
Regulation of extracellular vesicles for protein secretion in Aspergillus nidulans
This study reveals that Aspergillus nidulans boosts extracellular vesicle production when ER-trafficked enzymes are induced, uncovering how fungi remodel their secretome through vesicle-mediated secretion to adapt to changing environments and biofilm formation.

Transcriptomic response to different heme sources in Trypanosoma cruzi epimastigotes
This study uncovers how the Chagas disease parasite adapts to changes in heme, an essential molecule for its survival, providing transcriptional clues to heme metabolism and identifying a previously unreported heme-binding protein in T. cruzi.

Luminal acetylation of microtubules is not essential for Plasmodium berghei and Toxoplasma gondii survival
Acetylation of α-tubulin at lysine 40 is not essential for cytoskeletal stability in Plasmodium berghei or Toxoplasma gondii, suggesting redundancy and plasticity in microtubule regulation in these parasites.

The dual-site agonist for human M2 muscarinic receptors Iper-8-naphtalimide induces mitochondrial dysfunction in Saccharomyces cerevisiae
S. cerevisiae is a model to study human GPCRs. N-8-Iper, active against glioblastoma via M2 receptor, causes mitochondrial damage in yeast by binding Ste2, highlighting evolutionary conservation of GPCRs.

Integrative Omics reveals changes in the cellular landscape of peroxisome-deficient pex3 yeast cells
To uncover the consequences of peroxisome deficiency, we compared Saccharomyces cerevisiae wild-type with pex3 cells, which lack peroxisomes, employing quantitative proteomics and transcriptomics technologies.

The mechanism of Tat-dependent protein translocation
Brüser and SandersThis review integrates mechanistically relevant biochemical, molecular, and structural studies on Tat-dependent translocation of folded proteins into an in its molecular detail new comprehensive explanation of how the Tat system mediates protein transport.

TOR-dependent regulation of the yeast homolog of the juvenile Batten Disease-associated gene CLN3
Pillalamarri et al.This study identifies conditions and genes that induce BTN1 expression in yeast. We show that BTN1 expression is regulated by translational control and by the mTOR1 pathway. An understanding of when and why BTN1 expression will aid in understanding the expression of CLN3, which may be helpful in the treatment of this devastating disease.

Overcoming phagocytosis resistance of hypervirulent Klebsiella pneumoniae by directly targeting capsules
Tsubaki et al.This study highlights a promising strategy for disarming hypervirulent K. pneumoniae by directly targeting its key virulence factors and provides novel insights into antibacterial therapeutic approaches against this clinically significant pathogen.
Protein arginine methyltransferases in protozoan parasites: a new path for antiparasitic chemotherapy?
Campagnaro et al.This review discusses the activity and the relevance of arginine methyltransferases for the survival of pathogenic kinetoplastids, apicomplexans and amoebas, and how these enzymes could be exploited as drug targets.

VapA/Scs2 sustains polarized growth in Aspergillus nidulans by maintaining AP-2-mediated apical endocytosis
Georgiou et al.To explore the functional significance of ER–PM contact sites in filamentous fungi, we identified and genetically characterized all Aspergillus nidulans proteins homologous to Snc2/VAP, Ist2, or tricalbins.

Genetic make-up and regulation of the L-lysine biosynthesis pathway in Vibrio natriegens
Straube et al.This study analysed the make-up and regulation of the biosynthetic pathway for L-lysine and related L-aspartate family amino acids (AFAAs) in Vibrio natriegens DSM759 to provide a comprehensive basis for future metabolic engineering endeavours aiming at developing this strain into an amino acid overproducer.
Airborne bacteria in show caves from Southern Spain
This study analyzes the factors conditioning the diversity of airborne bacteria recorded in three Andalusian show caves, subjected to different managements.
Landscapes and bacterial signatures of mucosa-associated intestinal microbiota in Chilean and Spanish patients with inflammatory bowel disease
This study investigates the landscapes and alterations of mucosa-associated intestinal microbiota in patients with inflammatory bowel diseases, which cause chronic inflammation of the gut, including ulcerative colitis and Crohn’s disease.
Genome, transcriptome and secretome analyses of the antagonistic, yeast-like fungus Aureobasidium pullulans to identify potential biocontrol genes
This study highlights the value of a sequential approach starting with genome mining and consecutive transcriptome and secretome analyses in order to identify a limited number of potential target genes for detailed, functional analyses in Aureobasidium pullulans.
Proanthocyanidin-enriched cranberry extract induces resilient bacterial community dynamics in a gnotobiotic mouse model
This study investigates the effect of a water-soluble, proanthocyanidin-rich cranberry juice extract on the short-term dynamics of a human-derived bacterial community in a gnotobiotic mouse model.
Dry biocleaning of artwork: an innovative methodology for Cultural Heritage recovery?
This work proposes an innovative methodology based on applied biotechnology for the recovery of altered stonework: the “dry biocleaning”, which envisages the use of dehydrated microbial cells without the use of free water or gel-based matrices.
Aeration mitigates endoplasmic reticulum stress in Saccharomyces cerevisiae even without mitochondrial respiration
This work demonstrates a scenario, in which aeration acts beneficially on Saccharmyces cerevisiae cells even under fermentative conditions.
A novel BR-SMAD is required for larval development in barber’s pole worm Haemonchus contortus
The herein presented results show a BMP-like receptor-regulated SMAD in Haemonchus contortus that is required for larval differentiation and underscore an adaptive functional repurposing of BMP-signaling in parasitic worms.
Nutrient sensing and cAMP signaling in yeast: G-protein coupled receptor versus transceptor activation of PKA
The herein presented work supports a model, in which nutrient transceptors are evolutionary ancestors of GPCRs, employing a more primitive direct signaling mechanism compared to the indirect cAMP second-messenger signaling mechanism used by GPCRs for activation of PKA.
Regulation of anti-microbial autophagy by factors of the complement system
Viret et al.This review explores emerging evidence that components of the complement system, beyond their traditional immune roles, modulate autophagy – particularly xenophagy – thereby influencing cell-autonomous antimicrobial responses during host-pathogen interactions.
More than flipping the lid: Cdc50 contributes to echinocandin resistance by regulating calcium homeostasis in Cryptococcus neoformans
Cao and XueIn this article, the authors comment on the study “A mechanosensitive channel governs lipid flippase-mediated echinocandin resistance in Cryptococcus neoformans” by Cao et al. (mBio, 2019), which uncovers a dual role for the lipid flippase subunit Cdc50 in Cryptococcus neoformans, linking lipid translocation and calcium signaling via its interaction with the mechanosensitive channel Crm1, thereby contributing to innate resistance against the antifungal drug caspofungin.
New insights in the mode of action of anti-leishmanial drugs by using chemical mutagenesis screens coupled to next-generation sequencing
Bhattacharya et al.In this article, the authors comment on the study “Coupling chemical mutagenesis to next generation sequencing for the identification of drug resistance mutations in Leishmania” by Bhattacharya et al. (Nat Commun, 2019), which introduces Mut-seq, a chemical mutagenesis and sequencing approach, to uncover drug resistance mechanisms in Leishmania, revealing links between lipid metabolism genes and miltefosine resistance, and a protein kinase involved in translation conferring paromomycin resistance.
Microfluidic techniques for separation of bacterial cells via taxis
Gurung et al.Microfluidic tools, ideal for studying microbial motility due to their control over laminar flows at microscopic scales, enable precise analysis of various taxis behaviors and have advanced applications in synthetic biology, directed evolution, and medical microbiology.
Influence of delivery and feeding mode in oral fungi colonization – a systematic review
Azevedo et al.A systematic review of oral fungal colonization in infants found that while breastfeeding did not significantly affect the oral mycobiome, vaginal delivery was associated with higher oral yeast colonization, particularly of Candida albicans.
A holobiont view on thrombosis: unravelling the microbiota’s influence on arterial thrombus growth
Pontarollo et al.In this article, the authors comment on the study “The microbiota promotes arterial thrombosis in low-density lipoprotein receptor-deficient mice” by Kiouptsi et al. (mBio, 2019) that showed that commensal microbiota, intricately linked to host physiology, may influence cardiovascular disease, as shown by studies using germ-free atherosclerosis-prone mice to examine how microbial presence and diet affect arterial thrombosis and lesion development.
The role of Lactobacillus species in the control of Candida via biotrophic interactions
Zangl et al.Microbial communities, including Candida and Lactobacillus species, play a crucial role in human health, particularly in the context of mucosal infections, but our understanding of their interactions and effects is still incomplete due to the variability of species and isolates as well as the complexity of the human host.
Tribal warfare: Commensal Neisseria kill pathogen Neisseria gonorrhoeae using its DNA
So and RendónThis article comments on work published by Kim et al (Cell Host Microbe, 2019), which adds a new dimension to the concept of commensal protection. It shows that commensal Neisseria kill the closely related pathogen N. gonorrhoeae through an unexpected mechanism, one that involves genetic competence, DNA methylation state and recombination.
Targeting GATA transcription factors – a novel strategy for anti-aging interventions?
Zimmermann et al.This article comments on work published by Carmona-Gutierrez et al. (Nat Commun., 2019), which identified a natural compound, 4,4′-dimethoxychalcone, inducing autophagy and prolonging lifespan in different organisms through a mechanism that involves GATA transcription factors.
In the beginning was the word: How terminology drives our understanding of endosymbiotic organelles
OborníkThis In the Pit article argues that the naming conventions for biological entities influence research perspectives and methodologies, advocating for mitochondria and plastids to be classified and named as bacteria due to their endosymbiotic origins, with potential implications for our understanding of bacterial prevalence, definitions of the microbiome and multicellularity, and the concept of endosymbiotic domestication.
What’s in a name? How organelles of endosymbiotic origin can be distinguished from endosymbionts
GruberThis In the Pit article suggests redefining the relationship between hosts and endosymbionts, like mitochondria and plastids, as a single species based on “sexual symbiont integration,” the loss of independent speciation, and congruence in genetic recombination and population sizes, rather than solely on historic classifications or structural properties.
Microbial wars: competition in ecological niches and within the microbiome
Bauer et al.In this Editorial Bauer et al. provide a brief overview on microbial competition and discuss some of its roles and consequences that directly affect humans.
Exploring the mechanism of amebic trogocytosis: the role of amebic lysosomes
Gilmartin and PetriIn this article, the authors comment on the study “Inhibition of Amebic Lysosomal Acidification Blocks Amebic Trogocytosis and Cell Killing” by Gilmartin et al. (MBio, 2017), discussing the the role of amebic lysosomes in Trogocytosis, the intracellular transfer of fragments of cell material.
Uncovering the hidden: complexity and strategies for diagnosing latent tuberculosis
Flores-ValdezThis editorial postulates that advanced proteomic and transcriptomic techniques are evolving and may enhance the detection of latent tuberculosis, thereby distinguishing true M. tuberculosis infections from other conditions, which is vital for controlling potential reactivation and transmission.
The Yin & Yang of Mitochondrial Architecture – Interplay of MICOS and F1Fo-ATP synthase in cristae formation
Rampelt and van der LaanThis Editorial posits that mitochondrial cristae architecture is shaped by the interplay of MICOS and ATP synthase, with a recent study illuminating their roles in cristae formation and maintenance.
When a ribosomal protein grows up – the ribosome assembly path of Rps3
Brigitte PertschyThis article comments on two papers by Mitterer et al., which followed yeast protein Rps3, highlighting the sophisticated mechanisms for protein protection, nuclear transport, and integration into pre-ribosomal particles for final assembly with 40S subunits.
Can’t find what you’re looking for?
You can browse all our issues and published articles here.
FAQs
Peer-reviewed, open-access research using unicellular organisms (and multicellular microorganisms) to understand cellular responses and human disease.
The journal (founded in 2014) is led by its Editors-in-Chief Frank Madeo, Didac Carmona-Gutierrez, and Guido Kroemer
Microbial Cell has been publishing original scientific literature since 2014, and from the very beginning has been managed by active scientists through an independent Publishing House (Shared science Publishers). The journal was conceived as a platform to acknowledge the importance of unicellular organisms, both as model systems as well as in the biological context of human health and disease.
Ever since, Microbial Cell has very positively developed and strongly grown into a respected journal in the unicellular research community and even beyond. This scientific impact is reflected in the yearly number of citations obtained by articles published in Microbial Cell, as recorded by the Web of Science (Clarivate, formerly Thomson/Reuters):
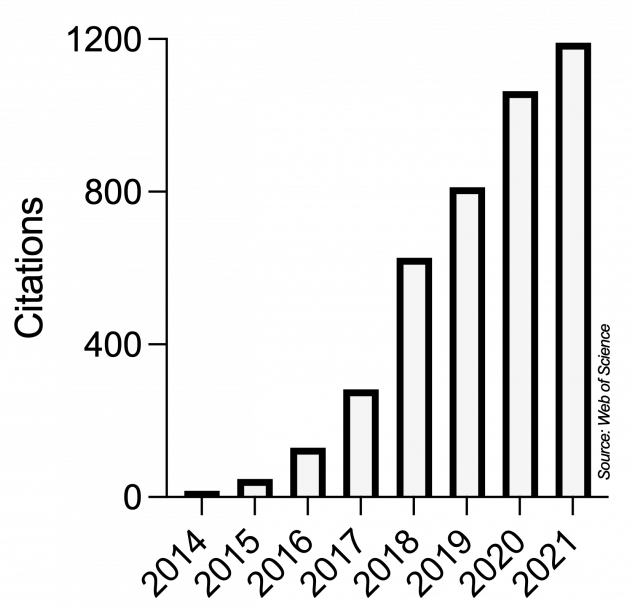
The scientific impact of Microbial Cell is also mirrored in a series of milestones:
2015: Microbial Cell is included in the Emerging Sources Citation Index (ESCI), a selection of developing journals drafted by Clarivate Analytics based on the candidate’s publishing standards, quality, editorial content, and citation data. Note: As an ESCI-selected journal, Microbial Cell is currently being evaluated in a rigorous and long process to determine an inclusion in the Science Citation Index Expanded (SCIE), which allows the official calculation of Clarivate Analytics’ impact factor.
2016: Microbial Cell is awarded the so-called DOAJ Seal by the selective Directory of Open Access Journals (DOAJ). The DOAJ Seal is an exclusive mark of certification for open access journals granted by DOAJ to journals that adhere to outstanding best practice and achieve an extra high and clear commitment to open access and high publishing standards.
2017: Microbial Cell is included in Pubmed Central (PMC), allowing the archiving of all the journal’s articles in PMC and PubMed.
2019: Microbial Cell is indexed in the prestigious abstract and citation database Scopus after a thorough selection process. This also means that Microbial Cell obtains, for the first time, an official Scopus CiteScore as well as an official journal ranking in the Scimago Journal and Country Ranking.
2022: Microbial Cell’s CiteScore reaches a value of 7.2 for the year 2021, positioning Microbial Cell among the top microbiology journals (previously available CiteScores: 2019: 5.4; 2020: 5.1).
2022: Microbial Cell is indexed in the highly selective Science Citation Index Expanded™, which covers approx. 9,500 of the world’s most impactful journals across 178 scientific disciplines. In their journal selection and curation process, Clarivate´s editors apply 24 ‘quality’ criteria and four ‘impact’ criteria to select the most influential journals in their respective fields. This selection is also a pre-requisite for inclusion in the JCR, which features the impact factor.
2022: Microbial Cell is listed in the Journal Citation Reports™ (JCR), and obtains its first official Journal Impact Factor™ (JIF) for the year 2021: 5.316.Check Article Types and Manuscript Preparation guidelines. Submit online via Scholastica.



Sulfur dioxide resistance in Saccharomyces cerevisiae: beyond SSU1
García-Ríos and GuillamónThis article discusses the importance of understanding sulfite resistance in Saccharomyces cerevisiae due to its use in winemaking and the potential role of the transcription factor Com2. While the SSU1 gene and its activity have been correlated with sulfite tolerance, the work by Lage et al. (2019) indicates that Com2 might control a large percentage of the genes activated by SO2 and contribute to the yeast’s protective response, offering new insights into the molecular factors influencing this oenological trait.