitic chemotherapy?
Campagnaro et al.This review discusses the activity and the relevance of arginine methyltransferases for the survival of pathogenic kinetoplastids, apicomplexans and amoebas, and how these enzymes could be exploited as drug targets.

VapA/Scs2 sustains polarized growth in Aspergillus nidulans by maintaining AP-2-mediated apical endocytosis
Georgiou et al.To explore the functional significance of ER–PM contact sites in filamentous fungi, we identified and genetically characterized all Aspergillus nidulans proteins homologous to Snc2/VAP, Ist2, or tricalbins.

Genetic make-up and regulation of the L-lysine biosynthesis pathway in Vibrio natriegens
Straube et al.This study analysed the make-up and regulation of the biosynthetic pathway for L-lysine and related L-aspartate family amino acids (AFAAs) in Vibrio natriegens DSM759 to provide a comprehensive basis for future metabolic engineering endeavours aiming at developing this strain into an amino acid overproducer.
Airborne bacteria in show caves from Southern Spain
This study analyzes the factors conditioning the diversity of airborne bacteria recorded in three Andalusian show caves, subjected to different managements.
Landscapes and bacterial signatures of mucosa-associated intestinal microbiota in Chilean and Spanish patients with inflammatory bowel disease
This study investigates the landscapes and alterations of mucosa-associated intestinal microbiota in patients with inflammatory bowel diseases, which cause chronic inflammation of the gut, including ulcerative colitis and Crohn’s disease.
Genome, transcriptome and secretome analyses of the antagonistic, yeast-like fungus Aureobasidium pullulans to identify potential biocontrol genes
This study highlights the value of a sequential approach starting with genome mining and consecutive transcriptome and secretome analyses in order to identify a limited number of potential target genes for detailed, functional analyses in Aureobasidium pullulans.
Proanthocyanidin-enriched cranberry extract induces resilient bacterial community dynamics in a gnotobiotic mouse model
This study investigates the effect of a water-soluble, proanthocyanidin-rich cranberry juice extract on the short-term dynamics of a human-derived bacterial community in a gnotobiotic mouse model.
Dry biocleaning of artwork: an innovative methodology for Cultural Heritage recovery?
This work proposes an innovative methodology based on applied biotechnology for the recovery of altered stonework: the “dry biocleaning”, which envisages the use of dehydrated microbial cells without the use of free water or gel-based matrices.
Aeration mitigates endoplasmic reticulum stress in Saccharomyces cerevisiae even without mitochondrial respiration
This work demonstrates a scenario, in which aeration acts beneficially on Saccharmyces cerevisiae cells even under fermentative conditions.
A novel BR-SMAD is required for larval development in barber’s pole worm Haemonchus contortus
The herein presented results show a BMP-like receptor-regulated SMAD in Haemonchus contortus that is required for larval differentiation and underscore an adaptive functional repurposing of BMP-signaling in parasitic worms.
Nutrient sensing and cAMP signaling in yeast: G-protein coupled receptor versus transceptor activation of PKA
The herein presented work supports a model, in which nutrient transceptors are evolutionary ancestors of GPCRs, employing a more primitive direct signaling mechanism compared to the indirect cAMP second-messenger signaling mechanism used by GPCRs for activation of PKA.
pH homeostasis links the nutrient sensing PKA/TORC1/Sch9 ménage-à-trois to stress tolerance and longevity
Deprez et al.In this article, Deprez et al. discuss accumulating evidence indicates that pH homeostasis plays a prominent role in the determination of ageing and longevity, thereby providing new perspectives and avenues to explore the underlying molecular mechanisms.
Guidelines and recommendations on yeast cell death nomenclature
Carmona-Gutierrez et al.In this review, we propose unified criteria for the definition of accidental, regulated, and programmed forms of cell death in yeast based on a series of morphological and biochemical criteria. Specifically, we provide consensus guidelines on the differential definition of terms including apoptosis, regulated necrosis, and autophagic cell death, as we refer to additional cell death routines that are relevant for the biology of yeast.
Burkholderia gladioli strain NGJ1 deploys a prophage tail-like protein for mycophagy
Kumar et al.In this article, the authors comment on the study “A prophage tail-like protein is deployed by Burkholderia bacteria to feed on fungi” by Swain et al. (Nature Communications, 2017), discussing that a prophage tail-like protein (Bg_9562) is essential for mycophagy. The protein may help the bacteria to survive in certain ecological niches and, considering its broad-spectrum antifungal activity, may be potentially useful in biotechnological applications to control fungal diseases.
Ras signalling in pathogenic yeasts
Pentland et al.In this article Pentland et al. review the roles of Ras protein function and signalling in the major human yeast pathogens Candida albicans and Cryptococcus neoformans and discuss the potential for targeting Ras as a novel approach to anti-fungal therapy.
A novel basolateral type IV secretion model for the CagA oncoprotein of Helicobacter pylori
Wessler and BackertIn this article, the authors comment on the study “Helicobacter pylori Employs a Unique Basolateral Type IV Secretion Mechanism for CagA Delivery” by Tegtmeyer et al. (Cell Host Microbe, 2017), discussing that the finding of a T4SS receptor suggests the presence of a sophisticated control mechanism for the injection of CagA and the possible impact of this novel signaling cascade on pathogenesis during infection with Helicobacter pylori.
A new role for the nuclear basket network
Gallardo et al.This article comments on work published by Salas-Pino et al. (J Cell Biol, 2017), which describes a novel function of the fission yeast nuclear basket component – the translocated promoter region (TPR) nucleoporin Alm1 – in proper localization of the proteasome to the nuclear envelope.
VAMP8 mucin exocytosis attenuates intestinal pathogenesis by Entamoeba histolytica
Cornick et al.This article comments on work published by Cornick et al. (mBio, 2017), which nominates SNARE-mediated exocytosis as the putative mechanism responsible for pathogen-induced mucus secretion from goblet cells.
Shutdown of interferon signaling by a viral-hijacked E3 ubiquitin ligase
Davis and PattonThis article comments on work published by Davis et al. (mBio, 2017), which describes molecular requirements that govern NSP1 recognition of β-TrCP, including an essential degron phosphorylation event, and the step-wise incorporation of NSP1 into hijacked cullin-RING E3 ligases (CRLs) that ubiquitinate and tag β-TrCP for degradation.
The emerging role of complex modifications of tRNALysUUU in signaling pathways
Patrick C. Thiaville and Valérie de Crécy-LagardThis comment discusses the article “Loss of wobble uridine modification in tRNA anticodons interferes with TOR pathway signaling” by Scheidt et al (Microbial Cell, 2014).
Only functional localization is faithful localization
Roland LillThis article comments on work published by Peleh et al. (Microbial Cell 2014), which analyzes the localization of Dre2 in Saccharomyces cerevisiae.
One cell, one love: a journal for microbial research
Didac Carmona-Gutierrez et al.In this inaugural article of Microbial Cell, we highlight the importance of microbial research in general and the journal’s intention to serve as a publishing forum that supports and enfolds the scientific diversity in this area as it provides a unique, high-quality and universally accessible source of information and inspiration.
What’s the role of autophagy in trypanosomes?
Katherine Figarella and Néstor L. UzcáteguiThis article comments on Proto et al. (Microbial Cell, 2014), who report first insights into the molecular mechanism of autophagy in African trypanosomes by generating reporter bloodstream form cell lines.
Can’t find what you’re looking for?
You can browse all our issues and published articles here.
FAQs
Peer-reviewed, open-access research using unicellular organisms (and multicellular microorganisms) to understand cellular responses and human disease.
The journal (founded in 2014) is led by its Editors-in-Chief Frank Madeo, Didac Carmona-Gutierrez, and Guido Kroemer
Microbial Cell has been publishing original scientific literature since 2014, and from the very beginning has been managed by active scientists through an independent Publishing House (Shared science Publishers). The journal was conceived as a platform to acknowledge the importance of unicellular organisms, both as model systems as well as in the biological context of human health and disease.
Ever since, Microbial Cell has very positively developed and strongly grown into a respected journal in the unicellular research community and even beyond. This scientific impact is reflected in the yearly number of citations obtained by articles published in Microbial Cell, as recorded by the Web of Science (Clarivate, formerly Thomson/Reuters):
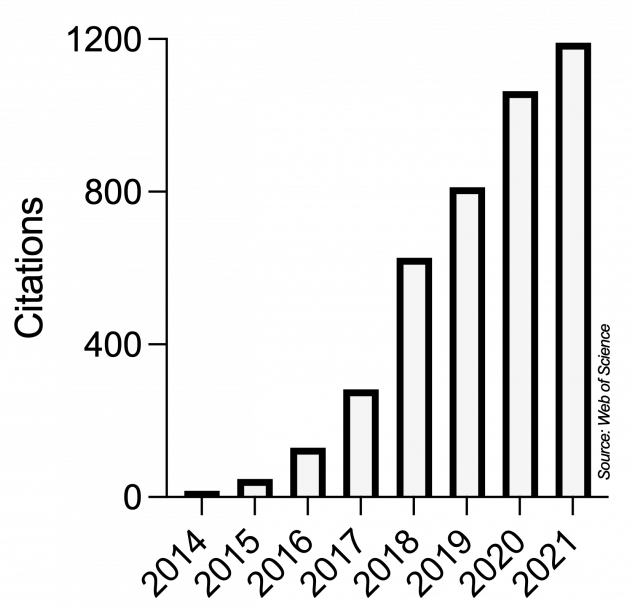
The scientific impact of Microbial Cell is also mirrored in a series of milestones:
2015: Microbial Cell is included in the Emerging Sources Citation Index (ESCI), a selection of developing journals drafted by Clarivate Analytics based on the candidate’s publishing standards, quality, editorial content, and citation data. Note: As an ESCI-selected journal, Microbial Cell is currently being evaluated in a rigorous and long process to determine an inclusion in the Science Citation Index Expanded (SCIE), which allows the official calculation of Clarivate Analytics’ impact factor.
2016: Microbial Cell is awarded the so-called DOAJ Seal by the selective Directory of Open Access Journals (DOAJ). The DOAJ Seal is an exclusive mark of certification for open access journals granted by DOAJ to journals that adhere to outstanding best practice and achieve an extra high and clear commitment to open access and high publishing standards.
2017: Microbial Cell is included in Pubmed Central (PMC), allowing the archiving of all the journal’s articles in PMC and PubMed.
2019: Microbial Cell is indexed in the prestigious abstract and citation database Scopus after a thorough selection process. This also means that Microbial Cell obtains, for the first time, an official Scopus CiteScore as well as an official journal ranking in the Scimago Journal and Country Ranking.
2022: Microbial Cell’s CiteScore reaches a value of 7.2 for the year 2021, positioning Microbial Cell among the top microbiology journals (previously available CiteScores: 2019: 5.4; 2020: 5.1).
2022: Microbial Cell is indexed in the highly selective Science Citation Index Expanded™, which covers approx. 9,500 of the world’s most impactful journals across 178 scientific disciplines. In their journal selection and curation process, Clarivate´s editors apply 24 ‘quality’ criteria and four ‘impact’ criteria to select the most influential journals in their respective fields. This selection is also a pre-requisite for inclusion in the JCR, which features the impact factor.
2022: Microbial Cell is listed in the Journal Citation Reports™ (JCR), and obtains its first official Journal Impact Factor™ (JIF) for the year 2021: 5.316.Check Article Types and Manuscript Preparation guidelines. Submit online via Scholastica.


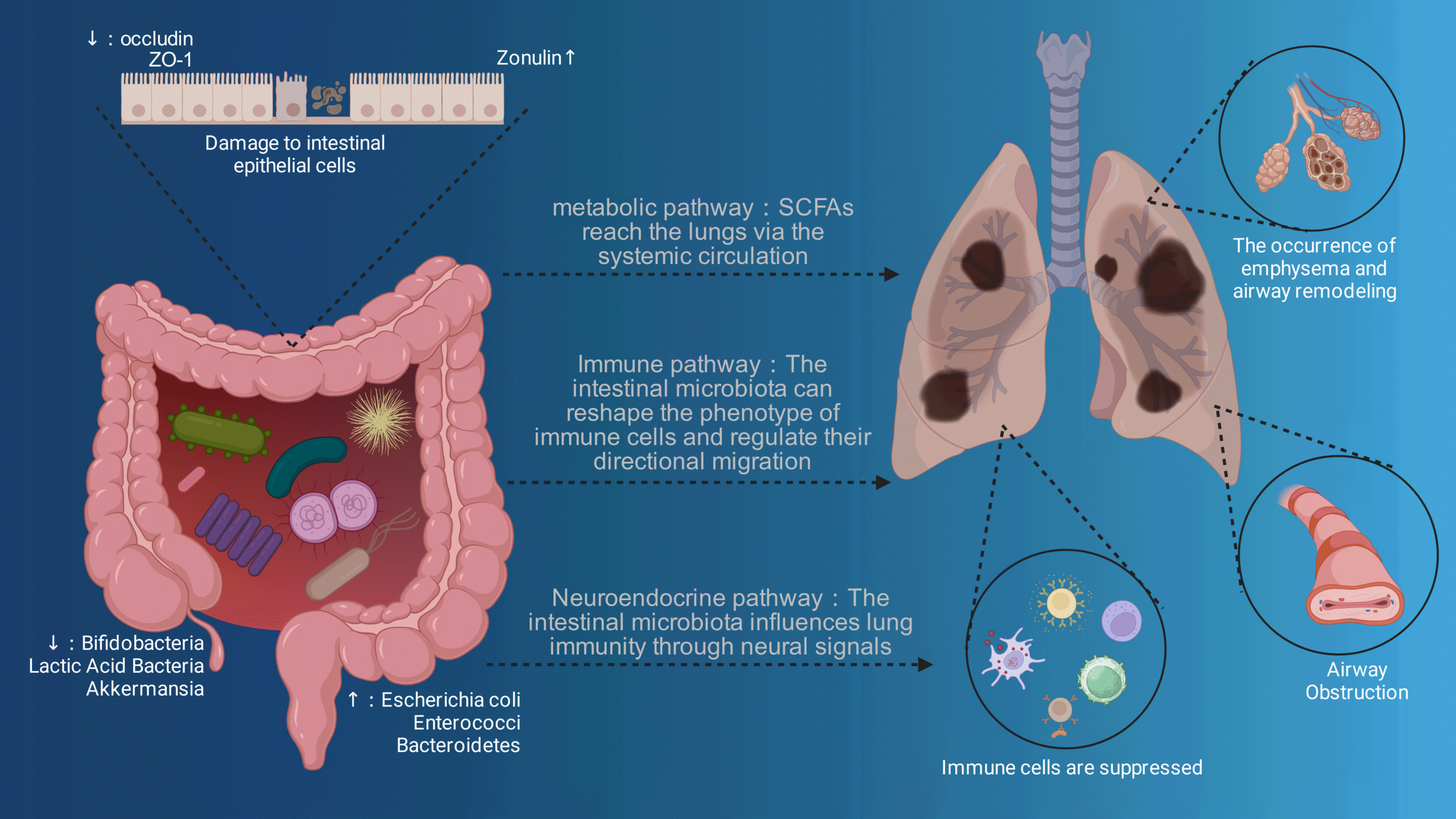

Metabolic pathways further increase the complexity of cell size control in budding yeast
Jorrit M. EnserinkThis article comments on work published by Soma et al. (Microbial Cell, 2014), which teased apart the effect of metabolism and growth rate on setting of critical cell size in Saccharomyces cerevisiae.